- How it works


Academic Assignment Samples and Examples
Are you looking for someone to write your academic assignment for you? This is the right place for you. To showcase the quality of the work that can be expected from ResearchProspect, we have curated a few samples of academic assignments. These examples have been developed by professional writers here. Place your order with us now.
Assignment Sample
Discipline: Sociology
Quality: Approved / Passed
Discipline: Construction
Quality: 1st / 78%
Discipline: Accounting & Finance
Quality: 2:1 / 69%
Undergraduate
Discipline: Bio-Medical
Quality: 1st / 76%
Discipline: Statistics
Quality: 1st / 73%
Discipline: Health and Safety
Quality: 2:1 / 68%
Discipline: Business
Quality: 2:1 / 67%
Discipline: Medicine
Quality: 2:1 / 66%
Discipline: Religion Theology
Quality: 2:1 / 64%
Discipline: Project Management
Quality: 2:1 / 63%
Discipline: Website Development
Discipline: Fire and Construction
Discipline: Environmental Management
Discipline: Early Child Education
Quality: 1st / 72%
Analysis of a Business Environment: Coffee and Cake Ltd (CC Ltd)
Business Strategy
Application of Project Management Using the Agile Approach ….
Project Management
Assessment of British Airways Social Media Posts
Critical annotation, global business environment (reflective report assignment), global marketing strategies, incoterms, ex (exw), free (fob, fca), cost (cpt, cip), delivery …., it systems strategy – the case of oxford university, management and organisation in global environment, marketing plan for “b airlines”, prepare a portfolio review and remedial options and actions …., systematic identification, analysis, and assessment of risk …., the exploratory problem-solving play and growth mindset for …..
Childhood Development
The Marketing Plan- UK Sustainable Energy Limited
Law assignment.
Law Case Study
To Analyse User’s Perception towards the Services Provided by Their…
Assignment Samples
Research Methodology
Discipline: Civil Engineering
Discipline: Health & Manangement
Our Assignment Writing Service Features
Subject specialists.
We have writers specialising in their respective fields to ensure rigorous quality control.
We are reliable as we deliver all your work to you and do not use it in any future work.
We ensure that our work is 100% plagiarism free and authentic and all references are cited.
Thoroughly Researched
We perform thorough research to get accurate content for you with proper citations.
Excellent Customer Service
To resolve your issues and queries, we provide 24/7 customer service
Our prices are kept at a level that is affordable for everyone to ensure maximum help.
Loved by over 100,000 students
Thousands of students have used ResearchProspect academic support services to improve their grades. Why are you waiting?
"I am glad I gave my order to ResearchProspect after seeing their academic assignment sample. Really happy with the results. "

Law Student
"I am grateful to them for doing my academic assignment. Got high grades."

Economics Student
Frequently Ask Questions?
How can these samples help you.
The assignment writing samples we provide help you by showing you versions of the finished item. It’s like having a picture of the cake you’re aiming to make when following a recipe.
Assignments that you undertake are a key part of your academic life; they are the usual way of assessing your knowledge on the subject you’re studying.
There are various types of assignments: essays, annotated bibliographies, stand-alone literature reviews, reflective writing essays, etc. There will be a specific structure to follow for each of these. Before focusing on the structure, it is best to plan your assignment first. Your school will have its own guidelines and instructions, you should align with those. Start by selecting the essential aspects that need to be included in your assignment.
Based on what you understand from the assignment in question, evaluate the critical points that should be made. If the task is research-based, discuss your aims and objectives, research method, and results. For an argumentative essay, you need to construct arguments relevant to the thesis statement.
Your assignment should be constructed according to the outline’s different sections. This is where you might find our samples so helpful; inspect them to understand how to write your assignment.
Adding headings to sections can enhance the clarity of your assignment. They are like signposts telling the reader what’s coming next.
Where structure is concerned, our samples can be of benefit. The basic structure is of three parts: introduction, discussion, and conclusion. It is, however, advisable to follow the structural guidelines from your tutor.
For example, our master’s sample assignment includes lots of headings and sub-headings. Undergraduate assignments are shorter and present a statistical analysis only.
If you are still unsure about how to approach your assignment, we are here to help, and we really can help you. You can start by just asking us a question with no need to commit. Our writers are able to assist by guiding you through every step of your assignment.
Who will write my assignment?
We have a cherry-picked writing team. They’ve been thoroughly tested and checked out to verify their skills and credentials. You can be sure our writers have proved they can write for you.
What if I have an urgent assignment? Do your delivery days include the weekends?
No problem. Christmas, Boxing Day, New Year’s Eve – our only days off. We know you want weekend delivery, so this is what we do.
Explore More Samples
View our professional samples to be certain that we have the portofilio and capabilities to deliver what you need.
USEFUL LINKS
LEARNING RESOURCES

COMPANY DETAILS

- How It Works

Sample Papers
This page contains sample papers formatted in seventh edition APA Style. The sample papers show the format that authors should use to submit a manuscript for publication in a professional journal and that students should use to submit a paper to an instructor for a course assignment. You can download the Word files to use as templates and edit them as needed for the purposes of your own papers.
Most guidelines in the Publication Manual apply to both professional manuscripts and student papers. However, there are specific guidelines for professional papers versus student papers, including professional and student title page formats. All authors should check with the person or entity to whom they are submitting their paper (e.g., publisher or instructor) for guidelines that are different from or in addition to those specified by APA Style.
Sample papers from the Publication Manual
The following two sample papers were published in annotated form in the Publication Manual and are reproduced here as PDFs for your ease of use. The annotations draw attention to content and formatting and provide the relevant sections of the Publication Manual (7th ed.) to consult for more information.
- Student sample paper with annotations (PDF, 5MB)
- Professional sample paper with annotations (PDF, 2.7MB)
We also offer these sample papers in Microsoft Word (.docx) format with the annotations as comments to the text.
- Student sample paper with annotations as comments (DOCX, 42KB)
- Professional sample paper with annotations as comments (DOCX, 103KB)
Finally, we offer these sample papers in Microsoft Word (.docx) format without the annotations.
- Student sample paper without annotations (DOCX, 36KB)
- Professional sample paper without annotations (DOCX, 96KB)
Sample professional paper templates by paper type
These sample papers demonstrate APA Style formatting standards for different professional paper types. Professional papers can contain many different elements depending on the nature of the work. Authors seeking publication should refer to the journal’s instructions for authors or manuscript submission guidelines for specific requirements and/or sections to include.
- Literature review professional paper template (DOCX, 47KB)
- Mixed methods professional paper template (DOCX, 68KB)
- Qualitative professional paper template (DOCX, 72KB)
- Quantitative professional paper template (DOCX, 77KB)
- Review professional paper template (DOCX, 112KB)
Sample papers are covered in the seventh edition APA Style manuals in the Publication Manual Chapter 2 and the Concise Guide Chapter 1
Related handouts
- Heading Levels Template: Student Paper (PDF, 257KB)
- Heading Levels Template: Professional Paper (PDF, 213KB)
Other instructional aids
- Journal Article Reporting Standards (JARS)
- APA Style Tutorials and Webinars
- Handouts and Guides
- Paper Format
View all instructional aids
Sample student paper templates by paper type
These sample papers demonstrate APA Style formatting standards for different student paper types. Students may write the same types of papers as professional authors (e.g., quantitative studies, literature reviews) or other types of papers for course assignments (e.g., reaction or response papers, discussion posts), dissertations, and theses.
APA does not set formal requirements for the nature or contents of an APA Style student paper. Students should follow the guidelines and requirements of their instructor, department, and/or institution when writing papers. For instance, an abstract and keywords are not required for APA Style student papers, although an instructor may request them in student papers that are longer or more complex. Specific questions about a paper being written for a course assignment should be directed to the instructor or institution assigning the paper.
- Discussion post student paper template (DOCX, 31KB)
- Literature review student paper template (DOCX, 37KB)
- Quantitative study student paper template (DOCX, 53KB)
Sample papers in real life
Although published articles differ in format from manuscripts submitted for publication or student papers (e.g., different line spacing, font, margins, and column format), articles published in APA journals provide excellent demonstrations of APA Style in action.
APA journals began publishing papers in seventh edition APA Style in 2020. Professional authors should check the author submission guidelines for the journal to which they want to submit their paper for any journal-specific style requirements.
Credits for sample professional paper templates
Quantitative professional paper template: Adapted from “Fake News, Fast and Slow: Deliberation Reduces Belief in False (but Not True) News Headlines,” by B. Bago, D. G. Rand, and G. Pennycook, 2020, Journal of Experimental Psychology: General , 149 (8), pp. 1608–1613 ( https://doi.org/10.1037/xge0000729 ). Copyright 2020 by the American Psychological Association.
Qualitative professional paper template: Adapted from “‘My Smartphone Is an Extension of Myself’: A Holistic Qualitative Exploration of the Impact of Using a Smartphone,” by L. J. Harkin and D. Kuss, 2020, Psychology of Popular Media , 10 (1), pp. 28–38 ( https://doi.org/10.1037/ppm0000278 ). Copyright 2020 by the American Psychological Association.
Mixed methods professional paper template: Adapted from “‘I Am a Change Agent’: A Mixed Methods Analysis of Students’ Social Justice Value Orientation in an Undergraduate Community Psychology Course,” by D. X. Henderson, A. T. Majors, and M. Wright, 2019, Scholarship of Teaching and Learning in Psychology , 7 (1), 68–80. ( https://doi.org/10.1037/stl0000171 ). Copyright 2019 by the American Psychological Association.
Literature review professional paper template: Adapted from “Rethinking Emotions in the Context of Infants’ Prosocial Behavior: The Role of Interest and Positive Emotions,” by S. I. Hammond and J. K. Drummond, 2019, Developmental Psychology , 55 (9), pp. 1882–1888 ( https://doi.org/10.1037/dev0000685 ). Copyright 2019 by the American Psychological Association.
Review professional paper template: Adapted from “Joining the Conversation: Teaching Students to Think and Communicate Like Scholars,” by E. L. Parks, 2022, Scholarship of Teaching and Learning in Psychology , 8 (1), pp. 70–78 ( https://doi.org/10.1037/stl0000193 ). Copyright 2020 by the American Psychological Association.
Credits for sample student paper templates
These papers came from real students who gave their permission to have them edited and posted by APA.
- Privacy Policy

Home » Assignment – Types, Examples and Writing Guide
Assignment – Types, Examples and Writing Guide
Table of Contents

Definition:
Assignment is a task given to students by a teacher or professor, usually as a means of assessing their understanding and application of course material. Assignments can take various forms, including essays, research papers, presentations, problem sets, lab reports, and more.
Assignments are typically designed to be completed outside of class time and may require independent research, critical thinking, and analysis. They are often graded and used as a significant component of a student’s overall course grade. The instructions for an assignment usually specify the goals, requirements, and deadlines for completion, and students are expected to meet these criteria to earn a good grade.
History of Assignment
The use of assignments as a tool for teaching and learning has been a part of education for centuries. Following is a brief history of the Assignment.
- Ancient Times: Assignments such as writing exercises, recitations, and memorization tasks were used to reinforce learning.
- Medieval Period : Universities began to develop the concept of the assignment, with students completing essays, commentaries, and translations to demonstrate their knowledge and understanding of the subject matter.
- 19th Century : With the growth of schools and universities, assignments became more widespread and were used to assess student progress and achievement.
- 20th Century: The rise of distance education and online learning led to the further development of assignments as an integral part of the educational process.
- Present Day: Assignments continue to be used in a variety of educational settings and are seen as an effective way to promote student learning and assess student achievement. The nature and format of assignments continue to evolve in response to changing educational needs and technological innovations.
Types of Assignment
Here are some of the most common types of assignments:
An essay is a piece of writing that presents an argument, analysis, or interpretation of a topic or question. It usually consists of an introduction, body paragraphs, and a conclusion.
Essay structure:
- Introduction : introduces the topic and thesis statement
- Body paragraphs : each paragraph presents a different argument or idea, with evidence and analysis to support it
- Conclusion : summarizes the key points and reiterates the thesis statement
Research paper
A research paper involves gathering and analyzing information on a particular topic, and presenting the findings in a well-structured, documented paper. It usually involves conducting original research, collecting data, and presenting it in a clear, organized manner.
Research paper structure:
- Title page : includes the title of the paper, author’s name, date, and institution
- Abstract : summarizes the paper’s main points and conclusions
- Introduction : provides background information on the topic and research question
- Literature review: summarizes previous research on the topic
- Methodology : explains how the research was conducted
- Results : presents the findings of the research
- Discussion : interprets the results and draws conclusions
- Conclusion : summarizes the key findings and implications
A case study involves analyzing a real-life situation, problem or issue, and presenting a solution or recommendations based on the analysis. It often involves extensive research, data analysis, and critical thinking.
Case study structure:
- Introduction : introduces the case study and its purpose
- Background : provides context and background information on the case
- Analysis : examines the key issues and problems in the case
- Solution/recommendations: proposes solutions or recommendations based on the analysis
- Conclusion: Summarize the key points and implications
A lab report is a scientific document that summarizes the results of a laboratory experiment or research project. It typically includes an introduction, methodology, results, discussion, and conclusion.
Lab report structure:
- Title page : includes the title of the experiment, author’s name, date, and institution
- Abstract : summarizes the purpose, methodology, and results of the experiment
- Methods : explains how the experiment was conducted
- Results : presents the findings of the experiment
Presentation
A presentation involves delivering information, data or findings to an audience, often with the use of visual aids such as slides, charts, or diagrams. It requires clear communication skills, good organization, and effective use of technology.
Presentation structure:
- Introduction : introduces the topic and purpose of the presentation
- Body : presents the main points, findings, or data, with the help of visual aids
- Conclusion : summarizes the key points and provides a closing statement
Creative Project
A creative project is an assignment that requires students to produce something original, such as a painting, sculpture, video, or creative writing piece. It allows students to demonstrate their creativity and artistic skills.
Creative project structure:
- Introduction : introduces the project and its purpose
- Body : presents the creative work, with explanations or descriptions as needed
- Conclusion : summarizes the key elements and reflects on the creative process.
Examples of Assignments
Following are Examples of Assignment templates samples:
Essay template:
I. Introduction
- Hook: Grab the reader’s attention with a catchy opening sentence.
- Background: Provide some context or background information on the topic.
- Thesis statement: State the main argument or point of your essay.
II. Body paragraphs
- Topic sentence: Introduce the main idea or argument of the paragraph.
- Evidence: Provide evidence or examples to support your point.
- Analysis: Explain how the evidence supports your argument.
- Transition: Use a transition sentence to lead into the next paragraph.
III. Conclusion
- Restate thesis: Summarize your main argument or point.
- Review key points: Summarize the main points you made in your essay.
- Concluding thoughts: End with a final thought or call to action.
Research paper template:
I. Title page
- Title: Give your paper a descriptive title.
- Author: Include your name and institutional affiliation.
- Date: Provide the date the paper was submitted.
II. Abstract
- Background: Summarize the background and purpose of your research.
- Methodology: Describe the methods you used to conduct your research.
- Results: Summarize the main findings of your research.
- Conclusion: Provide a brief summary of the implications and conclusions of your research.
III. Introduction
- Background: Provide some background information on the topic.
- Research question: State your research question or hypothesis.
- Purpose: Explain the purpose of your research.
IV. Literature review
- Background: Summarize previous research on the topic.
- Gaps in research: Identify gaps or areas that need further research.
V. Methodology
- Participants: Describe the participants in your study.
- Procedure: Explain the procedure you used to conduct your research.
- Measures: Describe the measures you used to collect data.
VI. Results
- Quantitative results: Summarize the quantitative data you collected.
- Qualitative results: Summarize the qualitative data you collected.
VII. Discussion
- Interpretation: Interpret the results and explain what they mean.
- Implications: Discuss the implications of your research.
- Limitations: Identify any limitations or weaknesses of your research.
VIII. Conclusion
- Review key points: Summarize the main points you made in your paper.
Case study template:
- Background: Provide background information on the case.
- Research question: State the research question or problem you are examining.
- Purpose: Explain the purpose of the case study.
II. Analysis
- Problem: Identify the main problem or issue in the case.
- Factors: Describe the factors that contributed to the problem.
- Alternative solutions: Describe potential solutions to the problem.
III. Solution/recommendations
- Proposed solution: Describe the solution you are proposing.
- Rationale: Explain why this solution is the best one.
- Implementation: Describe how the solution can be implemented.
IV. Conclusion
- Summary: Summarize the main points of your case study.
Lab report template:
- Title: Give your report a descriptive title.
- Date: Provide the date the report was submitted.
- Background: Summarize the background and purpose of the experiment.
- Methodology: Describe the methods you used to conduct the experiment.
- Results: Summarize the main findings of the experiment.
- Conclusion: Provide a brief summary of the implications and conclusions
- Background: Provide some background information on the experiment.
- Hypothesis: State your hypothesis or research question.
- Purpose: Explain the purpose of the experiment.
IV. Materials and methods
- Materials: List the materials and equipment used in the experiment.
- Procedure: Describe the procedure you followed to conduct the experiment.
- Data: Present the data you collected in tables or graphs.
- Analysis: Analyze the data and describe the patterns or trends you observed.
VI. Discussion
- Implications: Discuss the implications of your findings.
- Limitations: Identify any limitations or weaknesses of the experiment.
VII. Conclusion
- Restate hypothesis: Summarize your hypothesis or research question.
- Review key points: Summarize the main points you made in your report.
Presentation template:
- Attention grabber: Grab the audience’s attention with a catchy opening.
- Purpose: Explain the purpose of your presentation.
- Overview: Provide an overview of what you will cover in your presentation.
II. Main points
- Main point 1: Present the first main point of your presentation.
- Supporting details: Provide supporting details or evidence to support your point.
- Main point 2: Present the second main point of your presentation.
- Main point 3: Present the third main point of your presentation.
- Summary: Summarize the main points of your presentation.
- Call to action: End with a final thought or call to action.
Creative writing template:
- Setting: Describe the setting of your story.
- Characters: Introduce the main characters of your story.
- Rising action: Introduce the conflict or problem in your story.
- Climax: Present the most intense moment of the story.
- Falling action: Resolve the conflict or problem in your story.
- Resolution: Describe how the conflict or problem was resolved.
- Final thoughts: End with a final thought or reflection on the story.
How to Write Assignment
Here is a general guide on how to write an assignment:
- Understand the assignment prompt: Before you begin writing, make sure you understand what the assignment requires. Read the prompt carefully and make note of any specific requirements or guidelines.
- Research and gather information: Depending on the type of assignment, you may need to do research to gather information to support your argument or points. Use credible sources such as academic journals, books, and reputable websites.
- Organize your ideas : Once you have gathered all the necessary information, organize your ideas into a clear and logical structure. Consider creating an outline or diagram to help you visualize your ideas.
- Write a draft: Begin writing your assignment using your organized ideas and research. Don’t worry too much about grammar or sentence structure at this point; the goal is to get your thoughts down on paper.
- Revise and edit: After you have written a draft, revise and edit your work. Make sure your ideas are presented in a clear and concise manner, and that your sentences and paragraphs flow smoothly.
- Proofread: Finally, proofread your work for spelling, grammar, and punctuation errors. It’s a good idea to have someone else read over your assignment as well to catch any mistakes you may have missed.
- Submit your assignment : Once you are satisfied with your work, submit your assignment according to the instructions provided by your instructor or professor.
Applications of Assignment
Assignments have many applications across different fields and industries. Here are a few examples:
- Education : Assignments are a common tool used in education to help students learn and demonstrate their knowledge. They can be used to assess a student’s understanding of a particular topic, to develop critical thinking skills, and to improve writing and research abilities.
- Business : Assignments can be used in the business world to assess employee skills, to evaluate job performance, and to provide training opportunities. They can also be used to develop business plans, marketing strategies, and financial projections.
- Journalism : Assignments are often used in journalism to produce news articles, features, and investigative reports. Journalists may be assigned to cover a particular event or topic, or to research and write a story on a specific subject.
- Research : Assignments can be used in research to collect and analyze data, to conduct experiments, and to present findings in written or oral form. Researchers may be assigned to conduct research on a specific topic, to write a research paper, or to present their findings at a conference or seminar.
- Government : Assignments can be used in government to develop policy proposals, to conduct research, and to analyze data. Government officials may be assigned to work on a specific project or to conduct research on a particular topic.
- Non-profit organizations: Assignments can be used in non-profit organizations to develop fundraising strategies, to plan events, and to conduct research. Volunteers may be assigned to work on a specific project or to help with a particular task.
Purpose of Assignment
The purpose of an assignment varies depending on the context in which it is given. However, some common purposes of assignments include:
- Assessing learning: Assignments are often used to assess a student’s understanding of a particular topic or concept. This allows educators to determine if a student has mastered the material or if they need additional support.
- Developing skills: Assignments can be used to develop a wide range of skills, such as critical thinking, problem-solving, research, and communication. Assignments that require students to analyze and synthesize information can help to build these skills.
- Encouraging creativity: Assignments can be designed to encourage students to be creative and think outside the box. This can help to foster innovation and original thinking.
- Providing feedback : Assignments provide an opportunity for teachers to provide feedback to students on their progress and performance. Feedback can help students to understand where they need to improve and to develop a growth mindset.
- Meeting learning objectives : Assignments can be designed to help students meet specific learning objectives or outcomes. For example, a writing assignment may be designed to help students improve their writing skills, while a research assignment may be designed to help students develop their research skills.
When to write Assignment
Assignments are typically given by instructors or professors as part of a course or academic program. The timing of when to write an assignment will depend on the specific requirements of the course or program, but in general, assignments should be completed within the timeframe specified by the instructor or program guidelines.
It is important to begin working on assignments as soon as possible to ensure enough time for research, writing, and revisions. Waiting until the last minute can result in rushed work and lower quality output.
It is also important to prioritize assignments based on their due dates and the amount of work required. This will help to manage time effectively and ensure that all assignments are completed on time.
In addition to assignments given by instructors or professors, there may be other situations where writing an assignment is necessary. For example, in the workplace, assignments may be given to complete a specific project or task. In these situations, it is important to establish clear deadlines and expectations to ensure that the assignment is completed on time and to a high standard.
Characteristics of Assignment
Here are some common characteristics of assignments:
- Purpose : Assignments have a specific purpose, such as assessing knowledge or developing skills. They are designed to help students learn and achieve specific learning objectives.
- Requirements: Assignments have specific requirements that must be met, such as a word count, format, or specific content. These requirements are usually provided by the instructor or professor.
- Deadline: Assignments have a specific deadline for completion, which is usually set by the instructor or professor. It is important to meet the deadline to avoid penalties or lower grades.
- Individual or group work: Assignments can be completed individually or as part of a group. Group assignments may require collaboration and communication with other group members.
- Feedback : Assignments provide an opportunity for feedback from the instructor or professor. This feedback can help students to identify areas of improvement and to develop their skills.
- Academic integrity: Assignments require academic integrity, which means that students must submit original work and avoid plagiarism. This includes citing sources properly and following ethical guidelines.
- Learning outcomes : Assignments are designed to help students achieve specific learning outcomes. These outcomes are usually related to the course objectives and may include developing critical thinking skills, writing abilities, or subject-specific knowledge.
Advantages of Assignment
There are several advantages of assignment, including:
- Helps in learning: Assignments help students to reinforce their learning and understanding of a particular topic. By completing assignments, students get to apply the concepts learned in class, which helps them to better understand and retain the information.
- Develops critical thinking skills: Assignments often require students to think critically and analyze information in order to come up with a solution or answer. This helps to develop their critical thinking skills, which are important for success in many areas of life.
- Encourages creativity: Assignments that require students to create something, such as a piece of writing or a project, can encourage creativity and innovation. This can help students to develop new ideas and perspectives, which can be beneficial in many areas of life.
- Builds time-management skills: Assignments often come with deadlines, which can help students to develop time-management skills. Learning how to manage time effectively is an important skill that can help students to succeed in many areas of life.
- Provides feedback: Assignments provide an opportunity for students to receive feedback on their work. This feedback can help students to identify areas where they need to improve and can help them to grow and develop.
Limitations of Assignment
There are also some limitations of assignments that should be considered, including:
- Limited scope: Assignments are often limited in scope, and may not provide a comprehensive understanding of a particular topic. They may only cover a specific aspect of a topic, and may not provide a full picture of the subject matter.
- Lack of engagement: Some assignments may not engage students in the learning process, particularly if they are repetitive or not challenging enough. This can lead to a lack of motivation and interest in the subject matter.
- Time-consuming: Assignments can be time-consuming, particularly if they require a lot of research or writing. This can be a disadvantage for students who have other commitments, such as work or extracurricular activities.
- Unreliable assessment: The assessment of assignments can be subjective and may not always accurately reflect a student’s understanding or abilities. The grading may be influenced by factors such as the instructor’s personal biases or the student’s writing style.
- Lack of feedback : Although assignments can provide feedback, this feedback may not always be detailed or useful. Instructors may not have the time or resources to provide detailed feedback on every assignment, which can limit the value of the feedback that students receive.
About the author
Muhammad Hassan
Researcher, Academic Writer, Web developer
You may also like

References in Research – Types, Examples and...

APA Table of Contents – Format and Example

Future Research – Thesis Guide

Table of Contents – Types, Formats, Examples

Research Gap – Types, Examples and How to...

Research Problem – Examples, Types and Guide

Understanding Assignments
What this handout is about.
The first step in any successful college writing venture is reading the assignment. While this sounds like a simple task, it can be a tough one. This handout will help you unravel your assignment and begin to craft an effective response. Much of the following advice will involve translating typical assignment terms and practices into meaningful clues to the type of writing your instructor expects. See our short video for more tips.
Basic beginnings
Regardless of the assignment, department, or instructor, adopting these two habits will serve you well :
- Read the assignment carefully as soon as you receive it. Do not put this task off—reading the assignment at the beginning will save you time, stress, and problems later. An assignment can look pretty straightforward at first, particularly if the instructor has provided lots of information. That does not mean it will not take time and effort to complete; you may even have to learn a new skill to complete the assignment.
- Ask the instructor about anything you do not understand. Do not hesitate to approach your instructor. Instructors would prefer to set you straight before you hand the paper in. That’s also when you will find their feedback most useful.
Assignment formats
Many assignments follow a basic format. Assignments often begin with an overview of the topic, include a central verb or verbs that describe the task, and offer some additional suggestions, questions, or prompts to get you started.
An Overview of Some Kind
The instructor might set the stage with some general discussion of the subject of the assignment, introduce the topic, or remind you of something pertinent that you have discussed in class. For example:
“Throughout history, gerbils have played a key role in politics,” or “In the last few weeks of class, we have focused on the evening wear of the housefly …”
The Task of the Assignment
Pay attention; this part tells you what to do when you write the paper. Look for the key verb or verbs in the sentence. Words like analyze, summarize, or compare direct you to think about your topic in a certain way. Also pay attention to words such as how, what, when, where, and why; these words guide your attention toward specific information. (See the section in this handout titled “Key Terms” for more information.)
“Analyze the effect that gerbils had on the Russian Revolution”, or “Suggest an interpretation of housefly undergarments that differs from Darwin’s.”
Additional Material to Think about
Here you will find some questions to use as springboards as you begin to think about the topic. Instructors usually include these questions as suggestions rather than requirements. Do not feel compelled to answer every question unless the instructor asks you to do so. Pay attention to the order of the questions. Sometimes they suggest the thinking process your instructor imagines you will need to follow to begin thinking about the topic.
“You may wish to consider the differing views held by Communist gerbils vs. Monarchist gerbils, or Can there be such a thing as ‘the housefly garment industry’ or is it just a home-based craft?”
These are the instructor’s comments about writing expectations:
“Be concise”, “Write effectively”, or “Argue furiously.”
Technical Details
These instructions usually indicate format rules or guidelines.
“Your paper must be typed in Palatino font on gray paper and must not exceed 600 pages. It is due on the anniversary of Mao Tse-tung’s death.”
The assignment’s parts may not appear in exactly this order, and each part may be very long or really short. Nonetheless, being aware of this standard pattern can help you understand what your instructor wants you to do.
Interpreting the assignment
Ask yourself a few basic questions as you read and jot down the answers on the assignment sheet:
Why did your instructor ask you to do this particular task?
Who is your audience.
- What kind of evidence do you need to support your ideas?
What kind of writing style is acceptable?
- What are the absolute rules of the paper?
Try to look at the question from the point of view of the instructor. Recognize that your instructor has a reason for giving you this assignment and for giving it to you at a particular point in the semester. In every assignment, the instructor has a challenge for you. This challenge could be anything from demonstrating an ability to think clearly to demonstrating an ability to use the library. See the assignment not as a vague suggestion of what to do but as an opportunity to show that you can handle the course material as directed. Paper assignments give you more than a topic to discuss—they ask you to do something with the topic. Keep reminding yourself of that. Be careful to avoid the other extreme as well: do not read more into the assignment than what is there.
Of course, your instructor has given you an assignment so that they will be able to assess your understanding of the course material and give you an appropriate grade. But there is more to it than that. Your instructor has tried to design a learning experience of some kind. Your instructor wants you to think about something in a particular way for a particular reason. If you read the course description at the beginning of your syllabus, review the assigned readings, and consider the assignment itself, you may begin to see the plan, purpose, or approach to the subject matter that your instructor has created for you. If you still aren’t sure of the assignment’s goals, try asking the instructor. For help with this, see our handout on getting feedback .
Given your instructor’s efforts, it helps to answer the question: What is my purpose in completing this assignment? Is it to gather research from a variety of outside sources and present a coherent picture? Is it to take material I have been learning in class and apply it to a new situation? Is it to prove a point one way or another? Key words from the assignment can help you figure this out. Look for key terms in the form of active verbs that tell you what to do.
Key Terms: Finding Those Active Verbs
Here are some common key words and definitions to help you think about assignment terms:
Information words Ask you to demonstrate what you know about the subject, such as who, what, when, where, how, and why.
- define —give the subject’s meaning (according to someone or something). Sometimes you have to give more than one view on the subject’s meaning
- describe —provide details about the subject by answering question words (such as who, what, when, where, how, and why); you might also give details related to the five senses (what you see, hear, feel, taste, and smell)
- explain —give reasons why or examples of how something happened
- illustrate —give descriptive examples of the subject and show how each is connected with the subject
- summarize —briefly list the important ideas you learned about the subject
- trace —outline how something has changed or developed from an earlier time to its current form
- research —gather material from outside sources about the subject, often with the implication or requirement that you will analyze what you have found
Relation words Ask you to demonstrate how things are connected.
- compare —show how two or more things are similar (and, sometimes, different)
- contrast —show how two or more things are dissimilar
- apply—use details that you’ve been given to demonstrate how an idea, theory, or concept works in a particular situation
- cause —show how one event or series of events made something else happen
- relate —show or describe the connections between things
Interpretation words Ask you to defend ideas of your own about the subject. Do not see these words as requesting opinion alone (unless the assignment specifically says so), but as requiring opinion that is supported by concrete evidence. Remember examples, principles, definitions, or concepts from class or research and use them in your interpretation.
- assess —summarize your opinion of the subject and measure it against something
- prove, justify —give reasons or examples to demonstrate how or why something is the truth
- evaluate, respond —state your opinion of the subject as good, bad, or some combination of the two, with examples and reasons
- support —give reasons or evidence for something you believe (be sure to state clearly what it is that you believe)
- synthesize —put two or more things together that have not been put together in class or in your readings before; do not just summarize one and then the other and say that they are similar or different—you must provide a reason for putting them together that runs all the way through the paper
- analyze —determine how individual parts create or relate to the whole, figure out how something works, what it might mean, or why it is important
- argue —take a side and defend it with evidence against the other side
More Clues to Your Purpose As you read the assignment, think about what the teacher does in class:
- What kinds of textbooks or coursepack did your instructor choose for the course—ones that provide background information, explain theories or perspectives, or argue a point of view?
- In lecture, does your instructor ask your opinion, try to prove their point of view, or use keywords that show up again in the assignment?
- What kinds of assignments are typical in this discipline? Social science classes often expect more research. Humanities classes thrive on interpretation and analysis.
- How do the assignments, readings, and lectures work together in the course? Instructors spend time designing courses, sometimes even arguing with their peers about the most effective course materials. Figuring out the overall design to the course will help you understand what each assignment is meant to achieve.
Now, what about your reader? Most undergraduates think of their audience as the instructor. True, your instructor is a good person to keep in mind as you write. But for the purposes of a good paper, think of your audience as someone like your roommate: smart enough to understand a clear, logical argument, but not someone who already knows exactly what is going on in your particular paper. Remember, even if the instructor knows everything there is to know about your paper topic, they still have to read your paper and assess your understanding. In other words, teach the material to your reader.
Aiming a paper at your audience happens in two ways: you make decisions about the tone and the level of information you want to convey.
- Tone means the “voice” of your paper. Should you be chatty, formal, or objective? Usually you will find some happy medium—you do not want to alienate your reader by sounding condescending or superior, but you do not want to, um, like, totally wig on the man, you know? Eschew ostentatious erudition: some students think the way to sound academic is to use big words. Be careful—you can sound ridiculous, especially if you use the wrong big words.
- The level of information you use depends on who you think your audience is. If you imagine your audience as your instructor and they already know everything you have to say, you may find yourself leaving out key information that can cause your argument to be unconvincing and illogical. But you do not have to explain every single word or issue. If you are telling your roommate what happened on your favorite science fiction TV show last night, you do not say, “First a dark-haired white man of average height, wearing a suit and carrying a flashlight, walked into the room. Then a purple alien with fifteen arms and at least three eyes turned around. Then the man smiled slightly. In the background, you could hear a clock ticking. The room was fairly dark and had at least two windows that I saw.” You also do not say, “This guy found some aliens. The end.” Find some balance of useful details that support your main point.
You’ll find a much more detailed discussion of these concepts in our handout on audience .
The Grim Truth
With a few exceptions (including some lab and ethnography reports), you are probably being asked to make an argument. You must convince your audience. It is easy to forget this aim when you are researching and writing; as you become involved in your subject matter, you may become enmeshed in the details and focus on learning or simply telling the information you have found. You need to do more than just repeat what you have read. Your writing should have a point, and you should be able to say it in a sentence. Sometimes instructors call this sentence a “thesis” or a “claim.”
So, if your instructor tells you to write about some aspect of oral hygiene, you do not want to just list: “First, you brush your teeth with a soft brush and some peanut butter. Then, you floss with unwaxed, bologna-flavored string. Finally, gargle with bourbon.” Instead, you could say, “Of all the oral cleaning methods, sandblasting removes the most plaque. Therefore it should be recommended by the American Dental Association.” Or, “From an aesthetic perspective, moldy teeth can be quite charming. However, their joys are short-lived.”
Convincing the reader of your argument is the goal of academic writing. It doesn’t have to say “argument” anywhere in the assignment for you to need one. Look at the assignment and think about what kind of argument you could make about it instead of just seeing it as a checklist of information you have to present. For help with understanding the role of argument in academic writing, see our handout on argument .
What kind of evidence do you need?
There are many kinds of evidence, and what type of evidence will work for your assignment can depend on several factors–the discipline, the parameters of the assignment, and your instructor’s preference. Should you use statistics? Historical examples? Do you need to conduct your own experiment? Can you rely on personal experience? See our handout on evidence for suggestions on how to use evidence appropriately.
Make sure you are clear about this part of the assignment, because your use of evidence will be crucial in writing a successful paper. You are not just learning how to argue; you are learning how to argue with specific types of materials and ideas. Ask your instructor what counts as acceptable evidence. You can also ask a librarian for help. No matter what kind of evidence you use, be sure to cite it correctly—see the UNC Libraries citation tutorial .
You cannot always tell from the assignment just what sort of writing style your instructor expects. The instructor may be really laid back in class but still expect you to sound formal in writing. Or the instructor may be fairly formal in class and ask you to write a reflection paper where you need to use “I” and speak from your own experience.
Try to avoid false associations of a particular field with a style (“art historians like wacky creativity,” or “political scientists are boring and just give facts”) and look instead to the types of readings you have been given in class. No one expects you to write like Plato—just use the readings as a guide for what is standard or preferable to your instructor. When in doubt, ask your instructor about the level of formality they expect.
No matter what field you are writing for or what facts you are including, if you do not write so that your reader can understand your main idea, you have wasted your time. So make clarity your main goal. For specific help with style, see our handout on style .
Technical details about the assignment
The technical information you are given in an assignment always seems like the easy part. This section can actually give you lots of little hints about approaching the task. Find out if elements such as page length and citation format (see the UNC Libraries citation tutorial ) are negotiable. Some professors do not have strong preferences as long as you are consistent and fully answer the assignment. Some professors are very specific and will deduct big points for deviations.
Usually, the page length tells you something important: The instructor thinks the size of the paper is appropriate to the assignment’s parameters. In plain English, your instructor is telling you how many pages it should take for you to answer the question as fully as you are expected to. So if an assignment is two pages long, you cannot pad your paper with examples or reword your main idea several times. Hit your one point early, defend it with the clearest example, and finish quickly. If an assignment is ten pages long, you can be more complex in your main points and examples—and if you can only produce five pages for that assignment, you need to see someone for help—as soon as possible.
Tricks that don’t work
Your instructors are not fooled when you:
- spend more time on the cover page than the essay —graphics, cool binders, and cute titles are no replacement for a well-written paper.
- use huge fonts, wide margins, or extra spacing to pad the page length —these tricks are immediately obvious to the eye. Most instructors use the same word processor you do. They know what’s possible. Such tactics are especially damning when the instructor has a stack of 60 papers to grade and yours is the only one that low-flying airplane pilots could read.
- use a paper from another class that covered “sort of similar” material . Again, the instructor has a particular task for you to fulfill in the assignment that usually relates to course material and lectures. Your other paper may not cover this material, and turning in the same paper for more than one course may constitute an Honor Code violation . Ask the instructor—it can’t hurt.
- get all wacky and “creative” before you answer the question . Showing that you are able to think beyond the boundaries of a simple assignment can be good, but you must do what the assignment calls for first. Again, check with your instructor. A humorous tone can be refreshing for someone grading a stack of papers, but it will not get you a good grade if you have not fulfilled the task.
Critical reading of assignments leads to skills in other types of reading and writing. If you get good at figuring out what the real goals of assignments are, you are going to be better at understanding the goals of all of your classes and fields of study.
You may reproduce it for non-commercial use if you use the entire handout and attribute the source: The Writing Center, University of North Carolina at Chapel Hill
Make a Gift
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, generate accurate citations for free.
- Knowledge Base
The Beginner's Guide to Writing an Essay | Steps & Examples
An academic essay is a focused piece of writing that develops an idea or argument using evidence, analysis, and interpretation.
There are many types of essays you might write as a student. The content and length of an essay depends on your level, subject of study, and course requirements. However, most essays at university level are argumentative — they aim to persuade the reader of a particular position or perspective on a topic.
The essay writing process consists of three main stages:
- Preparation: Decide on your topic, do your research, and create an essay outline.
- Writing : Set out your argument in the introduction, develop it with evidence in the main body, and wrap it up with a conclusion.
- Revision: Check your essay on the content, organization, grammar, spelling, and formatting of your essay.
Instantly correct all language mistakes in your text
Upload your document to correct all your mistakes in minutes
Table of contents
Essay writing process, preparation for writing an essay, writing the introduction, writing the main body, writing the conclusion, essay checklist, lecture slides, frequently asked questions about writing an essay.
The writing process of preparation, writing, and revisions applies to every essay or paper, but the time and effort spent on each stage depends on the type of essay .
For example, if you’ve been assigned a five-paragraph expository essay for a high school class, you’ll probably spend the most time on the writing stage; for a college-level argumentative essay , on the other hand, you’ll need to spend more time researching your topic and developing an original argument before you start writing.
| 1. Preparation | 2. Writing | 3. Revision |
|---|---|---|
| , organized into Write the | or use a for language errors |
Receive feedback on language, structure, and formatting
Professional editors proofread and edit your paper by focusing on:
- Academic style
- Vague sentences
- Style consistency
See an example
Before you start writing, you should make sure you have a clear idea of what you want to say and how you’re going to say it. There are a few key steps you can follow to make sure you’re prepared:
- Understand your assignment: What is the goal of this essay? What is the length and deadline of the assignment? Is there anything you need to clarify with your teacher or professor?
- Define a topic: If you’re allowed to choose your own topic , try to pick something that you already know a bit about and that will hold your interest.
- Do your research: Read primary and secondary sources and take notes to help you work out your position and angle on the topic. You’ll use these as evidence for your points.
- Come up with a thesis: The thesis is the central point or argument that you want to make. A clear thesis is essential for a focused essay—you should keep referring back to it as you write.
- Create an outline: Map out the rough structure of your essay in an outline . This makes it easier to start writing and keeps you on track as you go.
Once you’ve got a clear idea of what you want to discuss, in what order, and what evidence you’ll use, you’re ready to start writing.
The introduction sets the tone for your essay. It should grab the reader’s interest and inform them of what to expect. The introduction generally comprises 10–20% of the text.
1. Hook your reader
The first sentence of the introduction should pique your reader’s interest and curiosity. This sentence is sometimes called the hook. It might be an intriguing question, a surprising fact, or a bold statement emphasizing the relevance of the topic.
Let’s say we’re writing an essay about the development of Braille (the raised-dot reading and writing system used by visually impaired people). Our hook can make a strong statement about the topic:
The invention of Braille was a major turning point in the history of disability.
2. Provide background on your topic
Next, it’s important to give context that will help your reader understand your argument. This might involve providing background information, giving an overview of important academic work or debates on the topic, and explaining difficult terms. Don’t provide too much detail in the introduction—you can elaborate in the body of your essay.
3. Present the thesis statement
Next, you should formulate your thesis statement— the central argument you’re going to make. The thesis statement provides focus and signals your position on the topic. It is usually one or two sentences long. The thesis statement for our essay on Braille could look like this:
As the first writing system designed for blind people’s needs, Braille was a groundbreaking new accessibility tool. It not only provided practical benefits, but also helped change the cultural status of blindness.
4. Map the structure
In longer essays, you can end the introduction by briefly describing what will be covered in each part of the essay. This guides the reader through your structure and gives a preview of how your argument will develop.
The invention of Braille marked a major turning point in the history of disability. The writing system of raised dots used by blind and visually impaired people was developed by Louis Braille in nineteenth-century France. In a society that did not value disabled people in general, blindness was particularly stigmatized, and lack of access to reading and writing was a significant barrier to social participation. The idea of tactile reading was not entirely new, but existing methods based on sighted systems were difficult to learn and use. As the first writing system designed for blind people’s needs, Braille was a groundbreaking new accessibility tool. It not only provided practical benefits, but also helped change the cultural status of blindness. This essay begins by discussing the situation of blind people in nineteenth-century Europe. It then describes the invention of Braille and the gradual process of its acceptance within blind education. Subsequently, it explores the wide-ranging effects of this invention on blind people’s social and cultural lives.
Write your essay introduction
The body of your essay is where you make arguments supporting your thesis, provide evidence, and develop your ideas. Its purpose is to present, interpret, and analyze the information and sources you have gathered to support your argument.
Length of the body text
The length of the body depends on the type of essay. On average, the body comprises 60–80% of your essay. For a high school essay, this could be just three paragraphs, but for a graduate school essay of 6,000 words, the body could take up 8–10 pages.
Paragraph structure
To give your essay a clear structure , it is important to organize it into paragraphs . Each paragraph should be centered around one main point or idea.
That idea is introduced in a topic sentence . The topic sentence should generally lead on from the previous paragraph and introduce the point to be made in this paragraph. Transition words can be used to create clear connections between sentences.
After the topic sentence, present evidence such as data, examples, or quotes from relevant sources. Be sure to interpret and explain the evidence, and show how it helps develop your overall argument.
Lack of access to reading and writing put blind people at a serious disadvantage in nineteenth-century society. Text was one of the primary methods through which people engaged with culture, communicated with others, and accessed information; without a well-developed reading system that did not rely on sight, blind people were excluded from social participation (Weygand, 2009). While disabled people in general suffered from discrimination, blindness was widely viewed as the worst disability, and it was commonly believed that blind people were incapable of pursuing a profession or improving themselves through culture (Weygand, 2009). This demonstrates the importance of reading and writing to social status at the time: without access to text, it was considered impossible to fully participate in society. Blind people were excluded from the sighted world, but also entirely dependent on sighted people for information and education.
See the full essay example
Here's why students love Scribbr's proofreading services
Discover proofreading & editing
The conclusion is the final paragraph of an essay. It should generally take up no more than 10–15% of the text . A strong essay conclusion :
- Returns to your thesis
- Ties together your main points
- Shows why your argument matters
A great conclusion should finish with a memorable or impactful sentence that leaves the reader with a strong final impression.
What not to include in a conclusion
To make your essay’s conclusion as strong as possible, there are a few things you should avoid. The most common mistakes are:
- Including new arguments or evidence
- Undermining your arguments (e.g. “This is just one approach of many”)
- Using concluding phrases like “To sum up…” or “In conclusion…”
Braille paved the way for dramatic cultural changes in the way blind people were treated and the opportunities available to them. Louis Braille’s innovation was to reimagine existing reading systems from a blind perspective, and the success of this invention required sighted teachers to adapt to their students’ reality instead of the other way around. In this sense, Braille helped drive broader social changes in the status of blindness. New accessibility tools provide practical advantages to those who need them, but they can also change the perspectives and attitudes of those who do not.
Write your essay conclusion
Checklist: Essay
My essay follows the requirements of the assignment (topic and length ).
My introduction sparks the reader’s interest and provides any necessary background information on the topic.
My introduction contains a thesis statement that states the focus and position of the essay.
I use paragraphs to structure the essay.
I use topic sentences to introduce each paragraph.
Each paragraph has a single focus and a clear connection to the thesis statement.
I make clear transitions between paragraphs and ideas.
My conclusion doesn’t just repeat my points, but draws connections between arguments.
I don’t introduce new arguments or evidence in the conclusion.
I have given an in-text citation for every quote or piece of information I got from another source.
I have included a reference page at the end of my essay, listing full details of all my sources.
My citations and references are correctly formatted according to the required citation style .
My essay has an interesting and informative title.
I have followed all formatting guidelines (e.g. font, page numbers, line spacing).
Your essay meets all the most important requirements. Our editors can give it a final check to help you submit with confidence.
Open Google Slides Download PowerPoint
An essay is a focused piece of writing that explains, argues, describes, or narrates.
In high school, you may have to write many different types of essays to develop your writing skills.
Academic essays at college level are usually argumentative : you develop a clear thesis about your topic and make a case for your position using evidence, analysis and interpretation.
The structure of an essay is divided into an introduction that presents your topic and thesis statement , a body containing your in-depth analysis and arguments, and a conclusion wrapping up your ideas.
The structure of the body is flexible, but you should always spend some time thinking about how you can organize your essay to best serve your ideas.
Your essay introduction should include three main things, in this order:
- An opening hook to catch the reader’s attention.
- Relevant background information that the reader needs to know.
- A thesis statement that presents your main point or argument.
The length of each part depends on the length and complexity of your essay .
A thesis statement is a sentence that sums up the central point of your paper or essay . Everything else you write should relate to this key idea.
The thesis statement is essential in any academic essay or research paper for two main reasons:
- It gives your writing direction and focus.
- It gives the reader a concise summary of your main point.
Without a clear thesis statement, an essay can end up rambling and unfocused, leaving your reader unsure of exactly what you want to say.
A topic sentence is a sentence that expresses the main point of a paragraph . Everything else in the paragraph should relate to the topic sentence.
At college level, you must properly cite your sources in all essays , research papers , and other academic texts (except exams and in-class exercises).
Add a citation whenever you quote , paraphrase , or summarize information or ideas from a source. You should also give full source details in a bibliography or reference list at the end of your text.
The exact format of your citations depends on which citation style you are instructed to use. The most common styles are APA , MLA , and Chicago .
Is this article helpful?
Other students also liked.
- How long is an essay? Guidelines for different types of essay
- How to write an essay introduction | 4 steps & examples
- How to conclude an essay | Interactive example
More interesting articles
- Checklist for academic essays | Is your essay ready to submit?
- Comparing and contrasting in an essay | Tips & examples
- Example of a great essay | Explanations, tips & tricks
- Generate topic ideas for an essay or paper | Tips & techniques
- How to revise an essay in 3 simple steps
- How to structure an essay: Templates and tips
- How to write a descriptive essay | Example & tips
- How to write a literary analysis essay | A step-by-step guide
- How to write a narrative essay | Example & tips
- How to write a rhetorical analysis | Key concepts & examples
- How to Write a Thesis Statement | 4 Steps & Examples
- How to write an argumentative essay | Examples & tips
- How to write an essay outline | Guidelines & examples
- How to write an expository essay
- How to write the body of an essay | Drafting & redrafting
- Kinds of argumentative academic essays and their purposes
- Organizational tips for academic essays
- The four main types of essay | Quick guide with examples
- Transition sentences | Tips & examples for clear writing
Get unlimited documents corrected
✔ Free APA citation check included ✔ Unlimited document corrections ✔ Specialized in correcting academic texts
How to Write a Perfect Assignment: Step-By-Step Guide
Table of contents
- 1 How to Structure an Assignment?
- 2.1 The research part
- 2.2 Planning your text
- 2.3 Writing major parts
- 3 Expert Tips for your Writing Assignment
- 4 Will I succeed with my assignments?
- 5 Conclusion
How to Structure an Assignment?
To cope with assignments, you should familiarize yourself with the tips on formatting and presenting assignments or any written paper, which are given below. It is worth paying attention to the content of the paper, making it structured and understandable so that ideas are not lost and thoughts do not refute each other.
If the topic is free or you can choose from the given list — be sure to choose the one you understand best. Especially if that could affect your semester score or scholarship. It is important to select an engaging title that is contextualized within your topic. A topic that should captivate you or at least give you a general sense of what is needed there. It’s easier to dwell upon what interests you, so the process goes faster.
To construct an assignment structure, use outlines. These are pieces of text that relate to your topic. It can be ideas, quotes, all your thoughts, or disparate arguments. Type in everything that you think about. Separate thoughts scattered across the sheets of Word will help in the next step.
Then it is time to form the text. At this stage, you have to form a coherent story from separate pieces, where each new thought reinforces the previous one, and one idea smoothly flows into another.
Main Steps of Assignment Writing
These are steps to take to get a worthy paper. If you complete these step-by-step, your text will be among the most exemplary ones.
The research part
If the topic is unique and no one has written about it yet, look at materials close to this topic to gain thoughts about it. You should feel that you are ready to express your thoughts. Also, while reading, get acquainted with the format of the articles, study the details, collect material for your thoughts, and accumulate different points of view for your article. Be careful at this stage, as the process can help you develop your ideas. If you are already struggling here, pay for assignment to be done , and it will be processed in a split second via special services. These services are especially helpful when the deadline is near as they guarantee fast delivery of high-quality papers on any subject.
If you use Google to search for material for your assignment, you will, of course, find a lot of information very quickly. Still, the databases available on your library’s website will give you the clearest and most reliable facts that satisfy your teacher or professor. Be sure you copy the addresses of all the web pages you will use when composing your paper, so you don’t lose them. You can use them later in your bibliography if you add a bit of description! Select resources and extract quotes from them that you can use while working. At this stage, you may also create a request for late assignment if you realize the paper requires a lot of effort and is time-consuming. This way, you’ll have a backup plan if something goes wrong.
Planning your text
Assemble a layout. It may be appropriate to use the structure of the paper of some outstanding scientists in your field and argue it in one of the parts. As the planning progresses, you can add suggestions that come to mind. If you use citations that require footnotes, and if you use single spacing throughout the paper and double spacing at the end, it will take you a very long time to make sure that all the citations are on the exact pages you specified! Add a reference list or bibliography. If you haven’t already done so, don’t put off writing an essay until the last day. It will be more difficult to do later as you will be stressed out because of time pressure.
Writing major parts
It happens that there is simply no mood or strength to get started and zero thoughts. In that case, postpone this process for 2-3 hours, and, perhaps, soon, you will be able to start with renewed vigor. Writing essays is a great (albeit controversial) way to improve your skills. This experience will not be forgotten. It will certainly come in handy and bring many benefits in the future. Do your best here because asking for an extension is not always possible, so you probably won’t have time to redo it later. And the quality of this part defines the success of the whole paper.
Writing the major part does not mean the matter is finished. To review the text, make sure that the ideas of the introduction and conclusion coincide because such a discrepancy is the first thing that will catch the reader’s eye and can spoil the impression. Add or remove anything from your intro to edit it to fit the entire paper. Also, check your spelling and grammar to ensure there are no typos or draft comments. Check the sources of your quotes so that your it is honest and does not violate any rules. And do not forget the formatting rules.
with the right tips and guidance, it can be easier than it looks. To make the process even more straightforward, students can also use an assignment service to get the job done. This way they can get professional assistance and make sure that their assignments are up to the mark. At PapersOwl, we provide a professional writing service where students can order custom-made assignments that meet their exact requirements.
Expert Tips for your Writing Assignment
Want to write like a pro? Here’s what you should consider:
- Save the document! Send the finished document by email to yourself so you have a backup copy in case your computer crashes.
- Don’t wait until the last minute to complete a list of citations or a bibliography after the paper is finished. It will be much longer and more difficult, so add to them as you go.
- If you find a lot of information on the topic of your search, then arrange it in a separate paragraph.
- If possible, choose a topic that you know and are interested in.
- Believe in yourself! If you set yourself up well and use your limited time wisely, you will be able to deliver the paper on time.
- Do not copy information directly from the Internet without citing them.
Writing assignments is a tedious and time-consuming process. It requires a lot of research and hard work to produce a quality paper. However, if you are feeling overwhelmed or having difficulty understanding the concept, you may want to consider getting accounting homework help online . Professional experts can assist you in understanding how to complete your assignment effectively. PapersOwl.com offers expert help from highly qualified and experienced writers who can provide you with the homework help you need.
Will I succeed with my assignments?
Anyone can learn how to be good at writing: follow simple rules of creating the structure and be creative where it is appropriate. At one moment, you will need some additional study tools, study support, or solid study tips. And you can easily get help in writing assignments or any other work. This is especially useful since the strategy of learning how to write an assignment can take more time than a student has.
Therefore all students are happy that there is an option to order your paper at a professional service to pass all the courses perfectly and sleep still at night. You can also find the sample of the assignment there to check if you are on the same page and if not — focus on your papers more diligently.
So, in the times of studies online, the desire and skill to research and write may be lost. Planning your assignment carefully and presenting arguments step-by-step is necessary to succeed with your homework. When going through your references, note the questions that appear and answer them, building your text. Create a cover page, proofread the whole text, and take care of formatting. Feel free to use these rules for passing your next assignments.
When it comes to writing an assignment, it can be overwhelming and stressful, but Papersowl is here to make it easier for you. With a range of helpful resources available, Papersowl can assist you in creating high-quality written work, regardless of whether you’re starting from scratch or refining an existing draft. From conducting research to creating an outline, and from proofreading to formatting, the team at Papersowl has the expertise to guide you through the entire writing process and ensure that your assignment meets all the necessary requirements.
Readers also enjoyed

WHY WAIT? PLACE AN ORDER RIGHT NOW!
Just fill out the form, press the button, and have no worries!
We use cookies to give you the best experience possible. By continuing we’ll assume you board with our cookie policy.
Essay samples for every taste and need
Find the perfect essay sample that you can reference for educational purposes. Need a unique one?

Choose samples by essay type
The font type, compare and contrast, how to craft a good essay writing sample.
Additionally, it's important to understand the essay prompt and requirements before starting to write. Brainstorming, creating an outline, and conducting research are also crucial steps. Once these steps are completed, you can begin writing your essay by following the outline and incorporating your research. Finally, proofread and edit your essay to ensure it is error-free and flows well.If you need more guidance, you can always seek the help of a professional writer from a custom essay writing service. Alternatively, you can find inspiration and guidance by looking for high school sample essays that cover a similar topic to the one you're working on. These can provide you with ideas on how to structure your essay, what to include, and how to approach your argument. So, if you're struggling to write an essay , don't hesitate to find a great sample essay or ask a professional writer to write an essay for you!
- Choose both an essay type and its format
- Brainstorm to find a topic
- Develop a thesis statement
- Make an essay outline
- Write the first draft
- Expand on the drafts
- Edit and proofread
Regular new academic essay sample
William shakespear romeo and juliet, how to stop cyber bullying essay, what rights should animals have, essay about police brutality, overpopulation as a global environmental crisis: concept and prevention, teenage pregnancy prevention programs, how to use our essay samples.
Think of the essays on this website as inspirational research. They are not meant to be presented as your own work. Any sample of essay writing found here is intended to be a tool to help students further understand the topic and develop their own ideas. Copying sections of an essay sample for college would be considered plagiarism and would violate the honor code and may result in disciplinary action. Please think of the material as an additional study aid, but we strongly recommend that students develop their own ideas and write their own words.
Writing is challenging, so use our samples as a basis for new viewpoints, how to structure your essay, and as a source of inspiration.

Frequently asked questions
Where to find a free essay sample online, where to find good sample essay papers to ignite your inspiration, how do i properly cite my sources, is it legal, how to avoid accidentally plagiarizing when i paraphrase a quote, why are your samples better, examples as study materials.
Essay samples have a proven track record of improving a students academic performance. Many high-schools and higher education institutes give essay examples as study materials, especially to explore a topic more deeply. Many academic institutions don't have access to a diverse range of high quality essay samples though, and that's where Studyfy steps in! Our samples cover a huge variety of topics and will be invaluable in helping you to study.
Always here to help
Having trouble with finding the right sample for your topic? Don't worry! The first thing you should do is reach out to a member of our support team and ask them if they can source the right sample. If they can't, then you can always order an essay sample from one of our professional writers. We are committed to helping students with the assignments that they find challenging!
No more struggle
Over the course of your higher education, you will be asked to write papers on a wide variety of topics. While certain topics might be of interest to you, others won't be. It's often these papers that students struggle with the most. Maybe you're not a fan of classic literature, maybe the dreaded foundation or general education courses have forced you to take a science class. Don't let a single course affect your grades! Essay samples, your creativity, Studyfy homework help, online proofreading and essay editing service, custom essay , admission essay writing service , and " write paper for me " services are the perfect tools to give you inspiration and help you with unpleasant essays!
Free features
- Search entire site
- Search for a course
- Browse study areas
Analytics and Data Science
- Data Science and Innovation
- Postgraduate Research Courses
- Business Research Programs
- Undergraduate Business Programs
- Entrepreneurship
- MBA Programs
- Postgraduate Business Programs
Communication
- Animation Production
- Business Consulting and Technology Implementation
- Digital and Social Media
- Media Arts and Production
- Media Business
- Media Practice and Industry
- Music and Sound Design
- Social and Political Sciences
- Strategic Communication
- Writing and Publishing
- Postgraduate Communication Research Degrees
Design, Architecture and Building
- Architecture
- Built Environment
- DAB Research
- Public Policy and Governance
- Secondary Education
- Education (Learning and Leadership)
- Learning Design
- Postgraduate Education Research Degrees
- Primary Education
Engineering
- Civil and Environmental
- Computer Systems and Software
- Engineering Management
- Mechanical and Mechatronic
- Systems and Operations
- Telecommunications
- Postgraduate Engineering courses
- Undergraduate Engineering courses
- Sport and Exercise
- Palliative Care
- Public Health
- Nursing (Undergraduate)
- Nursing (Postgraduate)
- Health (Postgraduate)
- Research and Honours
- Health Services Management
- Child and Family Health
- Women's and Children's Health
Health (GEM)
- Coursework Degrees
- Clinical Psychology
- Genetic Counselling
- Good Manufacturing Practice
- Physiotherapy
- Speech Pathology
- Research Degrees
Information Technology
- Business Analysis and Information Systems
- Computer Science, Data Analytics/Mining
- Games, Graphics and Multimedia
- IT Management and Leadership
- Networking and Security
- Software Development and Programming
- Systems Design and Analysis
- Web and Cloud Computing
- Postgraduate IT courses
- Postgraduate IT online courses
- Undergraduate Information Technology courses
- International Studies
- Criminology
- International Relations
- Postgraduate International Studies Research Degrees
- Sustainability and Environment
- Practical Legal Training
- Commercial and Business Law
- Juris Doctor
- Legal Studies
- Master of Laws
- Intellectual Property
- Migration Law and Practice
- Overseas Qualified Lawyers
- Postgraduate Law Programs
- Postgraduate Law Research
- Undergraduate Law Programs
- Life Sciences
- Mathematical and Physical Sciences
- Postgraduate Science Programs
- Science Research Programs
- Undergraduate Science Programs
Transdisciplinary Innovation
- Creative Intelligence and Innovation
- Diploma in Innovation
- Postgraduate Research Degree
- Transdisciplinary Learning
Sample written assignments
Look at sample assignments to help you develop and enhance your academic writing skills.
How to use this page
This page features authentic sample assignments that you can view or download to help you develop and enhance your academic writing skills.
PLEASE NOTE: Comments included in these sample written assignments are intended as an educational guide only. Always check with academic staff which referencing convention you should follow. All sample assignments have been submitted using Turnitin® (anti-plagiarism software). Under no circumstances should you copy from these or any other texts.
Annotated bibliography
Annotated Bibliography: Traditional Chinese Medicine (PDF, 103KB)
Essay: Business - "Culture is a Tool Used by Management" (PDF, 496KB)
Essay: Business - "Integrating Business Perspectives - Wicked Problem" (PDF, 660KB)
Essay: Business - "Overconsumption and Sustainability" (PDF, 762KB)
Essay: Business - "Post bureaucracy vs Bureaucracy" (PDF, 609KB)
Essay: Design, Architecture & Building - "Ideas in History - Postmodernism" (PDF, 545KB)
Essay: Design, Architecture & Building - "The Context of Visual Communication Design Research Project" (PDF, 798KB)
Essay: Design, Architecture & Building - "Ideas in History - The Nurses Walk and Postmodernism" (PDF, 558KB)
Essay: Health (Childhood Obesity ) (PDF, 159KB)
Essay: Health (Improving Quality and Safety in Healthcare) (PDF, 277KB)
Essay: Health (Organisational Management in Healthcare) (PDF, 229KB)
UTS HELPS annotated Law essay
(PDF, 250KB)
Essay: Science (Traditional Chinese Medicine) (PDF, 153KB)
Literature review
Literature Review: Education (Critical Pedagogy) (PDF, 165KB)
Reflective writing
Reflective Essay: Business (Simulation Project) (PDF, 119KB)
Reflective Essay: Nursing (Professionalism in Context) (PDF, 134KB)
Report: Business (Management Decisions and Control) (PDF, 244KB)
Report: Education (Digital Storytelling) (PDF, 145KB)
Report: Education (Scholarly Practice) (PDF, 261KB)
Report: Engineering Communication (Flood Mitigation & Water Storage) (PDF, 1MB)
UTS acknowledges the Gadigal people of the Eora Nation, the Boorooberongal people of the Dharug Nation, the Bidiagal people and the Gamaygal people, upon whose ancestral lands our university stands. We would also like to pay respect to the Elders both past and present, acknowledging them as the traditional custodians of knowledge for these lands.

Search form
How to write the best college assignments.
By Lois Weldon
When it comes to writing assignments, it is difficult to find a conceptualized guide with clear and simple tips that are easy to follow. That’s exactly what this guide will provide: few simple tips on how to write great assignments, right when you need them. Some of these points will probably be familiar to you, but there is no harm in being reminded of the most important things before you start writing the assignments, which are usually determining on your credits.
The most important aspects: Outline and Introduction
Preparation is the key to success, especially when it comes to academic assignments. It is recommended to always write an outline before you start writing the actual assignment. The outline should include the main points of discussion, which will keep you focused throughout the work and will make your key points clearly defined. Outlining the assignment will save you a lot of time because it will organize your thoughts and make your literature searches much easier. The outline will also help you to create different sections and divide up the word count between them, which will make the assignment more organized.
The introduction is the next important part you should focus on. This is the part that defines the quality of your assignment in the eyes of the reader. The introduction must include a brief background on the main points of discussion, the purpose of developing such work and clear indications on how the assignment is being organized. Keep this part brief, within one or two paragraphs.
This is an example of including the above mentioned points into the introduction of an assignment that elaborates the topic of obesity reaching proportions:
Background : The twenty first century is characterized by many public health challenges, among which obesity takes a major part. The increasing prevalence of obesity is creating an alarming situation in both developed and developing regions of the world.
Structure and aim : This assignment will elaborate and discuss the specific pattern of obesity epidemic development, as well as its epidemiology. Debt, trade and globalization will also be analyzed as factors that led to escalation of the problem. Moreover, the assignment will discuss the governmental interventions that make efforts to address this issue.
Practical tips on assignment writing
Here are some practical tips that will keep your work focused and effective:
– Critical thinking – Academic writing has to be characterized by critical thinking, not only to provide the work with the needed level, but also because it takes part in the final mark.
– Continuity of ideas – When you get to the middle of assignment, things can get confusing. You have to make sure that the ideas are flowing continuously within and between paragraphs, so the reader will be enabled to follow the argument easily. Dividing the work in different paragraphs is very important for this purpose.
– Usage of ‘you’ and ‘I’ – According to the academic writing standards, the assignments should be written in an impersonal language, which means that the usage of ‘you’ and ‘I’ should be avoided. The only acceptable way of building your arguments is by using opinions and evidence from authoritative sources.
– Referencing – this part of the assignment is extremely important and it takes a big part in the final mark. Make sure to use either Vancouver or Harvard referencing systems, and use the same system in the bibliography and while citing work of other sources within the text.
– Usage of examples – A clear understanding on your assignment’s topic should be provided by comparing different sources and identifying their strengths and weaknesses in an objective manner. This is the part where you should show how the knowledge can be applied into practice.
– Numbering and bullets – Instead of using numbering and bullets, the academic writing style prefers the usage of paragraphs.
– Including figures and tables – The figures and tables are an effective way of conveying information to the reader in a clear manner, without disturbing the word count. Each figure and table should have clear headings and you should make sure to mention their sources in the bibliography.
– Word count – the word count of your assignment mustn’t be far above or far below the required word count. The outline will provide you with help in this aspect, so make sure to plan the work in order to keep it within the boundaries.
The importance of an effective conclusion
The conclusion of your assignment is your ultimate chance to provide powerful arguments that will impress the reader. The conclusion in academic writing is usually expressed through three main parts:
– Stating the context and aim of the assignment
– Summarizing the main points briefly
– Providing final comments with consideration of the future (discussing clear examples of things that can be done in order to improve the situation concerning your topic of discussion).
Normal 0 false false false EN-US X-NONE X-NONE /* Style Definitions */ table.MsoNormalTable {mso-style-name:"Table Normal"; mso-tstyle-rowband-size:0; mso-tstyle-colband-size:0; mso-style-noshow:yes; mso-style-priority:99; mso-style-parent:""; mso-padding-alt:0in 5.4pt 0in 5.4pt; mso-para-margin:0in; mso-para-margin-bottom:.0001pt; mso-pagination:widow-orphan; font-size:11.0pt; font-family:"Calibri","sans-serif"; mso-ascii-font-family:Calibri; mso-ascii-theme-font:minor-latin; mso-hansi-font-family:Calibri; mso-hansi-theme-font:minor-latin;}
Lois Weldon is writer at Uk.bestdissertation.com . Lives happily at London with her husband and lovely daughter. Adores writing tips for students. Passionate about Star Wars and yoga.
7 comments on “How To Write The Best College Assignments”
Extremely useful tip for students wanting to score well on their assignments. I concur with the writer that writing an outline before ACTUALLY starting to write assignments is extremely important. I have observed students who start off quite well but they tend to lose focus in between which causes them to lose marks. So an outline helps them to maintain the theme focused.
Hello Great information…. write assignments
Well elabrated
Thanks for the information. This site has amazing articles. Looking forward to continuing on this site.
This article is certainly going to help student . Well written.
Really good, thanks
Practical tips on assignment writing, the’re fantastic. Thank you!
Leave a comment
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
- Stanford Home
- Maps & Directions
- Search Stanford
- Emergency Info
- Terms of Use
- Non-Discrimination
- Accessibility
© Stanford University , Stanford , California 94305 .

A step-by-step guide for creating and formatting APA Style student papers
The start of the semester is the perfect time to learn how to create and format APA Style student papers. This article walks through the formatting steps needed to create an APA Style student paper, starting with a basic setup that applies to the entire paper (margins, font, line spacing, paragraph alignment and indentation, and page headers). It then covers formatting for the major sections of a student paper: the title page, the text, tables and figures, and the reference list. Finally, it concludes by describing how to organize student papers and ways to improve their quality and presentation.
The guidelines for student paper setup are described and shown using annotated diagrams in the Student Paper Setup Guide (PDF, 3.40MB) and the A Step-by-Step Guide to APA Style Student Papers webinar . Chapter 1 of the Concise Guide to APA Style and Chapter 2 of the Publication Manual of the American Psychological Association describe the elements, format, and organization for student papers. Tables and figures are covered in Chapter 7 of both books. Information on paper format and tables and figures and a full sample student paper are also available on the APA Style website.
Basic setup
The guidelines for basic setup apply to the entire paper. Perform these steps when you first open your document, and then you do not have to worry about them again while writing your paper. Because these are general aspects of paper formatting, they apply to all APA Style papers, student or professional. Students should always check with their assigning instructor or institution for specific guidelines for their papers, which may be different than or in addition to APA Style guidelines.
Seventh edition APA Style was designed with modern word-processing programs in mind. Most default settings in programs such as Academic Writer, Microsoft Word, and Google Docs already comply with APA Style. This means that, for most paper elements, you do not have to make any changes to the default settings of your word-processing program. However, you may need to make a few adjustments before you begin writing.
Use 1-in. margins on all sides of the page (top, bottom, left, and right). This is usually how papers are automatically set.
Use a legible font. The default font of your word-processing program is acceptable. Many sans serif and serif fonts can be used in APA Style, including 11-point Calibri, 11-point Arial, 12-point Times New Roman, and 11-point Georgia. You can also use other fonts described on the font page of the website.
Line spacing
Double-space the entire paper including the title page, block quotations, and the reference list. This is something you usually must set using the paragraph function of your word-processing program. But once you do, you will not have to change the spacing for the entirety of your paper–just double-space everything. Do not add blank lines before or after headings. Do not add extra spacing between paragraphs. For paper sections with different line spacing, see the line spacing page.
Paragraph alignment and indentation
Align all paragraphs of text in the body of your paper to the left margin. Leave the right margin ragged. Do not use full justification. Indent the first line of every paragraph of text 0.5-in. using the tab key or the paragraph-formatting function of your word-processing program. For paper sections with different alignment and indentation, see the paragraph alignment and indentation page.
Page numbers
Put a page number in the top right of every page header , including the title page, starting with page number 1. Use the automatic page-numbering function of your word-processing program to insert the page number in the top right corner; do not type the page numbers manually. The page number is the same font and font size as the text of your paper. Student papers do not require a running head on any page, unless specifically requested by the instructor.
Title page setup
Title page elements.
APA Style has two title page formats: student and professional (for details, see title page setup ). Unless instructed otherwise, students should use the student title page format and include the following elements, in the order listed, on the title page:
- Paper title.
- Name of each author (also known as the byline).
- Affiliation for each author.
- Course number and name.
- Instructor name.
- Assignment due date.
- Page number 1 in the top right corner of the page header.
The format for the byline depends on whether the paper has one author, two authors, or three or more authors.
- When the paper has one author, write the name on its own line (e.g., Jasmine C. Hernandez).
- When the paper has two authors, write the names on the same line and separate them with the word “and” (e.g., Upton J. Wang and Natalia Dominguez).
- When the paper has three or more authors, separate the names with commas and include “and” before the final author’s name (e.g., Malia Mohamed, Jaylen T. Brown, and Nia L. Ball).
Students have an academic affiliation, which identities where they studied when the paper was written. Because students working together on a paper are usually in the same class, they will have one shared affiliation. The affiliation consists of the name of the department and the name of the college or university, separated by a comma (e.g., Department of Psychology, George Mason University). The department is that of the course to which the paper is being submitted, which may be different than the department of the student’s major. Do not include the location unless it is part of the institution’s name.
Write the course number and name and the instructor name as shown on institutional materials (e.g., the syllabus). The course number and name are often separated by a colon (e.g., PST-4510: History and Systems Psychology). Write the assignment due date in the month, date, and year format used in your country (e.g., Sept. 10, 2020).
Title page line spacing
Double-space the whole title page. Place the paper title three or four lines down from the top of the page. Add an extra double-spaced blank like between the paper title and the byline. Then, list the other title page elements on separate lines, without extra lines in between.
Title page alignment
Center all title page elements (except the right-aligned page number in the header).
Title page font
Write the title page using the same font and font size as the rest of your paper. Bold the paper title. Use standard font (i.e., no bold, no italics) for all other title page elements.
Text elements
Repeat the paper title at the top of the first page of text. Begin the paper with an introduction to provide background on the topic, cite related studies, and contextualize the paper. Use descriptive headings to identify other sections as needed (e.g., Method, Results, Discussion for quantitative research papers). Sections and headings vary depending on the paper type and its complexity. Text can include tables and figures, block quotations, headings, and footnotes.
Text line spacing
Double-space all text, including headings and section labels, paragraphs of text, and block quotations.

Text alignment
Center the paper title on the first line of the text. Indent the first line of all paragraphs 0.5-in.
Left-align the text. Leave the right margin ragged.
Block quotation alignment
Indent the whole block quotation 0.5-in. from the left margin. Double-space the block quotation, the same as other body text. Find more information on the quotations page.
Use the same font throughout the entire paper. Write body text in standard (nonbold, nonitalic) font. Bold only headings and section labels. Use italics sparingly, for instance, to highlight a key term on first use (for more information, see the italics page).
Headings format
For detailed guidance on formatting headings, including headings in the introduction of a paper, see the headings page and the headings in sample papers .
- Alignment: Center Level 1 headings. Left-align Level 2 and Level 3 headings. Indent Level 4 and Level 5 headings like a regular paragraph.
- Font: Boldface all headings. Also italicize Level 3 and Level 5 headings. Create heading styles using your word-processing program (built into AcademicWriter, available for Word via the sample papers on the APA Style website).
Tables and figures setup
Tables and figures are only included in student papers if needed for the assignment. Tables and figures share the same elements and layout. See the website for sample tables and sample figures .
Table elements
Tables include the following four elements:
- Body (rows and columns)
- Note (optional if needed to explain elements in the table)
Figure elements
Figures include the following four elements:
- Image (chart, graph, etc.)
- Note (optional if needed to explain elements in the figure)
Table line spacing
Double-space the table number and title. Single-, 1.5-, or double-space the table body (adjust as needed for readability). Double-space the table note.
Figure line spacing
Double-space the figure number and title. The default settings for spacing in figure images is usually acceptable (but adjust the spacing as needed for readability). Double-space the figure note.
Table alignment
Left-align the table number and title. Center column headings. Left-align the table itself and left-align the leftmost (stub) column. Center data in the table body if it is short or left-align the data if it is long. Left-align the table note.
Figure alignment
Left-align the figure number and title. Left-align the whole figure image. The default alignment of the program in which you created your figure is usually acceptable for axis titles and data labels. Left-align the figure note.
Bold the table number. Italicize the table title. Use the same font and font size in the table body as the text of your paper. Italicize the word “Note” at the start of the table note. Write the note in the same font and font size as the text of your paper.
Figure font
Bold the figure number. Italicize the figure title. Use a sans serif font (e.g., Calibri, Arial) in the figure image in a size between 8 to 14 points. Italicize the word “Note” at the start of the figure note. Write the note in the same font and font size as the text of your paper.
Placement of tables and figures
There are two options for the placement of tables and figures in an APA Style paper. The first option is to place all tables and figures on separate pages after the reference list. The second option is to embed each table and figure within the text after its first callout. This guide describes options for the placement of tables and figures embedded in the text. If your instructor requires tables and figures to be placed at the end of the paper, see the table and figure guidelines and the sample professional paper .
Call out (mention) the table or figure in the text before embedding it (e.g., write “see Figure 1” or “Table 1 presents”). You can place the table or figure after the callout either at the bottom of the page, at the top of the next page, or by itself on the next page. Avoid placing tables and figures in the middle of the page.
Embedding at the bottom of the page
Include a callout to the table or figure in the text before that table or figure. Add a blank double-spaced line between the text and the table or figure at the bottom of the page.
Embedding at the top of the page
Include a callout to the table in the text on the previous page before that table or figure. The table or figure then appears at the top of the next page. Add a blank double-spaced line between the end of the table or figure and the text that follows.
Embedding on its own page
Embed long tables or large figures on their own page if needed. The text continues on the next page.
Reference list setup
Reference list elements.
The reference list consists of the “References” section label and the alphabetical list of references. View reference examples on the APA Style website. Consult Chapter 10 in both the Concise Guide and Publication Manual for even more examples.
Reference list line spacing
Start the reference list at the top of a new page after the text. Double-space the entire reference list (both within and between entries).
Reference list alignment
Center the “References” label. Apply a hanging indent of 0.5-in. to all reference list entries. Create the hanging indent using your word-processing program; do not manually hit the enter and tab keys.
Reference list font
Bold the “References” label at the top of the first page of references. Use italics within reference list entries on either the title (e.g., webpages, books, reports) or on the source (e.g., journal articles, edited book chapters).
Final checks
Check page order.
- Start each section on a new page.
- Arrange pages in the following order:
- Title page (page 1).
- Text (starts on page 2).
- Reference list (starts on a new page after the text).
Check headings
- Check that headings accurately reflect the content in each section.
- Start each main section with a Level 1 heading.
- Use Level 2 headings for subsections of the introduction.
- Use the same level of heading for sections of equal importance.
- Avoid having only one subsection within a section (have two or more, or none).
Check assignment instructions
- Remember that instructors’ guidelines supersede APA Style.
- Students should check their assignment guidelines or rubric for specific content to include in their papers and to make sure they are meeting assignment requirements.
Tips for better writing
- Ask for feedback on your paper from a classmate, writing center tutor, or instructor.
- Budget time to implement suggestions.
- Use spell-check and grammar-check to identify potential errors, and then manually check those flagged.
- Proofread the paper by reading it slowly and carefully aloud to yourself.
- Consult your university writing center if you need extra help.
About the author

Undergraduate student resources
More from Swinburne University
- Giving to Swinburne
- Student login
- Staff login
- Arts, Humanities and Social Sciences
- Built Environment and Architecture
- Engineering
- Film and Television
- Games and Animation
- Information Technology
- Media and Communication
- Trades and Apprenticeships
- Study online
- Transition to university from VCE
- Direct entry into university
- Returning to study
- Vocational Education and Training at Swinburne
- Early Entry Program
- University entry requirements
- Transferring to Swinburne
- Recognition of prior learning in the workplace
- Study Abroad in Melbourne
- Study support for indigenous students
- Guaranteed pathways from TAFE
- Short courses
- University certificates
- Pre-apprenticeships
- Apprenticeships
- Associate degrees
- Bachelor degrees
- Double degrees
- Certificates
- Traineeships
- Trade short courses
- Doctor of Philosophy
- Master degrees
- Graduate diploma courses
- Graduate certificate courses
- Studying outside of Australia
- Study on campus
- Loans and discounts for local students
- Fees for international students
- Fees for local students
- Student Services and Amenities Fee
- Scholarship conditions
- Scholarships for international students
- How to apply as a local student
- How to apply for a research degree
- How to apply as an international student
- Apply as an asylum seeker or refugee
- How to enrol
- Understanding your university offer
- Course planner
- Setting up your class timetable
- Enrol as a PhD or master degree student
- Why study in Australia?
- Plan your arrival in Melbourne
- Arriving in Melbourne
- Things to do in Melbourne
- Getting around Melbourne
- Money, living costs and banking in Australia
- International student stories
- Student email, password and Wi-Fi access
- Your student ID card and Swinburne login
- Student discounts and concessions
- Special consideration and extensions
- Accommodation
- Study and learning support
- Health and wellbeing
- Support for international students
- Independent advocacy for service
- Indigenous student services
- Financial support and advice
- AccessAbility services
- Legal advice for students
- Spiritual Wellbeing
- Assault reporting and help
- Asylum seeker and refugee support
- Care leaver support
- LGBTIQ+ community support
- Childcare for the Swinburne community
- Support for Students Guidelines: Policies, Procedures and Frameworks
- Industry-linked projects
- Internships
- Student stories
- Professional Degrees
- Industry study tours
- Get paid to podcast
- Real industry experience stories
- Overseas exchange
- Overseas study tours
- Overseas internships
- Students currently overseas
- Improve your employability
- Career services
- Professional Purpose program
- Partner Stories
- Hosting students with disabilities
- Work with our accreditation placement students
- Benefits of working with our students
- Apprenticeships and traineeships
- Workshops, events and outreach programs
- Work experience
- Knox Innovation, Opportunity and Sustainability Centre
- Australian Synchrotron Science Education
- PrimeSCI! science education
- Student projects
- Meet our facilitators
- Meet our consultants
- Meet our leadership and management teams
- Learning design and innovation
- Hybrid working solutions
- Training needs analysis
- Why partner with Swinburne
- 4 simple steps to setting up a partnership
- Achievements and success stories
- Research engagement
- Facilities and equipment
- Achievements and recognition
- Iverson Health Innovation Research Institute
- Social Innovation Research Institute
- Space Technology and Industry Institute
- Innovative Planet Research Institute
- Research centres, groups and clinics
- Platforms and initiatives
- Indigenous research projects
- Animal research
- Biosafety and Defence
- Data management
- Funding from tobacco companies
- Human research
- Intellectual property
Assignment writing guides and samples
If you're looking for useful guides for assignment writing and language skills check out our range of study skills resources
Essay writing
- Writing essays [PDF 240KB] . Tips on writing a great essay, including developing an argument, structure and appropriate referencing.
- Sample essay [PDF 330KB] . A sample of an essay that includes an annotated structure for your reference.
Writing a critical review
- Writing a critical review [PDF 260KB] . Tips on writing a great critical review, including structure, format and key questions to address when writing a review.
- Sample critical review [PDF 260KB] . A sample of a critical review that includes an annotated structure for your reference.
Writing a business-style report
- Writing a business-style report [PDF 330KB] . A resource for business and law students Find out how to write and format business-style reports.
- Sample of a business-style report [PDF 376 KB] . A resource for business and law students. A sample of a business-style report with an annotated format.
Investigative report sample
- Sample of an investigative report [PDF 500KB] . A resource for science, engineering and technology students. How to write an investigative report, including an annotated format.
Assignment topics and editing
- Interpreting assignment topics [PDF 370 KB] . Find out how to interpret an assignment topic, including understanding key words and concepts.
- How to edit your work [PDF 189KB] . A guide for all students about how to edit and review their work.
Language skills
- Building your word power (expanding your knowledge of words) [PDF 306KB]. A guide to expanding your knowledge of words and communicating your ideas in more interesting ways.
- Handy grammar hints [PDF 217KB] . A guide to getting grammar and style right in your assignments.
Resources relevant to your study area
Science, engineering and technology.
- Writing a critical review [PDF 260KB]. Tips on writing a great critical review, including structure, format and key questions to address when writing a review.
- Sample critical review [PDF 260KB] . A sample of a critical review that includes an annotated structure for your reference.
- Sample of an investigative report [PDF 500KB] . A resource for science, engineering and technology students. How to write an investigative report, including an annotated format.
- How to edit your work [PDF 189KB] . A guide for all students about how to edit and review their work.
- Building your word power (expanding your knowledge of words) [PDF 306KB]. A guide to expanding your knowledge of words and communicating your ideas in more interesting ways.
- Handy grammar hints [PDF 217KB] . A guide to getting grammar and style right in your assignments.
Health, Arts and Design
- Sample essay [PDF 330KB] . A sample of an essay that includes an annotated structure for your reference.
- Writing a critical review [PDF 260KB]. Tips on writing a great critical review, including structure, format and key questions to address when writing a review.
- Sample critical review [PDF 260KB]. A sample of a critical review that includes an annotated structure for your reference.
- How to edit your work [PDF 189KB] . A guide for all students about how to edit and review their work.
- Handy grammar hints [PDF 217KB]. A guide to getting grammar and style right in your assignments.
Business and Law
- Sample essay [PDF 330KB]. A sample of an essay that includes an annotated structure for your reference.
- Writing a business-style report [PDF 330KB]. A resource for business and law students. Find out how to write and format business-style reports.
- Sample of a business-style report [PDF 376 KB]. A resource for business and law students. A sample of a business-style report, with an annotated format.
- Interpreting assignment topics [PDF 370 KB]. Find out how to interpret an assignment topic, including understanding key words and concepts.
- How to edit your work [PDF 189KB]. A guide for all students about how to edit and review their work.

- Walden University
- Faculty Portal
Common Assignments: Journal Entries
Basics of journal entries, related webinar.
Didn't find what you need? Email us at [email protected] .
- Previous Page: Writing a Successful Response to Another's Post
- Next Page: Read the Prompt Carefully
- Office of Student Disability Services
Walden Resources
Departments.
- Academic Residencies
- Academic Skills
- Career Planning and Development
- Customer Care Team
- Field Experience
- Military Services
- Student Success Advising
- Writing Skills
Centers and Offices
- Center for Social Change
- Office of Academic Support and Instructional Services
- Office of Degree Acceleration
- Office of Research and Doctoral Services
- Office of Student Affairs
Student Resources
- Doctoral Writing Assessment
- Form & Style Review
- Quick Answers
- ScholarWorks
- SKIL Courses and Workshops
- Walden Bookstore
- Walden Catalog & Student Handbook
- Student Safety/Title IX
- Legal & Consumer Information
- Website Terms and Conditions
- Cookie Policy
- Accessibility
- Accreditation
- State Authorization
- Net Price Calculator
- Contact Walden
Walden University is a member of Adtalem Global Education, Inc. www.adtalem.com Walden University is certified to operate by SCHEV © 2024 Walden University LLC. All rights reserved.
Writing: Get your essay and assignment written from scratch by PhD expert
Rewriting: Paraphrase or rewrite your friend's essay with similar meaning at reduced cost
Editing: Proofread your work by experts and improve grade at Lowest cost
Enter phone no. to receive critical updates and urgent messages !
Error goes here
Please upload all relevant files for quick & complete assistance.
New User? Start here.
Essay Examples Crafted by Our Experts
Draw inspiration about any topic from our archive of 10K+ premium essays.
Get Comprehensive Assignment Help Services
We have the best minds and resources for you!
Get the best assistance with all essay genres – descriptive, analytical, and more.
Homework Help
From K12 to post doctorate – we offer aid with homework for all academic levels.
Dissertation Help
Our experts are here to help you at every stage of the dissertation writing process.
Coursework Help
Submit coursework on time with quality guidance from our subject matter experts.
Essay Typer
Simplify the task with topic suggestions and data-driven content from our tool.
Plagiarism Checker
Check your academic papers for duplicate text using our free and advanced tool.
Paraphrasing Tool
Restructure sentences accurately without altering the meaning with our free tool.
Conclusion Writing
Leave a lasting impression on readers with conclusions churned by our academic tool.
Essay Samples across ALL Disciplines
All disciplines, why do students look for essay assignment samples.
Students often seek essay samples online for various reasons. They primarily look to gain a better understanding of the order requirements, structure, and formatting. Here are some key motivations behind seeking sample assignments –
- Clarity on Expectations
Essay samples serve as practical examples. They offer clarity on what is expected in terms of content, style, and organization. You can reach our essay helpers for free samples. Students can analyze these references to comprehend the specific requirements of their homework.
- Learning Formatting and Structure
Many students struggle with the correct formatting and structure for academic homework. So, they seek university essay help with free references. By examining such materials, they can observe how to create an introduction, body, and conclusion. They can also show you how to incorporate citations and references properly.
- Ideas and Inspiration
Sometimes, students may find themselves stuck when it comes to generating ideas for their homework. Sample s can spark creativity and provide new perspectives on a particular topic.
- Quality Benchmark
Students often use homework examples as a benchmark for quality. However, not all free assignment help services can ensure top quality. By studying our well-written references, they can gauge the level of detail and critical thinking expected in their own work.
- Avoiding Plagiarism
It's crucial to avoid copying content directly from old materials. However, reviewing them can help students understand how to paraphrase and cite information correctly. You may lack this originality by seeking writing help . This leads to unintentional plagiarism.
- Time Management
With the pressure of deadlines, students may turn to online assignment samples to save time. Analyzing a well-structured reference can expedite their own writing process by providing a model to follow.
- Subject Familiarization
An essay sample can familiarize students with new or complex topics. By reviewing how others have approached similar subjects, students can gain insights into key concepts and terminology. Students can also compare their work to our high-quality essay examples . This allows students to self-assess and identify areas for improvement.
Get Essay References
Find Help with All Subjects with Our Essay Sample
We also cover all the major subjects and more. So, when you hire us, you get free references of these subjects –
- Ancient Civilizations’ Trade Routes
- Impact of World Wars on Society
- Renaissance Art and Culture
- The Industrial Revolution
- Women's Suffrage Movements
- Quantum Physics and its Paradoxes
- CRISPR Technology and Genetic Engineering
- Space Exploration and Colonisation
- Artificial Intelligence in Healthcare
- Climate Change and Environmental Science
- Dystopian Fiction: Exploring Alternate Realities
- Shakespearean Tragedies: Themes and Influences
- Magical Realism in Latin American Literature
- Modern Science Fiction: Themes and Trends
- Contemporary Women Writers: Breaking Stereotypes
- The Beauty of Fractals
- Applications of Game Theory
- History of Number Systems
- Cryptography and Number Theory
- Chaos Theory in Mathematics
- Cognitive Biases and Decision Making
- The Impact of Social Media on Mental Health
- Freudian Psychoanalysis: Concepts and Critiques
- Positive Psychology: The Science of Happiness
- Cross-Cultural Psychology
- Computer Science
- Quantum Computing: Future Possibilities
- Cyber Security Threats and Solutions
- Augmented Reality and Virtual Reality
- Ethics in Artificial Intelligence
- History and Evolution of Programming Languages
- Existentialism: Freedom and Responsibility
- Ethical Dilemmas in Bioethics
- Eastern Philosophy: Zen Buddhism and Taoism
- Political Philosophy: Concepts of Justice
- Philosophy of Mind: Consciousness and Identity
- Globalization and Its Effects on Economies
- Behavioral Economics: Understanding Human Behavior
- Economic Inequality and Redistribution
- Sustainable Development and Economics
- The History of Economic Thought
- Social Movements: Causes and Impacts
- Sociology of Deviance: Breaking Social Norms
- Globalization and Cultural Diversity
- Family Structures and Dynamics
- Urbanization: Challenges and Opportunities
- Impressionism: Capturing Light and Atmosphere
- Street Art and Graffiti as a Form of Expression
- Bauhaus Movement: Design and Architecture
- Photography as Art: Evolution and Impact
- Contemporary Art: Exploring New Mediums
Apart from them, you can log on to MyAssignmenthelp.com and unlock free references of various other subjects.
Search Your Samples
Can I Get Essay and Assignment Help for Free?
Most students live on a shoestring budget. Hence, it is common for them to look for sites that provide free essay assignments. There are various sites, that claims to provide online essays for students at free of cost. However, we never make such claims. Instead, we focus on delivering the best references.
We do get many requests from students asking, "Can you help me with the assignment for free?” As much as we regret not being able to help you with paper writing for free, you can certainly get the materials at no extra cost. We have an entire online library that students can use whenever needed.
Students simply need to hire our writers. Once they have availed of their services, they become eligible for a host of other benefits. From college assignment samples to university essay assignment samples, students can access all kinds of resources from our collection. Our top-qualified experts curate all these solved papers.
So, if you check a sample of our essay writing, you can immediately notice the difference in quality between us and a fake. We also regularly update our essay samples. So, if you are looking for essay writing examples on the latest topics, like "Hospitality after the COVID-19 pandemic" or "The latest developments in IT, " we have just the right resources for you.
Industry experts meticulously write all our materials by following the correct formatting and citations. Getting a high level of example of essays for no extra cost is certainly an offer you cannot miss. So, instead of looking for essay help for free online, reach us today. Simply hire a writer and get access to unlimited references for free.
Click For Free Materials
Use our Samples to score Straight As
Students can use sample assignments effectively to improve their grades. Follow this strategic approach to use our materials to get the highest grades –
Firstly, review the materials thoroughly. With our help, students get free access to the best essay examples. Students can refer to these high-quality references to understand the expected structure and format for their papers. This enables students to organize their ideas coherently, making their work more comprehensible to instructors.
Additionally, analyzing old reference materials help students grasp the tone and style appropriate for academic writing. This ensures that their own work aligns with the expected standards. This enhances its overall quality. By identifying a good sample assignment, students can incorporate similar language and vocabulary into their writing. This demonstrates a deeper understanding of the subject matter.
Furthermore, an essay sample can serve as a valuable resource for mastering citation styles. Proper referencing is crucial in academic writing. Studying well-cited free essay assignments assists students in correctly citing sources. Thus, students can easily avoid plagiarism, which bolsters the credibility of their work.
Learning from old references also enables students to recognize common mistakes. With our essay help, they get free references and guidance. They understand where others have faltered and can pre-emptively address such potential pitfalls in their own papers. Thereby, it improves the overall coherence and argumentative strength of their work.
Lastly, using an essay sample can act as a benchmark for research and analysis. Students can critically evaluate how evidence is presented in an old college assignment sample. They can check how the arguments are constructed in university essay assignment samples and refine their analytical skills. This helps you to write more compelling and well-supported arguments than any free essay assignments.
Click To Connect
Frequently Asked Questions
Join us before it’s too late..
That's our welcome gift for first time visitors
Sign up now and Get The Exclusive offer* Limited Time Only
- Mailing List
- Search Search
Username or Email Address
Remember Me

Resources for Teachers: Creating Writing Assignments
This page contains four specific areas:
Creating Effective Assignments
Checking the assignment, sequencing writing assignments, selecting an effective writing assignment format.
Research has shown that the more detailed a writing assignment is, the better the student papers are in response to that assignment. Instructors can often help students write more effective papers by giving students written instructions about that assignment. Explicit descriptions of assignments on the syllabus or on an “assignment sheet” tend to produce the best results. These instructions might make explicit the process or steps necessary to complete the assignment. Assignment sheets should detail:
- the kind of writing expected
- the scope of acceptable subject matter
- the length requirements
- formatting requirements
- documentation format
- the amount and type of research expected (if any)
- the writer’s role
- deadlines for the first draft and its revision
Providing questions or needed data in the assignment helps students get started. For instance, some questions can suggest a mode of organization to the students. Other questions might suggest a procedure to follow. The questions posed should require that students assert a thesis.
The following areas should help you create effective writing assignments.
Examining your goals for the assignment
- How exactly does this assignment fit with the objectives of your course?
- Should this assignment relate only to the class and the texts for the class, or should it also relate to the world beyond the classroom?
- What do you want the students to learn or experience from this writing assignment?
- Should this assignment be an individual or a collaborative effort?
- What do you want students to show you in this assignment? To demonstrate mastery of concepts or texts? To demonstrate logical and critical thinking? To develop an original idea? To learn and demonstrate the procedures, practices, and tools of your field of study?
Defining the writing task
- Is the assignment sequenced so that students: (1) write a draft, (2) receive feedback (from you, fellow students, or staff members at the Writing and Communication Center), and (3) then revise it? Such a procedure has been proven to accomplish at least two goals: it improves the student’s writing and it discourages plagiarism.
- Does the assignment include so many sub-questions that students will be confused about the major issue they should examine? Can you give more guidance about what the paper’s main focus should be? Can you reduce the number of sub-questions?
- What is the purpose of the assignment (e.g., review knowledge already learned, find additional information, synthesize research, examine a new hypothesis)? Making the purpose(s) of the assignment explicit helps students write the kind of paper you want.
- What is the required form (e.g., expository essay, lab report, memo, business report)?
- What mode is required for the assignment (e.g., description, narration, analysis, persuasion, a combination of two or more of these)?
Defining the audience for the paper
- Can you define a hypothetical audience to help students determine which concepts to define and explain? When students write only to the instructor, they may assume that little, if anything, requires explanation. Defining the whole class as the intended audience will clarify this issue for students.
- What is the probable attitude of the intended readers toward the topic itself? Toward the student writer’s thesis? Toward the student writer?
- What is the probable educational and economic background of the intended readers?
Defining the writer’s role
- Can you make explicit what persona you wish the students to assume? For example, a very effective role for student writers is that of a “professional in training” who uses the assumptions, the perspective, and the conceptual tools of the discipline.
Defining your evaluative criteria
1. If possible, explain the relative weight in grading assigned to the quality of writing and the assignment’s content:
- depth of coverage
- organization
- critical thinking
- original thinking
- use of research
- logical demonstration
- appropriate mode of structure and analysis (e.g., comparison, argument)
- correct use of sources
- grammar and mechanics
- professional tone
- correct use of course-specific concepts and terms.
Here’s a checklist for writing assignments:
- Have you used explicit command words in your instructions (e.g., “compare and contrast” and “explain” are more explicit than “explore” or “consider”)? The more explicit the command words, the better chance the students will write the type of paper you wish.
- Does the assignment suggest a topic, thesis, and format? Should it?
- Have you told students the kind of audience they are addressing — the level of knowledge they can assume the readers have and your particular preferences (e.g., “avoid slang, use the first-person sparingly”)?
- If the assignment has several stages of completion, have you made the various deadlines clear? Is your policy on due dates clear?
- Have you presented the assignment in a manageable form? For instance, a 5-page assignment sheet for a 1-page paper may overwhelm students. Similarly, a 1-sentence assignment for a 25-page paper may offer insufficient guidance.
There are several benefits of sequencing writing assignments:
- Sequencing provides a sense of coherence for the course.
- This approach helps students see progress and purpose in their work rather than seeing the writing assignments as separate exercises.
- It encourages complexity through sustained attention, revision, and consideration of multiple perspectives.
- If you have only one large paper due near the end of the course, you might create a sequence of smaller assignments leading up to and providing a foundation for that larger paper (e.g., proposal of the topic, an annotated bibliography, a progress report, a summary of the paper’s key argument, a first draft of the paper itself). This approach allows you to give students guidance and also discourages plagiarism.
- It mirrors the approach to written work in many professions.
The concept of sequencing writing assignments also allows for a wide range of options in creating the assignment. It is often beneficial to have students submit the components suggested below to your course’s STELLAR web site.
Use the writing process itself. In its simplest form, “sequencing an assignment” can mean establishing some sort of “official” check of the prewriting and drafting steps in the writing process. This step guarantees that students will not write the whole paper in one sitting and also gives students more time to let their ideas develop. This check might be something as informal as having students work on their prewriting or draft for a few minutes at the end of class. Or it might be something more formal such as collecting the prewriting and giving a few suggestions and comments.
Have students submit drafts. You might ask students to submit a first draft in order to receive your quick responses to its content, or have them submit written questions about the content and scope of their projects after they have completed their first draft.
Establish small groups. Set up small writing groups of three-five students from the class. Allow them to meet for a few minutes in class or have them arrange a meeting outside of class to comment constructively on each other’s drafts. The students do not need to be writing on the same topic.
Require consultations. Have students consult with someone in the Writing and Communication Center about their prewriting and/or drafts. The Center has yellow forms that we can give to students to inform you that such a visit was made.
Explore a subject in increasingly complex ways. A series of reading and writing assignments may be linked by the same subject matter or topic. Students encounter new perspectives and competing ideas with each new reading, and thus must evaluate and balance various views and adopt a position that considers the various points of view.
Change modes of discourse. In this approach, students’ assignments move from less complex to more complex modes of discourse (e.g., from expressive to analytic to argumentative; or from lab report to position paper to research article).
Change audiences. In this approach, students create drafts for different audiences, moving from personal to public (e.g., from self-reflection to an audience of peers to an audience of specialists). Each change would require different tasks and more extensive knowledge.
Change perspective through time. In this approach, students might write a statement of their understanding of a subject or issue at the beginning of a course and then return at the end of the semester to write an analysis of that original stance in the light of the experiences and knowledge gained in the course.
Use a natural sequence. A different approach to sequencing is to create a series of assignments culminating in a final writing project. In scientific and technical writing, for example, students could write a proposal requesting approval of a particular topic. The next assignment might be a progress report (or a series of progress reports), and the final assignment could be the report or document itself. For humanities and social science courses, students might write a proposal requesting approval of a particular topic, then hand in an annotated bibliography, and then a draft, and then the final version of the paper.
Have students submit sections. A variation of the previous approach is to have students submit various sections of their final document throughout the semester (e.g., their bibliography, review of the literature, methods section).
In addition to the standard essay and report formats, several other formats exist that might give students a different slant on the course material or allow them to use slightly different writing skills. Here are some suggestions:
Journals. Journals have become a popular format in recent years for courses that require some writing. In-class journal entries can spark discussions and reveal gaps in students’ understanding of the material. Having students write an in-class entry summarizing the material covered that day can aid the learning process and also reveal concepts that require more elaboration. Out-of-class entries involve short summaries or analyses of texts, or are a testing ground for ideas for student papers and reports. Although journals may seem to add a huge burden for instructors to correct, in fact many instructors either spot-check journals (looking at a few particular key entries) or grade them based on the number of entries completed. Journals are usually not graded for their prose style. STELLAR forums work well for out-of-class entries.
Letters. Students can define and defend a position on an issue in a letter written to someone in authority. They can also explain a concept or a process to someone in need of that particular information. They can write a letter to a friend explaining their concerns about an upcoming paper assignment or explaining their ideas for an upcoming paper assignment. If you wish to add a creative element to the writing assignment, you might have students adopt the persona of an important person discussed in your course (e.g., an historical figure) and write a letter explaining his/her actions, process, or theory to an interested person (e.g., “pretend that you are John Wilkes Booth and write a letter to the Congress justifying your assassination of Abraham Lincoln,” or “pretend you are Henry VIII writing to Thomas More explaining your break from the Catholic Church”).
Editorials . Students can define and defend a position on a controversial issue in the format of an editorial for the campus or local newspaper or for a national journal.
Cases . Students might create a case study particular to the course’s subject matter.
Position Papers . Students can define and defend a position, perhaps as a preliminary step in the creation of a formal research paper or essay.
Imitation of a Text . Students can create a new document “in the style of” a particular writer (e.g., “Create a government document the way Woody Allen might write it” or “Write your own ‘Modest Proposal’ about a modern issue”).
Instruction Manuals . Students write a step-by-step explanation of a process.
Dialogues . Students create a dialogue between two major figures studied in which they not only reveal those people’s theories or thoughts but also explore areas of possible disagreement (e.g., “Write a dialogue between Claude Monet and Jackson Pollock about the nature and uses of art”).
Collaborative projects . Students work together to create such works as reports, questions, and critiques.

Want to create or adapt books like this? Learn more about how Pressbooks supports open publishing practices.
43 Sample Writing Assignments
Process analysis.
- Be the expert and teach your reader how to do something. You will focus on writing the main steps to completing this process and organizing based on chronology or priority of steps. This essay could be demonstrative in nature (ex. How to bathe and groom a dog at home or how to make banana nut muffins) or philosophically-based (ex. How to not fail your freshman classes or how to survive being a camp counselor).
- Interview a person who has a compelling story to share or with whom you can focus on a particular angle. Interweave direct quotes, observations, and narrative elements to help readers understand this person or his/her perspective better. This should not be a full biography of this person or a career profile although biographical elements and details on jobs held may be woven in as appropriate. Perspective/angles can include but are not limited to this person’s relationship with: technology, ecology/the environment, politics, religion/spirituality, gender roles, family/what family means to them, love, betrayal, etc.
The purpose is to define a term, concept, or idea. You will typically lay the foundation with a dictionary definition (denotative) of the word but will move out to an extended definition (connotative). You are using a combination of the literal and implied meanings of a word or idea in addition to historical information to help readers understand the topic more effectively. You usually define a term or concept that is complex in nature or that can be misconstrued. Legal, business, and scientific terms or concepts typically work well for essays such as these. For example, take the term manslaughter; the literal and implied meanings can help to understand this sometimes misinterpreted crime. The definition paper can stand on its own or an abbreviated version can serve as part of a larger argumentative, analytical, or research paper.
- Take an abstract, complex, controversial word or a term that is personal to you. Using a dictionary definition as well as your own and others’ interpretations, craft an extended definition essay with the purpose of giving a more insightful, comprehensive, and layered understanding of this particular term.
Illustration
- Walk readers through a day, event, activity, or state of mind, making sure to focus on facts and authentic descriptions. Take a topic that you know something about, like video games. This expository essay on this topic could focus one’s addiction to video games. In one example, the writer might walk us through his day playing a video game while in another essay on the same topic, a writer could concentrate mostly on the reasons he or she became addicted to games and may touch briefly on how to prevent this addiction.
This prompt comes from: https://resources.instructure.com/courses/5/pages/summary-essay-prompt . This prompt could be easily modified by changing out the topic of digital literacy to another one of your choice. Whether working with popular articles or scholarly ones, summary writing is a key component of reading comprehension, setting up a foundation for a larger issue, and research.
“Digital literacy” may be a new term for you, but it’s probably not a new concept. Our personal and academic lives are being transformed by online content, and not everyone has the same innate level of skill at determining what and how to use this content.
Our first essay asks to you to summarize one of 3 short articles from the library library on the topic of digital literacy. (These articles can be found in the weekly modules.) The objectives of this assignment are to:
- Identify and restate the thesis of an author’s work
- Accurately portray the contents of an article
- Practice paraphrasing and quotation skills in formal writing
- Practice “neutral reporting”–being able to present the findings of others without making them appear as your own (most students find this to be the most challenging component of this assignment)
- Practice end citation methods (APA or MLA)
- This summary should be written for an audience that HAS NOT read the original article, and so you will report the major and minor ideas contained in the piece.
- Your summary should indicate the article’s thesis idea, if there is one. This thesis should be contained within your introduction. Be sure to also give the title of the article, the author(s), and where & when it was originally published.
- This summary should contain at least one direct quote from the article. Quotation marks should be used. Introduce the direct quote with a “signal” phrase, such as McMillan-Clifton writes… or the article states… or this website argues that… etc.
- This summary should contain at least one paraphrase from the article. Put the author’s ideas in your own words, but stay true to the original intent. Introduce the paraphrase with a tag phrase, as mentioned above. Remember that, as a general rule, phrases of 4 or more words that are exactly the same as the original text should be treated like a quote, not a paraphrase.
- Your summary will be NEUTRAL regarding the material contained within the article. While you should report any bias the author has, you yourself should not reveal your own opinions on the matter. Using “I” or “you” in this essay is not advised , unless it appears inside a quote.
- Normally I don’t mind if essay submissions exceed the maximum word limit, but this essay is an exception. Because one of the hallmarks of an effective summary is brevity, please do not exceed the maximum word count of 600 words.
- In-text citations will not be required in this assignment, though you are welcome to include them for practice.
- Your summary should have an end citation, APA or MLA
Write What Matters Copyright © 2020 by Liza Long; Amy Minervini; and Joel Gladd is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License , except where otherwise noted.
Share This Book

A to Z | Search
Writing Across the Curriculum
Sample assignments.
This page provides two downloadable documents: a set of Low Stakes writing assignments, and guidelines for High Stakes writing assignments. The documents are available in .docx copies to allow for revision and customization. You’re welcome to take what you need, please keep the Augsburg logo intact (other downloadable logos are available here ).
Click HERE to download a full set of sample Low Stakes assignment prompts.
Click HERE to download a set of sample High Stakes assignment guidelines.
You can learn more about the benefits of differentiating between low and high stakes assignments in Peter Elbow’s (1997) essay, “High stakes and low stakes in assigning and responding to writing” from Writing to Learn: Strategies for Assigning and Responding to Writing across the Discipline: New Directions for Teaching and Learning.
LOW STAKES WRITING
low stakes writing is:.
- Free writing in response to a simple prompt
- A simple, informal way to integrate writing in the classroom
- “Low effort, high impact”
- Easy to incorporate at the beginning or end of class
- Low-stress, and typically involves little to no grading
Low stakes writing helps:
- Describe, apply, and retain information
- Explore and personalize ideas
- Focus thoughts and questions
- Demonstrate the value of writing as a part of the learning process
- Informally engage each student in the classroom
- Improve high-stakes writing
- Efficiently assess student learning
A brief sample of low stakes prompts:
- What do you already know about this topic that can guide your learning?
- What have you learned from similar assignments that can help you succeed on this one?
- Summarize today’s lecture in one sentence.
- What do you feel like you learned today, and what lingering question do you have?
- Write an email to a friend who has been absent for a week and explain what they’ve missed. Aim to be comprehensive rather than writing a list.
HIGH STAKES WRITING
High stakes writing assignments:.
- Correspond to writing conventions in the discipline/genre
- Are typically formal and academic in style
- Develop over time through drafting and sequencing/scaffolding
- Require conducting effective research
- Depend on effective, close reading
- Synthesize complex information
- Are more sophisticated in thought and prose
Basic Guidelines
- Regard writing as a process rather than a product
- Clearly connect the assignment to course learning objectives
- Provide students with a clear assignment prompt detailing expectations
- Provide students with a rationale for those expectations
- Articulate the audience for the writer (Experts? A publication? You?)
- Use assignment sequencing/scaffolding (suggestions below and here )
- Include opportunities for feedback and related revision
- Provide effective feedback on drafts (suggestions here and here )
- Review suggested rubric options here
- Weight the assignment accordingly, usually assigning significant value in the overall course grading system
- Assign value (i.e. a grade or other form of credit) to reading assignments
High stakes writing helps to:
- Familiarize students with disciplinarity and writing in a genre
- Describe, apply, and retain complex disciplinary information
- Develop more advanced writing, thinking, learning, and process skills
- Develop self-assessment and revision skills
- Focus on developing depth rather than breadth
- Improve higher order learning/thinking
- Thoroughly assess student learning and content mastery
- Teach students to handle competing information and develop thesis
- Make use of in-class peer review activities to help crowd-source feedback
- Provide examples of previous work from students (with their permission) along with the original assignment description
- Focus on minimal comments in the margins and identify 1-3 strategies for improvement at the end of a draft
- Identify common strengths/weaknesses of the class and discuss those with the class as a whole
- Identify successful examples of student work in class for discussion
- Cover common mistakes in the original assignment description or when discussing the assignment, use low-stakes writing to reiterate the points
- If you don’t have time to teach a writing topic, such as citation style, link students to effective guides
Key high stakes writing resources:
- These writing guides are written for a student audience, they overview conventions of writing and conducting research in various academic disciplines across both the Sciences and Humanities.
- Search topically through hundreds of undergraduate and graduate courses by discipline or topic and access course syllabi, readings, and assignment documents.
- This webpage provides guides to some of the best online resources for helping instructors incorporate writing curriculum into their classrooms. Links address topics such as developing learning objectives, designing assignments, approaches to assessment, writing instruction handouts, and tutorials on references and citation.
Click HERE to download a more detailed set of sample High Stakes assignment guidelines.
- UWF Libraries
Literature Review: Conducting & Writing
- Sample Literature Reviews
- Steps for Conducting a Lit Review
- Finding "The Literature"
- Organizing/Writing
- APA Style This link opens in a new window
- Chicago: Notes Bibliography This link opens in a new window
- MLA Style This link opens in a new window
Sample Lit Reviews from Communication Arts
Have an exemplary literature review.
- Literature Review Sample 1
- Literature Review Sample 2
- Literature Review Sample 3
Have you written a stellar literature review you care to share for teaching purposes?
Are you an instructor who has received an exemplary literature review and have permission from the student to post?
Please contact Britt McGowan at [email protected] for inclusion in this guide. All disciplines welcome and encouraged.
- << Previous: MLA Style
- Next: Get Help! >>
- Last Updated: Mar 22, 2024 9:37 AM
- URL: https://libguides.uwf.edu/litreview
Purdue Online Writing Lab Purdue OWL® College of Liberal Arts
APA Sample Paper

Welcome to the Purdue OWL
This page is brought to you by the OWL at Purdue University. When printing this page, you must include the entire legal notice.
Copyright ©1995-2018 by The Writing Lab & The OWL at Purdue and Purdue University. All rights reserved. This material may not be published, reproduced, broadcast, rewritten, or redistributed without permission. Use of this site constitutes acceptance of our terms and conditions of fair use.
Note: This page reflects APA 6, which is now out of date. It will remain online until 2021, but will not be updated. The equivalent APA 7 page can be found here .
Media File: APA Sample Paper
This resource is enhanced by an Acrobat PDF file. Download the free Acrobat Reader
Click this link to download the PDF handout of the APA Sample Paper.
By continuing to browse the site you are agreeing to our use of cookies and similar tracking technologies described in our privacy policy .
Evaluating without Grading
/ AHA Resource Library
/ Evaluating without Grading
Published Date
September 10, 2021
Resource Type
For the Classroom
Teaching Methods
Teaching & Learning, Undergraduate Education
By Luke Clossey and Esther Souman
For more information on this course and specifications grading, read “ Evaluating without Grading: Encouraging Students to Master Skills with Specifications Grading ,” Perspectives on History , October 2021.
In our Introduction to Global History survey course at Simon Fraser University (SFU) in autumn 2020, students earned their grades by “unlocking” different levels of various assignments. We introduced a specifications grading system, evaluating individual assignments on a pass/fail basis, and created multiple pathways for students to move through the assignments and earn a final grade. Our goal was to ensure that every student mastered basic skills, and the more advanced students had opportunities to develop more creative projects. This system was adapted from Linda B. Nilson’s authoritative Specifications Grading (Stylus, 2015).
Assignments and Evaluations
There are no quizzes or examinations. This course uses a “specifications” grading model: Students undertake a number of short assignments, and receive a course grade based on the quantity and difficulty of the assignments successfully completed. Each assignment will involve options, so students can work on something relevant to their own lives. Each course grade also requires a specified minimal level of attendance and participation. To earn a grade, you must meet both the participation-score minimum and the minimum number/level of assignments. This approach has been shown to minimize stress and maximize long-term learning. Overall course grading will be on a curve that can only benefit students. If necessary, final grades will be increased to coincide with departmental averages.
| Grade | Participation Score Minimum | Minimum Number/Level of Assignments |
|---|---|---|
| A+ | 95 | L1 L2 L3 L4 |
| A | 90 | L1 L2 L3 L4 |
| A- | 80 | L1 L2 L3 L4 |
| B+ | 95 | L1 L2 L3 |
| B | 85 | L1 L2 L3 |
| B- | 75 | L1 L2 L3 |
| C+ | 80 | L1 L2 |
| C | 70 | L1 L2 |
| C- | 60 | L1 L2 |
| D | 60 | L1 |
You may submit one assignment (upload to Canvas) for each Intake, with the following deadlines:
First Intake: Oct. 7, 11:59 p.m.
Second Intake: Oct. 21, 11:59 p.m.
Third Intake: Nov. 11, 11:59 p.m.
Fourth Intake: Dec. 2, 11:59 p.m.
For the first Intake, everyone is invited to submit an L1 assignment. In subsequent Intakes, to submit an assignment of a specific level, you must have previously passed an assignment at the level beneath it. That is, to submit an L3 assignment, you must have already successfully completed an L2 assignment. To earn an A grade, you would submit a successful assignment for L1 at the first Intake, for L2 at the second, etc. To earn a C grade, you would pass an L1 and an L2 assignment, and would have no obligation to submit any others.
An assignment can be submitted late, with a participation-score penalty of 3 points per 24-hour-period, or fraction thereof, after the deadline. The teaching staff is happy to discuss assignment requirements, and to discuss how you might go about completing them, but cannot confirm in advance that a draft assignment fulfills the requirements. The teaching staff is not expert in the skills needed for the various assignments, so, depending on your choice, you may have to work with a significant degree of independence. You are encouraged to take advantage of the SFU Student Learning Commons for assistance with writing and study skills.
Requirements for All Assignments
- Completeness (all required items submitted)
- Professionalism in writing quality and presentation: clear organization, no sentence fragments or run-ons, no more than two errors per 250 words (including citations in Chicago style)
- Course relevance: must cite one or more podcasts, and one or more of the assigned sources
- No plagiarism (see “Academic Policies” below)
- All papers should be written according to the instructor’s “How to Write” and the Chicago Manual of Style or Turabian’s A Manual for Writers of Term Papers .
Level 1: Summarize!
Summarize the podcast(s) and sources from a single week of your choice. Do not include external information or analysis.
- Use prose, with paragraphs, as if in an essay (i.e., not in list-form)
- Between 500 and 750 words (include a word count)
- Organization should be clear, with transitions between paragraphs
- Information should be presented as a coherent whole
- Include citations for the podcast(s) and sources
Level 2: Analyze!
Analyze a source or a collection of sources. Make an explicit argument that demonstrates insight. The argument should be original, significant and interesting. (You should make clear to the reader the originality, significance, and interest.) The argument should not be a fact—if all scholars would agree with it, it is probably not an argument.
- Use prose, with paragraphs, as if in an essay
- Between 750 and 1,000 words (include a word count)
- You may include external information, properly cited, if it helps your argument
- Include a literature review, which you might use as a tool to demonstrate your originality
- Include a citation for the source(s) you analyze
| • podcast • assigned source • digital museum collection item (example) • image • sound recording (e.g., from radiooooo.com) • dataset (e.g., list of constitutions (example), www.slavevoyages.org, orbis.stanford.edu) • computer game | • lectio dificilior potius (to identify which of two versions of a text is older; we'll learn about this in Week 5) • global perspective: show the regional assumptions and motivations behind the source • find patterns in time and space |
Some project ideas
- Take a source of uncertain origins, and argue for a specific region or time of origin
- Choose 20 tracks from https://radiooooo.com , and find patterns across time or space (Is music in 1920s Africa more complex than music in 1930s Asia? Why?)
- Look at your diet for a week. How modern/premodern is it? How traditional/global? Write an essay about the history of one food you’ve eaten or about one quality found in several of your foods. How are other peoples’ dietary practices different from your own?
- Choose one of these texts. Make a list of all the assumptions it makes. Analyze each assumption from a global or premodern perspective.
Level 3: Create!
Your creation should demonstrate its clear, significant value, and have a professional appearance. Your creation should be accompanied by a commentary (300 to 500 words) explaining your goals for the project and how you sought to meet them (unless you are doing a proposal).
Personal Applications
- Create a global-history timeline, with events chosen for relevance to your own life.
- Create a global-history map, with places chosen for relevance to your own life.
- Choose things important to you (in categories such as technology, biology, ideas, recreation, food…) and then historicize and globalize them.
- Choose the most important events in global history. Justify your choice. Create chains of causality between them. Analyze the results.
- Identify an identity and trace it through space/time (e.g., your family, vegans, or whatever).
- Describe a particular world-perspective (e.g., Buddhism ca. 200 BC), and apply it to a modern problem.
Pedagogy Tools
- Create a infographic/visualization ( example 1 , example 2 )
- Create a textbook section
- Create a historical roleplaying session
- Create a map (e.g., Google map or a Carto map )
- Create pedagogical materials (with an instructor’s permission)
- Identify 4 to 6 learning objectives for HIST 130 and connect them to details in the course
- Design a final exam for HIST 130
Research Tools
- Write a research-paper proposal
- Compile an annotated bibliography of at least 25 items
- Create an annotated dataset of at least 25 items
- Write a Teaching and Learning Development Grant proposal
Level 4: Synthesize!
Your synthesis should demonstrate a clear, significant value, and have a professional appearance.
- Write a research paper (750 to 1,500 words). Explain the research question, why it is important, the state of the scholarship, and what sources or approached your paper uses to answer it.
- Write a policy proposal for the federal government (750-word limit).
- Sum up the history of the world in a single minute (video/audio) or in a 500-word-maximum essay. Include an essay (750 to 1,000 words) justifying the choices informing it.
- Write a small SSHRC grant. (Ignore Sections VI and VIII; you can fabricate any answers for Section I.)
Final Grades
This table shows the distribution of grades in the course, slightly randomized to preserve privacy.
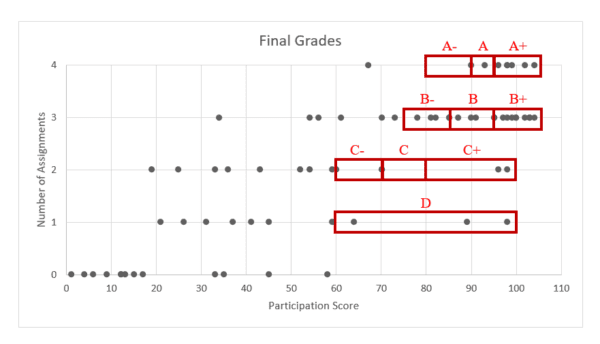
Related Resources

June 20, 2024
16 Months to Sumter: Newspaper Editorials on the Path to Secession

June 16, 2024
The History of Racism and Racist Violence: International Contexts and Comparisons
The history of racism and racist violence: monuments and museums, join the aha.
The AHA brings together historians from all specializations and all work contexts, embracing the breadth and variety of activity in history today.
Content Warning
This page contains words or ideas that might be offensive to modern readers. To maintain the accuracy of historical documentation, the content is reprinted in its entirety as it was originally published. This accurate reproduction of original historical texts therefore contains words and ideas that do not reflect the editorial decisions or views of the American Historical Association.
- Search Menu
- Sign in through your institution
- Advance articles
- Author Guidelines
- Submission Site
- Open Access Policy
- Self-Archiving Policy
- Why publish with this journal?
- About Bioinformatics Advances
- About the International Society for Computational Biology
- Editorial Board
- Advertising & Corporate Services
- Diversity, Equity and Inclusion
- Journals on Oxford Academic
- Books on Oxford Academic

Article Contents
1 introduction, 4 discussion, author contributions, supplementary data, conflict of interest.
- < Previous
Demultiplexing of single-cell RNA-sequencing data using interindividual variation in gene expression
- Article contents
- Figures & tables
- Supplementary Data
Isar Nassiri, Andrew J Kwok, Aneesha Bhandari, Katherine R Bull, Lucy C Garner, Paul Klenerman, Caleb Webber, Laura Parkkinen, Angela W Lee, Yanxia Wu, Benjamin Fairfax, Julian C Knight, David Buck, Paolo Piazza, Demultiplexing of single-cell RNA-sequencing data using interindividual variation in gene expression, Bioinformatics Advances , Volume 4, Issue 1, 2024, vbae085, https://doi.org/10.1093/bioadv/vbae085
- Permissions Icon Permissions
Pooled designs for single-cell RNA sequencing, where many cells from distinct samples are processed jointly, offer increased throughput and reduced batch variation. This study describes expression-aware demultiplexing (EAD), a computational method that employs differential co-expression patterns between individuals to demultiplex pooled samples without any extra experimental steps.
We use synthetic sample pools and show that the top interindividual differentially co-expressed genes provide a distinct cluster of cells per individual, significantly enriching the regulation of metabolism. Our application of EAD to samples of six isogenic inbred mice demonstrated that controlling genetic and environmental effects can solve interindividual variations related to metabolic pathways. We utilized 30 samples from both sepsis and healthy individuals in six batches to assess the performance of classification approaches. The results indicate that combining genetic and EAD results can enhance the accuracy of assignments (Min. 0.94, Mean 0.98, Max. 1). The results were enhanced by an average of 1.4% when EAD and barcoding techniques were combined (Min. 1.25%, Median 1.33%, Max. 1.74%). Furthermore, we demonstrate that interindividual differential co-expression analysis within the same cell type can be used to identify cells from the same donor in different activation states. By analysing single-nuclei transcriptome profiles from the brain, we demonstrate that our method can be applied to nonimmune cells.
EAD workflow is available at https://isarnassiri.github.io/scDIV/ as an R package called scDIV (acronym for single-cell RNA-sequencing data demultiplexing using interindividual variations).
Although single-cell analyses are beginning to unravel the molecular aetiology of diseases, most studies incorporate average gene expression profiles across bulk tissues, which frequently mask variation between individuals visible at single-cell resolution ( van der Wijst et al. 2018 , Yazar et al. 2022 ). The variability between individuals remains an understudied aspect of co-expression relationships between molecules at the single-cell level. Establishing molecular co-dependencies may break down due to genetic variation, uncoupling of transcription and splicing, or it could be caused by pathological insult. The regulation of downstream markers may be disrupted by altered interindividual co-expression relationships, which in turn affect the expression pattern of entire biological pathways ( Johansen et al. 2023 ).
Previous studies have shown that SNPs can alter co-expression relationships on an individual donor basis by looking for evidence of allele-specific correlation ( Fairfax et al. 2012 , van der Wijst et al. 2018 , Oelen et al. 2022 ). Here, we develop a computational method, expression-aware demultiplexing (EAD), that harnesses variation of interindividual co-expression signatures at single-cell resolution to stratify donors and improve the demultiplexing of pooled scRNA-seq data ( Fig. 1 ).

Workflow for computational demultiplexing of unrelated individuals in scRNA-seq. (a) First, we demultiplex pooled samples (vireo) ( Huang et al. 2019 ) using genetic differences inferred from scRNA-seq data (cellsnp-lite) ( Huang and Huang 2021 ). (b) Next, for each pair of individuals in the output of genetic demultiplexing, we estimate accurate gene expression values for all genes. ( g ) per cell using the gene expression recovery for single-cell RNA sequencing (SAVER) ( Huang et al. 2018 ). (c) We apply the LASSO to obtain the most representative subset of genes ( s ) ( Nassiri and McCall 2018 ) related to query gene ( q ). (d) We apply differential gene correlation analysis between pairs of query gene ( q ) and related genes selected by LASSO ( SG ), to identify the top first DCE genes interindividual (see section 2). (e) The co-expression patterns of the top first DCE genes ( q and g ) are used to fit a mixture model and reconstruct the sample identity of each cell.
The process of separating cells from multiple samples pooled in a single batch is referred to as demultiplexing in this article. Multiplexing can increase the number of donors that can be tested, reduce experiment costs, address batch effects, and make large-scale sample operations feasible. Multiple approaches can be employed to demultiplex pooled single-cell gene expression profiles, including DNA oligonucleotide tagging and natural genetic variation ( Huang et al. 2019 ).
Inspired by the differential co-expression (DCE) analysis of two genes between biological groups, we introduce an optimized statistical method for donor stratification guided by variation of co-expression signatures at single-cell resolution ( McKenzie et al. 2016 ).
Given cells with known sample identity, we demonstrate that differences in gene–gene relationships exist between individuals by providing multiple examples. The results show top interindividual DCE genes provide a distinct cluster of cells per individual and display the enrichment of cellular macromolecular super-complexes, or organelles related to metabolism (e.g. mitochondria and ribosomes).
We applied this approach to samples from isogenic inbred mice and showed that by controlling genetic and environmental effects, we can resolve interindividual variations related to metabolic pathways.
2.1 Sources of data
Sequencing data for evaluation and application covers 124 10× single-cell RNA-seq samples in 14 batches and 30 bulk RNA samples.
During library preparation for the first and second datasets (see sections 1.1.1 and 1.1.2), every sample (a group of cells that originated from the same donor) was assigned a specific index sequence. Sample indices were incorporated into the sequencing primers on Illumina sequencers. After sequencing, by using the 10× mkfastq pipeline the names of sample index sets were automatically identified and reads per sample were merged into the FASTQ files. Supplementary Tables 4 and 5 included reference tables with the complete set of index codes per donor. Accuracy and reliability of the embedded sample index sequences are hallmarks of the 10× Genomics platform. This provides a solid foundation for evaluating the performance of demultiplexing algorithms without introducing errors from the ground truth itself.
2.1.1 Dataset 1
Eight metastatic melanoma (MM) patients were included in the first scRNA-seq data set, which contains human peripheral blood mononuclear cells (PBMCs) with known donor labels ( Fairfax et al. 2020 ). In this dataset, sample indices were used to label cells per sequencing sample without errors. By using sample/donor indices, we were able to map demultiplexed cells using the expression-aware approach to their original sample donors and assess their performance in singlet assignment ( Supplementary Table 1 ). Two sample pools cover 16 samples pre- and posttreatment at day 21 (eight pre- and eight posttreatment) with immune checkpoint blockade therapies including Nivolumab (NIVO) + ipilimumab (IPI) or Pembrolizumab (Pembro) ( Fairfax et al. 2020 ). We chose this dataset to show that interindividual differences and no other confounding factors, such as cell type, underlie differential gene co-expression across individuals. We used the Cell Ranger pipeline (v7.0.1), the GRCh38 reference genome, and 5′ R2-only chemistry to process the data set. The analysis detected up to 36 842 cells per pool, 56 850 mean reads per cell, and 1768 median genes per cell. Raw data for single-cell sequencing datasets have been deposited on the European Genome–phenome Archive ( https://ega-archive.org/studies/EGAS00001004081 ).
2.1.2 Dataset 2
The second scRNAseq data set consists of a pooled sample with known donor labels from six isogenic mice. The samples were exposed to a topical TLR7 agonist, Imiquimod, to induce a systemic lupus erythematosus-like phenotype or vehicle control. In this dataset, cells were labelled per sequencing sample using sample indices ( Supplementary Table 3 ). We applied this dataset to show that by controlling genetic and environmental effects, we can resolve interindividual variations related to metabolic pathways. The Cell Ranger pipeline, mm10-2020-A reference genome, and single-cell 3′ v3 chemistry were employed for data processing. The analysis detected up to 12 685 cells per pool, with an average of 40 101 mean reads per cell and 1553 median genes per cell.
2.1.3 Dataset 3
The third data include five single-cell multi-omics (RNA-seq + ATAC-seq) batches with unknown donor labels. The dataset consists of circulating haematopoietic progenitor cells samples from seven healthy controls, 15 sepsis patients, and eight convalescent samples (six samples per batch) ( Kwok et al. 2023 ). The genetic demultiplexing pool samples were based on the use of an extra 30 bulk RNA-sequencing profiles from the same individuals. This dataset was chosen to evaluate the utility of EAD for challenging biological models involving a mixture of heterogeneous cell types and donors. The data set was processed using the Cell Ranger pipeline, GRCh38 reference genome, and Multiome chemistry. The analysis detected up to 14 806 cells per pool, 49 892 mean reads per cell, and 3250 median genes per cell. Raw data for single-cell sequencing datasets have been deposited on the European Genome–phenome Archive ( https://ega-archive.org/studies/EGAS00001006283 ) ( Kwok et al. 2023 ).
2.1.4 Dataset 4
The fourth dataset includes a scRNA-seq of human MAIT cells ( Garner et al. 2023 ). We used a subset of the original dataset, comprising five channels of a Chromium Next GEM Chip K. MAIT cells from three donors were either left unstimulated or activated with a TCR, cytokine, or dual TCR + cytokine stimulus. Cells from each donor-condition combination were labelled with TotalSeq-C hashtag antibodies (12 total), pooled, and split across the eight channels of the Chromium Chip ( Garner et al. 2023 ). The Cell Ranger (v7.0.1) count pipeline was used to process FASTQ files for gene expression. Next, hashtag oligo demultiplexing (HTO) and the extraction of singlets, doublets, and negative cells from multiplexing experiments were performed using the HTODemux algorithm ( Stoeckius et al. 2018 ) implemented in the Seurat tool ( Hao et al. 2024 ). We chose to utilize this dataset to demonstrate how interindividual DCE patterns can enhance cell hashing demultiplexing outcomes. The analysis detected up to 16 251 cells per pool, 53 413 mean reads per cell, and 3525 median genes per cell. Raw data for single-cell sequencing datasets have been deposited on the Gene Expression Omnibus (GEO) database ( https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE194187 and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE194189 ) ( Garner et al. 2023 ).
2.1.5 Dataset 5
The fifth dataset includes ventral and dorsal tiers of the substantia nigra (SN) and the cortex (middle frontal gyrus) from five healthy donors ( Agarwal et al. 2020 ). The total number of samples is 12, which includes two replicate samples of SN. The sample libraries contain genes that vary by nuclei from 607 to 3364, and mean Reads that range from 18 377 to 59 513 in both regions ( Agarwal et al. 2020 ). To obtain processed single-nuclei RNA-sequencing matrices, use the accession code https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE140231 from the Gene Expression Omnibus. This dataset was used to show that our method can be applied to nonimmune cells.
2.2 Quantification and gene expression analysis
Sequencing data was processed by Cell Ranger (v7.0) pipelines to create a feature barcoding and gene expression library. The filtered gene–cell matrix generated by the cell ranger was converted from an HDF5 gene–cell matrix to a gene–cell count matrix using the cell ranger mat2csv command provided by 10× genomics. We applied the scater package to filter out single-cell profiles that were outliers for any metrics, as they are considered low-quality libraries ( McCarthy et al. 2017 ). The SCRAN package was applied to detect and remove doublets using expression profiles as described in Dahlin et al. (2018) . Cell type annotation on scRNA-seq data was performed using the scQCEA tool ( Nassiri et al. 2023 ).
2.3 Gene expression recovery
Single-cell RNA-sequencing studies only sequence a small portion of the transcripts present in each cell. This leads to inaccurate quantification of genes with low or moderate expression levels. We used expression recovery methods to provide accurate expression estimates for all genes in order to address this challenge ( Huang et al. 2018 ). The SAVER method was our preference because it recovers the relationship between two marker genes, which we are aware do not correlate ( Huang et al. 2018 ).
2.4 Identification of interindividual variation in gene expression
We introduce an optimized statistical method to detect DCE patterns using single-cell data from two individuals ( McKenzie et al. 2016 ).
We use cellsnp-lite (v1.2.2) ( Huang and Huang 2021 ) and vireo (vireoSNP/0.3.2) ( Huang et al. 2019 ) to infer genetic variants from scRNA-seq data in the first step. Gene–cell count matrices for all possible pairs of individuals are generated using vireo’s best-proposed donor for each cell (or alternative resources, including the use of HTO). The SAVER ( Huang et al. 2018 ) tool is used to transform the gene–cell count matrix per pair of individuals (see section 1.2). The glmnet R package ( Engebretsen and Bohlin 2019 ) is used to select representative gene subsets in the expression profile for each pair of individuals. The calculation of correlation coefficients ( r ) for all possible pairs of selected genes is done using lasso. Interindividual differential gene correlation analysis (IDCA) was conducted for two donors ( D 1 and D 2 ) and genes ( G 1 and G 2 ) using correlation coefficients as described below.
2.5 Cell assignment to donors using a bivariate mixture model
We use a bivariate mixture model for uncovering correlation classes (donors) for each pair of differentially correlated genes. We reconstruct the identity of each cell by fitting a mixture model. The bivariate Gaussian mixture model is used to measure the joint variability between the expression profile of an indicated cell and two donors (donor label proposed by genetic-based demultiplexing) using maximum posterior probability. We applied flexmix R package( Grun and Leisch 2008 ) to perform mixtures of regression models using the Expectation Maximization algorithm and model-based clustering.
In order to reconstruct the sample identity of each cell, a mixture model is fitted using the co-expression patterns of the top DCE genes. A mixture model is utilized to predict a cluster of cells that are not labelled and to reconstruct their sample identity. We consider all possible donor pairs for an indicated cell. This procedure is repeated for the top 10 genes that are differentially co-expressed. We verify that the cell is from a donor if we are able to assign it to the donor most of the time (number of assignments to an indicated donor equal or greater than the total number of pairs of donors minus 1). Reconstructing the sample identification of cells can be done using more liberal thresholds, such as the total number of pairs of donors minus 2.
2.6 Calling genotypes from bulk RNA-sequencing data
Bulk RNA-seq profiles of 30 samples were used for genetic variant calling using GATK (Genome Analysis Toolkit) ( Deelen et al. 2015 ). Our first step was to align the bulk RNA-seq reads to the hg38 reference genome using the Burrows-Wheeler Alignment tool ( Li and Durbin 2009 ). Next, we used GATK’s HaplotypeCaller tool to find genetic variants in the aligned reads ( Van der Auwera et al. 2013 ). Using BCFtools, we removed false positives and low-quality variants from the variants after calling them ( Li 2011 ). Bulk RNA-Seq genotypes were inputted into the demuxlet tool with a posterior probability (PRB = 1) of singlet assignment ( Kang et al. 2018 ).
2.7 Calling genotypes from scRNA-sequencing data
The first step in assigning genetic donors to samples is to perform SNP genotyping using CellSNP-Lite (v1.2.3) ( Huang and Huang 2021 ) in a given data set. We followed the cellSNP-lite manual’s recommended default parameters. Using the cell data from cellSNP-Lite as input, we demultiplexed using vireoSNP (v0.5.8) ( Huang et al. 2019 ).
2.8 Data presentation
The pathway enrichment analysis ( Wu et al. 2021 ) of interindividual DCE genes ( McKenzie et al. 2016 ) was performed using R packages. Box plots and dot plots were generated using ggpubr (v0.2) and customizing ggplot2 ( Almeida et al. 2018 ).
2.9 Implementation
EAD workflow is available at https://isarnassiri.github.io/scDIV/ as an R package called scDIV (acronym for Single-Cell RNA-sequencing data Demultiplexing using Interindividual Variations). Our implementation with variable selection and proper data structures has made the EAD computationally efficient and can be run on a laptop with 16 Gb of memory and two 3.5-GHz CPUs. To run the tool on multiple servers simultaneously for large datasets, users should use a shell script. The package website has documentation that includes examples.
We developed a generic five-step workflow for demultiplexing scRNA-seq data using interindividual variation in gene expression ( Fig. 1 ). First, we infer genetic variants from scRNA-seq data ( Huang and Huang 2021 ) and demultiplex pooled samples ( Fig. 1a ). Gene–cell count matrices are generated for all possible pairs of individuals by utilizing the best-proposed donor for each cell in the previous step. The gene–cell count matrix is transformed by an expression recovery method per pair of individuals to provide precise gene expression values for all genes per cell ( Fig. 1b ) ( Huang et al. 2018 ). We apply lasso (least absolute shrinkage and selection operator) to find compact and representative gene subsets in the expression profiles to improve the accuracy and reduce the redundant downstream number of computations steps ( Fig. 1c ) ( Nassiri and McCall 2018 , Yang et al. 2021 ). Next, we performed an individual-specific co-expression analysis searching for altered co-expression patterns of gene pairs interindividual ( Fig. 1d ) (see section 2). A mixture distribution of correlation classes for each pair of differentially correlated genes is used to fit a mixture model. We reconstruct the identity of each cell based on the similarity of their gene expression with donor-specific clusters using the mixture model ( Fig. 1e ). Cells are assigned if expression-aware and genetic-based demultiplexing propose the same best singlet ( Fig. 2 and Supplementary Fig. 1 ). In this way, the combined method can achieve greater accuracy than the SNP-based or barcode-based methods by selecting the element-wise maximum of the two demultiplexing results.

An example of EAD is to assign an indicated cell to one of seven individuals using a mixture model. Genetic-based demultiplexing already suggested that the cell most likely belongs to donor-0 using the partial genotypic data from state of the individuals in a pool of donors. Now, using the gene expression profiles, we want to check the best-guess assignment obtained from genetic-based demultiplexing. (a) We select the top first pair of DCE genes per donor-0 compared to other donors. The expression pattern of a pair of DCE genes is used to create distinct clusters of cells across individuals. (b) We use a mixture model to predict a cluster of unlabelled cells and reconstruct their sample identity. (c) For an indicated cell (TACTCATCAGCTTCGG-1), we consider all possibilities and most of the time the cell is assigned to donor 0. We confirm that the cell belongs to donor 0 if we successfully assign it to donor 0 for an equal or greater number of pairs of donors, minus 1.
There are multiple methods that can be used to label cells or nuclei with antibodies-based oligonucleotides ( Stoeckius et al. 2018 , Gaublomme et al. 2019 ). By sequencing the cells’ RNA molecules with HTOs, a matrix of count for HTOs per droplet is produced and utilized for demultiplexing of the sample pools. Barcode-based multiplexing methods may be used to combine cells from the same donor and different stimulation conditions into sample pools ( Garner et al. 2023 ). In these circumstances, the genetic-based demultiplexing cannot differentiate between two samples that came from the same donor but had different stimulation conditions. Our EAD workflow uses alternative methods like HTODemux ( Stoeckius et al. 2018 ) to estimate gene expression for each pair of individuals ( Fig. 1b ) in these cases. In order to evaluate EAD, we utilize various datasets that employ diverse methods to label cells in scRNA-seq data for each sequencing donor. These approaches involve sample indices, genetic variation derived from bulk RNA-sequencing profiles of the same samples, and HTOs.
3.1 Model validation using synthetic sample pools
Our initial assessment of EAD was based on synthetic sample pools from eight MM patients ( Fairfax et al. 2020 ). To perform a preliminary evaluation of EAD, we mixed intact donor cells, and refer to them as the ‘synthetic sample pool’. A unique identification number was given to every cell in the synthetic sample pool to reveal its true origin. By using mixed cells with known donor identities, we were able to evaluate the performance of demultiplexing algorithms without introducing errors from the ground truth itself.
Peripheral blood samples were obtained from patients with MM who were treated with immune checkpoint blockade ( Fairfax et al. 2020 ). Samples were collected both before and after the initial treatment cycle. PBMCs were used to isolate monocyte and T cells. Monocyte and T cells were mixed in suspension and the Chromium 10× system was utilized to process the single-cell transcriptome. During library preparation for scRNA sequencing, libraries were tagged using unique indices per donor ( Supplementary Table 1 ). Each sequencing run involved pooling (multiplexing) multiple libraries and sequencing them together ( Fairfax et al. 2020 ). The sequencing run concluded with demultiplexing, and the reads produced were separated into different FASTQ files according to donor indices.
Sorting a pile of laundry (sequencing reads) with a tag (donor index) that identifies the owner (donor) is an analogy to demultiplexing in sequencing. This is not in line with the definition of single-cell sample demultiplexing in this article. In this article, demultiplexing is similar to sorting a pile of laundry (sequencing reads) without tags for each item, by incorporating other indicators like size or personal clothing style (genetic variation) ( Howitt et al. 2023 ).
We demultiplex monocyte and T cells separately in each synthetic sample pool ( Fig. 3a and b ). In the original paper, subsetting was performed to select T cells expressing CD8A, CD8B, and CD3D, and monocytes expressing CD14. We applied further subsetting to eliminate heterogeneous cell populations, including monocytes expressing CD3D, CD3E, CD3G, CD8E, or CD19, and T cells expressing CD14 or CD19.

(a) The results of quality control and the number of called variants for two single-cell sample pools made of 16 samples. (b) Example of top DCE genes (EEF1A and RPS4X) in PBMCs, distinguishes clusters of cells per individual. Pairwise correlations can be visualized as a network. The result shows that EEF1A and RPS4X display co-expression only in donor 1 (red arrow), which could not have been detected based on all individual cells or donor 2. (c) The results of ontology gene set enrichment analysis show a significant association of interindividual DCE genes with the regulation of metabolic processes. The enrichment analysis of cellular components in the dot plot shows associations with mitochondria, ribosomes, cellular macromolecular super complexes, or organelles related to metabolism. (d) Comparison between the methods in terms of prevalence (abbreviations: EAD, expression-aware demultiplexing; GD, genetic-based demultiplexing; mixed, a combination of ED and GD results). According to prevalence, a combination (mixed) of GD and EAD leads to better results for Monocytes and T cells. The colour density reflects the continuous data range to compare values. Lower values are shown to be the most profitable. The range of prevalence values is between 0 and 1.0.
We compared the performance of overall demultiplexing matrices generated based on genetic-based (GD), EAD, and mixed methods for demultiplexing results against that from the known sample labels ( Fig. 3d and Supplementary Table 2 ). We accept the best singlet proposed by GD for unassigned cells if it is consistent with the donor proposed by EAD and call it mixed demultiplexing ( Huang et al. 2019 ).
To calculate the accuracy and evaluate the results, we define true positive (TP) as a donor assignment result that correctly assigns the donor according to the known label. The term false positive refers to a donor assignment result that incorrectly assigns the donor based on the known label. True negative (TN) is characterized by an unassigned donor where the best guess incorrectly suggests a particular donor based on the known label. A false negative is defined as an unassigned donor result that correctly suggests a particular donor based on the known label.
The results showed that the combination of GD and EAD results improves the prevalence and harmonic mean of precision and sensitivity (F1 score) ( Fig. 3d and Supplementary Table 2 ). The accuracy of the two approaches is comparable, and mixed results improve accuracy ( Supplementary Table 2 ).
Enrichment analysis revealed that genes that are differentially co-expressed between individuals are strongly linked to the regulation of metabolic processes and represent metabolic differences between individuals ( Fig. 3c ). Therefore, our method to detect interindividual variation in gene expression could be applied to provide insight into challenges presented by interindividual differences in the responses to nutrition and obesity, cardiovascular and endocrine research, and comprehending the alterations that occur with age and the ensuing neurodegenerative conditions ( Manach et al. 2017 , Lotta et al. 2021 , Johansen et al. 2023 ). Figure 3b shows an example of top DCE genes ( EEF1A and RPS4X ) associated with memory decline in normal aging and Alzheimer’s disease, which distinguishes clusters of cells per individual ( Beckelman et al. 2016a , b ). Only donor 1 showed co-expression of EEF1A and RPS4X , which was not present in all cells or donor 2. Changes in interindividual co-expression relationships could have an impact on the regulation of downstream markers whose expression pattern influences entire biological pathways.
We found that 939 cells were not assigned to any donor. The location of detected unassigned cells on the knee plot showed aggregation after the inflection point ( Fig. 4a and b ). In addition, unassigned cells were mostly found in the bottom-left corner of the quantification plot, as shown in Fig. 4 . It means filtering out nonrelevant cells can improve the accuracy of demultiplexing, especially for samples with a low number of cells. In general, the UMAP projection and number of genes per cell plots showed that unassigned cells in EAD tend to be assigned to regions with low number of genes per cell (Wilcoxon test P < .001) ( Fig. 4 ).

(a, b) The knee, quantification, and UMAP plots show the location of unassigned cells in two sample pools. We ensure that the background red-coloured cells appear on top by dividing the points into different layers and plotting the red points after the black points ( Nassiri et al. 2023 ). Number of detected gene plots shows an association between cell assignment and the number of genes per cell. (c) Boxplots show a significant difference in the mean number of genes per cell across the classes of assigned and unassigned cells using EAD.
We were able to control confounding factors by removing single-cell profiles that did not fit any metric and in vitro separating T cells and monocyte cells. This allowed us to show that patterns of differential gene co-expression across individuals are the consequence of interindividual gene expression differences and no other factors such as cell type ( Fig. 3b ).
Synthetic sample pools are valuable for evaluating scRNA-seq demultiplexing approaches, but they have some shortcomings to consider. The complexity of real pooled single-cell RNA-seq data systems may not be fully captured by synthetic sample pools. Furthermore, a synthetic sample pool may not cover the full range of technical variations that can happen during actual sample pooling. If the synthetic sample pool does not fully replicate real-world scenarios, the observed patterns could reflect the limitations of the synthetic model.
We further evaluate the effectiveness of EAD using several heterogeneous cell populations in the following sections.
In this model, both treatment group and sex might be expected to alter the immune transcriptome. Therefore, we stratified individuals based on the type of treatment and sex as confounding variables to consider their influence on DCE patterns ( Bongen et al. 2019 ). We found no consensus interindividual DCE patterns for matched pairs based on sex (e.g. Male and Male) or type of treatment (e.g. Pembro and Pembro) ( Fig. 5a ).

(a) Confounding variables influence the variation in gene expression between individuals. We stratified melanoma patients based on the type of treatment with immune checkpoint blockades (Ipilimumab + Nivolumab (Ipi + Nivo) or Pembrolizumab (Pembro)) and sex (male or female). If these factors cause DCE patterns among individuals, we expect the appearance and disappearance of the top first examples of DCE genes across classes. We found no such an accumulation. For example, the expression of EEF1A1 and RPS4X represent individual 7 and we do not see a similar DCE pattern for matched pairs of individuals based on treatment or sex (e.g. Male and Female). (b) The results of quality control and the number of called variants per cell for a single-cell pool sample made of 6 isogenic mice. We applied the scater package to filter out single-cell profiles that were outliers for any metrics, as they are considered low-quality libraries ( McCarthy et al. 2017 ). (c) A pool of six samples with known cell labels from isogenic individuals was used as input for an EAD workflow. DCE patterns across pairs of donors could not distinguish interindividual differences in the gene expression including genes related to the metabolic pathway (e.g. ugp2 and Miox genes), and we only see differences related to the treatment (e.g. sample 4 versus sample 2).
We evaluated the effectiveness of the expression-aware approach in demultiplexing cells from isogenic individuals. We pooled cells with a known label from six isogenic mice exposed to a topical TLR7 agonist Imiquimod to induce a systemic lupus erythematosus-like phenotype or vehicle control and tried to perform demultiplexing using the expression-aware approach ( Fig. 5b and c ). Sample indices were used to label cells per sequencing sample/donor in this dataset ( Supplementary Table 3 ).
Interindividual differences were not discernible through DCE patterns across classes of isogenic samples ( Fig. 5c ). The results indicate that it is challenging to identify variations in metabolic pathways for sample pools with very little genetic diversity among donors. Therefore, demonstrating the efficacy of expression-aware approach for demultiplexing these pooled samples was not possible.
The assumption that genetic and environmental differences lead to DCE patterns between individuals is supported by the absence of individual-specific signatures in isogenic laboratory mice ( Figs 3 and 5c ). These variables can be controlled to resolve interindividual differences in gene co-expression related to metabolic pathways ( Fig. 5c ). In conclusion, nonisogenic individual samples are suitable to be pooled for demultiplexing scRNA-seq data using interindividual variations in genetic and gene expression.
3.2 Application to real pooled single-cell RNA-seq
We considered more challenging scenarios involving five pooled sample (batch) and 30 donors (six donors per batch) to test if the method would work well with more heterogeneous cell populations ( Fig. 6a and b ) ( Kwok et al. 2023 ). From every donor, the nuclei of CD34+ circulating hematopoietic stem and progenitor cells (HSPCs) were extracted. Six samples with equal numbers of cells were combined to form a single pool. Each pool was then subjected to cell lysis and nuclear extraction. RNA libraries were sequenced after loading each pool of nuclei across four channels of the 10× genomics lane on the chip ( Kwok et al. 2023 ). In this context, a 10× lane would represent a batch loaded onto the chip. To differentiate this dataset from other datasets, we use batch and 10× lane interchangeably.

Demultiplexing with genotype reference (Demuxlet) provides a source of ground truth for benchmarking the performance of demultiplexing algorithms. (a and b) The results of quality control including the percentage of filter-out cells as a doublet and the number of called variants per cell for five single-cell pool samples (10× lanes) made of 23 sepsis and seven healthy individuals. (c) Percentage of correct (TP and TN), and incorrect assigned cells (FP and FN) using EAD, GD demultiplexing (vireo), and the combination of genetic-based and expression-aware approaches (Mixed). The outcomes are given for each 10× lane and for all 10× lanes. (d) The confusion matrices were utilized to generate the key metrics and give a comprehensive assessment of demultiplexing methods that do not require a reference genome. Abbreviations: EAD, expression-aware demultiplexing; GD, genetic-based demultiplexing; Mixed-D, mixed demultiplexing; NPV, negative predictive value; CSI, critical success index.
10× lanes in the scRNA-seq experiment of HSPCs included samples from healthy donors, sepsis donors, and donors with both sepsis symptoms and COVID-19 infection ( Kwok et al. 2023 ).
There are four methods that can demultiplex pooled single-cell RNA-seq without relying on reference SNP genotypes: vireo ( Huang et al. 2019 ), scSplit ( Xu et al. 2019 ), Freemuxlet ( Kang et al. 2018 ), and Souporcell ( Heaton et al. 2020 ). According to available benchmark studies, vireo outperforms other tools when it comes to demultiplexing pooled single-cell RNA-seq data without genotype reference ( Neavin et al. 2024 , Cardiello et al. 2023 ). Furthermore, vireo outputs have a high level of consistency with the outcomes of other tools ( Neavin et al. 2024 , Cardiello et al. 2023 ). Therefore, we opted for vireo as the genotype-free demultiplexer for pooled single-cell RNA-seq.
We utilized the Demuxlet tool and reference genotypes derived from bulk RNA-sequencing profiles of the same samples to determine the donor identity of every singlet ( Kang et al. 2018 ). SNPs in each individual’s genome were used in demultiplexing with genotype reference (Demuxlet) to identify individual donor identities per cell with high accuracy. This allows for a precise assessment of how other algorithms compare to this ground truth. The performance of EAD and GD (GD—vireo) ( Huang et al. 2019 ) methods for demultiplexing, which do not require a reference SNP genotypes, was evaluated in the next step using Demuxlet results.
The TP rate is determined by the fraction of cells with the corresponding genetic assignment (Demuxlet) for each possible EAD, Mixed-D or GD assignment. The fraction of cells with EAD, Mixed-D or GD assignments that are different from their genetic assignments is called the false positive rate (FP). The false negative rate (FN) is the proportion of cells that have genetic assignment and EAD, Mixed-D or GD un-assignment, and the best guess of EAD, Mixed-D or GD accurately indicates a specific donor. The TN rate is the proportion of cells that have genetic un-assignment and EAD, Mixed-D or GD un-assignment.
We used the sensitivity and F-score to evaluate performance as our main metric. The proportion of positive cases that are correctly identified by the evaluation method is represented by sensitivity. The F-score refers to the harmonic mean of precision and recall, which can range from zero to one. A higher F-score indicates superior performance.
Six samples from genetically distinct donors were represented by cells in each of the five 10× lanes in the data set. The data from each 10× lane was subjected to demultiplexing methods. A cell could be assigned to singlet, a singlet that corresponds to one of the six unique samples or unassigned by each demultiplexing method. Misclassifying true singlets as doublets can be a more significant error source in downstream analysis than misclassifying true doublets ( Wolock et al. 2019 , Howitt et al. 2023 ). Therefore, for vireo, we treat the doublets as unassigned and do not attempt to reassign them through mixed demultiplexing.
We next evaluated the donor assignments made by expression-aware, vireo, and Mixed-D methods against the genetic assignments from Demuxlet that were considered to be ‘ground truth’. Figure 6 exhibits mosaic plots of the confusion matrix for demultiplexing methods, divided by lanes. In all 10× lanes, Mixed-D had an increased TP and FP rate, but a decreased FN rate. Mixed-D and vireo demonstrated a consistent correct assignment (TP+TN) in all batches ( Fig. 6c ).
To quantitatively evaluate the performance of demultiplexing methods, we calculated the key metrics derived from the confusion matrix ( Fig. 6d ). Mixed-D had the most significant mean sensitivity, prevalence, and negative predictive value among all methods for every 10× lane. All 10× lanes displayed consistent F-scores and accuracy for Mixed-D and vireo ( Fig. 6d ).
While expression-based methods could not significantly improve the percentage of correct assignment (TP+TN) in 10× lanes of this heterogeneous dataset, Mixed-D could provide more confirmation of the vireo results and improve the accuracy of assignments ( Fig. 6c ).
Despite the diversity of cells, the top interindividual DCE genes produced a distinct cluster of cells per individual ( Fig. 7a ). Consistent with previous results, ontology gene set enrichment analysis for DCE genes showed a significant association with the regulation of metabolic processes ( Supplementary Fig. 2 ). While the top interindividual DCE genes are related to metabolic functions or organelles, the breakdown of co-expression patterns can represent context-specific pathognomonic signatures. An illustrative example occurs in the differential co-expression of genes specific to sepsis in the context of COVID-19 infection such as NEDD9 ( Rizzo and Yuan 2022 ), BACE2 ( Tang et al. 2022 ), and DHX30 ( Apostolidou et al. 2021 ). The application of interindividual DCE genes allows for the differentiation of subtle pathology-induced patterns in a context-specific manner.

(a) Example of the top interindividual DCE genes provide a distinct cluster of cells per individual. (b) There are no significant differences in chromatin accessibility across states observed in the case of the top DCE gene. (c) Association between cell assignment and the number of genes per cell. Some unassigned cells show a low number of genes per cell, which means filtering out unassigned cells can improve the accuracy of cell calls.
We investigated the potential for identifying interindividual single-cell expression variability using multiome ATAC. The results showed that chromatin accessibility within the same cell does not provide a similar explicit model for co-expression relationships between molecules at single-cell level ( Fig. 7b ).
The tSNE projection plot revealed that unassigned cells were accumulating in regions with low numbers of genes per cell ( Fig. 7c ). Therefore, the accuracy of cell identification can be improved by removing cells which have very few genes per cell. Functional enrichment can be combined with unassigned cells to maintain statistical power for pooled scRNA-seq data analysis as an alternative solution ( Fairfax et al. 2020 ).
The authors in the original paper ( Kwok et al. 2023 ) used the combined gene expression profiles to group cells according to their gene expression and chromatin data (seven clusters labelled as C1–7). The differences in gene expression between the clusters of the dataset are likely driven by their skew towards different lineages (e.g. C4: lymphocyte progenitors, C5: emergency granulopoiesis, and C7: normal granulopoiesis). We did not observe strong correlation between the proportion of cells in each cluster and number of unassigned cells including singlets and doublets.
These results demonstrate the potential utility of demultiplexing pooled single-cell RNA-sequencing samples using interindividual variation in gene expression in various biological models.
3.3 Integrate the demultiplexing results from expression-based and barcode-based methods
Multiple methods are available for labelling cells or nuclei with oligonucleotide-barcoded antibodies ( Stoeckius et al. 2018 , Gaublomme et al. 2019 ). A matrix of counts per HTOs per droplet is created by sequencing the cells’ RNA molecules with HTOs.
Using a scRNA-seq dataset of stimulated human MAIT cells ( Garner et al. 2023 ), we present a demonstration of how expression-aware and barcode-based demultiplexing methods can complement each other ( Fig. 8 ). We used a subset of the original dataset, comprising five channels of a Chromium Next GEM Chip K. MAIT cells from three donors were either left unstimulated or activated with a TCR, cytokine, or dual TCR+cytokine stimulus. Cells from each donor-condition combination were labelled with TotalSeq-C hashtag antibodies (12 total), pooled, and split across the eight channels of the Chromium Chip ( Garner et al. 2023 ).

(a) A distinct cluster of cells from the same donor but a different stimulation condition is present in an example of the top DCE genes. (b) Summaries of cell hashing demultiplexing results showing the number of singlets called per 10× lane along with the percentage of doublets and negative cells (both filtered out). (c–e) Evaluating the impact of different thresholds for EA sample demultiplexing on accuracy, percentage of assigned cells, and percentage of genetic-based demultiplexing doublets in EAD assignments by specifying various thresholds ( T ).
The dataset contained three donors and four conditions per donor per 10× lane. Genetic-based demultiplexing (vireo) was unable to differentiate two samples from the same donor but different stimulation conditions ( Howitt et al. 2023 ). Therefore, we were unable to produce results for mixed demultiplexing generated using genetic-based and expression-aware methods on this dataset. To estimate gene expression for each pair of samples ( Fig. 1a ) in our EAD workflow, we utilized the output of Seurat HTODemux ( Stoeckius et al. 2018 ) instead of vireo. We chose HTODemux as our preferred tool because it is one of the top three most effective methods and performs fairly well on different types of datasets with minor differences between its output and other tools ( Howitt et al. 2023 ). The EAD technique was capable of separating all 12 samples per 10× lane, including those that were derived from the same donor but different stimulation conditions ( Fig. 8a ). Several methods were used for demultiplexing of cell hashing data: BFFcluster ( Boggy et al. 2022 ), BFFraw ( Boggy et al. 2022 ), GMM-Demux ( Xin et al. 2020 ), MULTI-seq ( McGinnis et al. 2019 ), HTODemux ( Stoeckius et al. 2018 ), demuxEM ( Gaublomme et al. 2019 ), and DropletUtils hashedDrops ( Griffiths et al. 2018 , Boggy et al. 2022 ). Each algorithm assigns cells as singlets (one hashtag antibody detected), doublets (two or more hashtag antibodies detected), or negative (no hashtag antibody detected) ( Fig. 8b ). We treat the doublets and negatives as unassigned and do not attempt to reassign them through EA demultiplexing. To assess the performance of the EAD method, we compared its assignments to singlet calls from the six HTO demultiplexing tools. To calculate accuracy, we compared EAD classifications to HTO algorithm classifications, and then divided the number of concordant classifications by the total number of classifications.
By using 70 000 singlets from five 10× lanes, EAD was able to assign the 90% (62 682/70 000) of cells with an average of 93.2% accuracy ( Fig. 8c ). EAD’s results were similar across all HTO tools, but there was a slight decline for DropletUtils and BFFcluster ( Fig. 8c ). BFFcluster and DropletUtils determine doublets by thresholding barcode counts, resulting in the algorithm’s performance being highly dependent on the correct selection of this parameter ( Boggy et al. 2022 ). The default parameters recommended by cellhashR were used in this study ( Boggy et al. 2022 ). Adjusting the parameters could potentially yield more consistent results. It is probable that there are problematic singlets for the expression-aware method as it did not consistently assign 6.8% of singlets. Our next step was to delve deeper into the cases where demultiplexing with hashtag oligos and EAD is not in accordance.
Figures 1 and 2c explain that EAD takes into account the assignments for an indicated cell across all possible donor pairs. A cell that is successfully assigned to a donor for an equal or greater number of donor pairs minus 1 (0.9 of all the donor pairs that are possible), is confirmed as belonging to the donor by EAD ( Fig. 2c ). It is possible to utilize either more liberal or rigid thresholds ( T ) for EA sample demultiplexing to evaluate its impact on accuracy ( Fig. 8c–e ). According to the results, EAD produces balanced output for accuracy and the percentage of assigned cells when T is ≥0.8. Increasing the threshold to 1 could lead to an increase in accuracy to 99.9%, but at the expense of decreasing the percentage of assigned cells to 46% on average ( Fig. 8d ). Since there are three genetically distinct donors per 10× lane, we can use SNP-based multiplexing to estimate cells with mismatched SNP profiles (doublets). According to the findings, elevating the threshold for EA sample demultiplexing consistently decreases the number of detected doublets by SNP-based demultiplexing (vireo) ( Fig. 8e ). This trend remains the same even with different thresholds that yield similar percentages of assigned cells (e.g. 0.1–0.6) ( Fig. 8d and e ). By using the EAD with a high threshold for sample demultiplexing (e.g. T = 1), it is possible to obtain reliable results for cells with sufficient barcode counts for accurate classification, as evidenced by this result ( Fig. 8c and d ).
The 1968 discordant assignments occurred totally among singlet calls from six HTO demultiplexing tools implemented in cellhashR ( Boggy et al. 2022 ). cellhashR defines discordant cells when there is at least one hashing algorithm that produces a different result. The majority of discordant results are caused by the algorithms having slightly different thresholds, as demonstrated in the previous study ( Boggy et al. 2022 ). To offer a demonstration of an EAD application that can be utilized in conjunction with existing barcode-based and SNP-based multiplexing methods, we attempted to assign cells using Mixed-D for each singlet that was discordant across HTO demultiplexing tools. When studying a biological phenomenon, it is typical to use several complementary techniques (such as Western blotting and qPCR) to enhance the reliability of findings. If the results of different techniques are similar, there is a boost in confidence in the overall conclusion. By using complementary techniques, we can analyse cell assignments from various angles, such as barcode-based and expression-based, and the convergence of results indicates the reliability of the assignments. Mixed-D only approved singlets proposed by EAD ( T = 1) that were identical to the proposed donor by at least three tools in cellhashR output. The mixed-D was able to save 50.4% (993/1968) of discordant singlets (Min. 40.95%, Median 50.00%, Max. 59.05%) and it improved the HTO demultiplexing results by an average of 1.4% (993/70 000) (Min. 1.25%, Median 1.33%, Max. 1.74%). Since three HTO demultiplexing tools have previously verified the Mixed-D assignments and we employ a threshold equal to 1 for EAD, we can anticipate secure assignments for the discordant cells.
The cost of single-cell sequencing is influenced by various factors, such as the specific technology employed, the number of cells sequenced, the sequencing depth, and the provider. In most cases, the cost of scRNA-seq is ∼£1–5 per cell. Considering the dataset of blood and liver MAIT cells as an example, mixed demultiplexing can reduce the number of discordant cells (from 1968 to 975) and save around £3000. Expression-based demultiplexing is a promising complementary approach to identifying individual samples in a pooled sequencing experiment. By combining the results of both methods, we can obtain a more precise picture of the actual demultiplexing results.
Interindividual DCE analysis within the same cell type can be used to identify cells from the same donor in different activation states, as indicated by the results from the MAIT cell dataset ( Garner et al. 2023 ). The use of interindividual DCE analysis for scRNA-seq is not limited to EAD. It can potentially be applied to discover small changes in gene expression that are linked to various conditions or cell states. Researchers who want to study the transcriptional heterogeneity of cell states within the same cell type can benefit from this technique ( Garner et al. 2023 ). These applications are covered in more detail in the following section.
3.4 Leveraging interindividual variation in gene expression for precision therapeutic strategies
In this section, we show that it is possible to identify genes that are differentially co-expressed for each pair of donors for nonimmune cells. Using DCE pairs between individuals allows for EAD. Furthermore, we examine examples of variations in co-expressed gene patterns among various individuals that can shed light on the molecular mechanisms underlying differences in therapeutic response.
To demonstrate that our method can be utilized for nonimmune cells, we used single-nuclei transcriptome profiles of the SN and cortex (middle frontal gyrus) (in total 12 samples, including two SN replicates) ( Agarwal et al. 2020 ). The analysis of cell-type enrichment showed that there were five different cell populations in each sample of the SN and six different cell populations in the cortex (e.g. astrocytes) ( Agarwal et al. 2020 ). We were able to identify distinct clusters of cells across individuals based on the expression pattern of DCE genes despite cell heterogeneity ( Fig. 9 ). In at least one pair of samples, we found 3207 genes with DCE (FDR < 1 −10 ), with 568 of them being detected in only one pair of samples for the indicated region. We did not observe DCE patterns between replicated samples (FDR < 1 −4 ). Our achievement of finding representative DCE genes for each pair of donors allows us to use EAD for this data type.

A few examples of the many interindividual DCE genes that have been identified in the substantia nigra and cortex. These genes play important roles in a variety of neurological processes, and their dysregulation can contribute to the development of neurological disorders. More investigation is required to fully comprehend the roles of these genes in the brain and their potential as therapeutic targets for neurological disorders. (a, b) Examples of interindividual DCE pattern in the substantia nigra and cortex. (c) The t-SNE project of transcriptionally and functionally distinct clusters, highlighting microglia and neuron cell type groups, is presented. Pink/Red cells have passed the threshold of cell type enrichment ( Nassiri et al. 2023 ). (d) Examples of DCE patterns between cell types in the substantia nigra and cortex. (e) A pair of genes that exhibit differential co-expression but not differential expression. The analyses were carried out by employing single-cell expression profiles across the SN and cortex regions of a donor. In each region, the single-cell expression distributions of CTNNA3 and FAM221A genes are visualized by a violin plot. The expression levels of both genes decrease from SN to cortex, but only CTNNA3 has a significant differential expression. (f) The CTNNA3 and FAM221A gene pairs exhibit DCE patterns in the SN and cortex.
One example is the differential co-expression of MALAT1 and LINGO1 in the cortex of different individuals ( Fig. 9a ). MALAT1 is a long noncoding RNA (lncRNA) that is highly expressed in the brain, particularly in the cortex. MALAT1 has the potential to be a promising therapeutic target for a variety of neurological disorders ( Wang et al. 2022 ). Neuronal survival and axonal growth are regulated by LINGO1 , which is a leucine-rich single transmembrane protein, in the cortex. It has been observed that it promotes the death of neurons and hinders the growth of axons. It is suggested that LINGO1 may be involved in both the normal pruning of neurons during development and the pathological loss of neurons that occurs in neurodegenerative diseases ( Inoue et al. 2007 ). LINGO1 has the potential to be a promising therapeutic target for several neurological disorders. MALAT1 and LINGO1 have been demonstrated to interact and have opposite effects on neuronal survival ( Fan et al. 2018 ). Neuronal survival is promoted by MALAT1 , while neuronal death is promoted by LINGO1 ( Inoue et al. 2007 , Wang et al. 2022 ). The balance between MALAT1 and LINGO1 expression may play a significant role in regulating neuronal survival in the cortex. Further research is needed to fully understand the differential co-expression relationship between upregulation of MALAT1 and downregulation of LINGO1 , as well as its roles in neurological disorders.
Other examples of interindividual DCE pairs include XIST and ARL17B in SN ( Fig. 9b ). The SN expresses XIST , which has been shown to regulate the expression of genes crucial for dopaminergic neuron function ( Wang et al. 2021 ). ARL17B is present in the SN and has been discovered to play a significant role in the survival of dopaminergic neurons ( Reus et al. 2021 ). Parkinson’s disease, which is characterized by the loss of dopaminergic neurons in the SN, has been linked to mutations in XIST and ARL17B ( Wang et al. 2021 ). Our result indicates that there is a settled co-expression relationship between XIST and ARL17B . Further exploration is required to fully comprehend the functional consequence of losing association between XIST and ARL17B expression in SN and their possible contribution to Parkinson’s disease.
The expression patterns of many genes are not directly co-regulated within cell types, but they are differentially co-expressed across cell types, as shown by our results. For instance, the interaction between ARID2 and CNTNAP2 resolves in glial cells of the SN ( Fig. 9c and d ). This demonstrates differential co-expression between cortical neurons and SN glial cell types. The regulation of gene expression and DNA replication by chromatin remodelling is facilitated by ARID2 (AT-rich interaction domain 2) protein ( Kang et al. 2021 ). It is particularly important for glial cells to maintain homeostasis and protect against infection and neurodegeneration. In glial cells, CNTNAP2 (contactin-associated protein 2) is a key component of glial-neuronal communication, neuroprotection, and cognitive function ( Gandhi et al. 2023 , St George-Hyslop et al. 2023 ). The interaction between CNTNAP2 and ARID2 leads to the stabilization of the ARID2 protein ( Moffat et al. 2022 ). ARID2 needs this stabilization to properly regulate gene expression ( Moffat et al. 2022 ). Our results suggest that the co-expression of ARID2 and CNTNAP2 is resolved in the microglia and astrocyte cells in SN ( Fig. 9d ). Further research is needed to fully elucidate the effects of losing the interaction between ARID2 and CNTNAP2 on microglial and astrocyte cell functions in SN.
Our observation revealed a distinct set of genes that represent each brain region and cell-type. In addition, we found examples of interindividual variations in co-expressed gene patterns, which provide insight into the challenges presented by variation in therapeutic response ( Crowell et al. 2020 , Guo et al. 2023 ).
Typical differential gene expression analysis (DiffEx) does not provide a similar explicit model for co-expression relations between molecules at the single-cell level. The focus of DiffEx is on the expression levels of individual genes ( Gaublomme et al. 2019 ). Differential co-expression analysis (DiffCoEx) is focused on discovering genes whose expression levels change significantly under various conditions. DiffCoEx offers insight into the coordinated response of genes and assists us in understanding the underlying biological networks ( McKenzie et al. 2016 , Crowell et al. 2020 ). As a demonstration, we compare DiffEx and DiffCoEx results across SN and cortex regions in a 56-year-old female donor ( Fig. 9e and f ). Differential expression of genes across the SN and cortex was found by the Seurat tool, and 64.8% (5483/8456) of them were also differentially co-expressed (FDR<1 −3 ) ( Hao et al. 2024 ). The example of CTNNA3 and FAM221A genes illustrates the distinction between differential expression and differential co-expression. In both SN and cortex contexts, Fig. 9e illustrates where the CTNNA3 and FAM221A genes are expressed. The expression values of both genes decrease from SN to cortex, but FAM221A is not a significant differentially expressed gene ( P = 1), as shown in Fig. 9e . These genes have significant differential co-expression in the SN and cortex, which cannot be predicted solely based on differential expression relationships ( Fig. 9f ).
The capability of multiplex scRNA-seq samples has attracted research attention to lowering experiment costs and addressing batch effects. Several multiplexing methods and bioinformatics tools have been developed for demultiplexing pooled datasets ( Kang et al. 2018 , Stoeckius et al. 2018 , Gaublomme et al. 2019 , Guo et al. 2019 ). The main approaches include barcode-based ( Boggy et al. 2022 ) and SNP-based sample pool demultiplexing ( Kang et al. 2018 , Huang et al. 2019 ). Compared to other approaches, EAD facilitates feature selection (marker gene selection) for clustering single-cell data by obtaining individual-specific variability ( Ranjan et al. 2021 ). EAD has added benefits to previous methods by improving the accuracy of cell assignments into individual samples, without the need for additional experimental steps ( Nassiri et al. 2023 ).
The accurate assigning of cells to their respective donors can provide valuable insights for researchers into the biological processes and disease mechanisms that vary across individuals. The findings indicate that combining expression-based demultiplexing with SNP-based or barcode-based methods is the most accurate approach for demultiplexing single-cell and single-nuclei sequencing data. The combined demultiplexing results are more accurate due to the fact that they take into account the strengths of methods. For example, cell-multiplexing oligos using cells or nuclei samples can be technically challenging and have limited performance ( Stoeckius et al. 2018 ). On the other hand, the identity of donors (donor-specific information) is not specified by EAD. When cells are assigned to donor groups using EAD, the identity of donors can be specified by incorporating demultiplexing results provided by barcode-based multiplexing assays ( Kim et al. 2022 ).
Each person has a distinct set of SNPs. In the event that two cells from different individuals are combined to form a doublet, the combined SNP profile will appear unbalanced. SNP demultiplexing algorithms have the ability to identify these inconsistencies and label them as potential doublets ( Kang et al. 2018 , Huang et al. 2019 ). SNP demultiplexing for doublet detection is not perfect and can lead to false positives. Factors like sequencing depth and quality, and completeness of reference panels used for SNP calling determine the accuracy of doublet detection ( Kang et al. 2018 ). False positives can be caused by sequencing errors, allelic dropout, batch effects, somatic mutations, and natural variability in gene expression ( Huang et al. 2019 ). While expression-based demultiplexing cannot detect doublets, the combination of SNP and expression demultiplexing can offer more confirmation or reassign doublet predictions. Filtering out cells that are classified as true doublets using tools like SCAN is recommended before reassigning a predicted doublet using Mixed-D ( Dahlin et al. 2018 ).
The validation approach for characterization of co-expression variations is limited by the use of datasets from the 10× Genomics Chromium platform. In general, it is plausible that targeted scRNA-seq is less likely to obtain enough SNPs for SNP-based demultiplexing and has a limited number of genes for DCE analysis. By incorporating other single-cell RNA and protein expression technologies that have been developed, our approach can be easily enhanced.
The application of interindividual DCE analysis of scRNA-seq goes beyond EAD. It can potentially be applied to reveal biological activity that is useful for patient stratification, identifying biomarkers of target engagement, and connecting genomic programs with biological functions ( McKenzie et al. 2016 , Badia-i-Mompel et al. 2023 ). The study of interindividual variation in single-cell RNA-sequencing data is a constantly evolving field that involves continuous methodological advancement and refinement ( Kumasaka et al. 2023 ). Most single-cell studies employ average gene expression profiles across cell types or states of interest, which often obscures differences among individuals that are apparent at single-cell resolution ( Murdock and Tsai 2023 ). Variability between individuals remains an understudied aspect of relationships between molecules at the single-cell level ( Crowell et al. 2020 ). Cell-level mixed models are currently employed for differential state analysis across multiple samples (e.g. donors) and experimental conditions using cell-level measurements ( Crowell et al. 2020 ). The cell-level mixed models generate log-fold changes and the proportion of cells that have expressed a particular gene in each sample or group. These methods greatly underestimate the differences in gene expression between different cell subpopulations, such as those with low expression ( Crowell et al. 2020 , Auerbach et al. 2021 ). The cell-level solution to improve this issue is provided by DCE analysis, which directly models gene expression. In addition, it provides the possibility of modelling related gene regulatory networks and following each pair of DCE genes along the axis of samples. While the interindividual differential co-expression approach identifies how disease signatures vary between clinical subtypes, these changes ultimately might converge on shared signalling pathways that present biomarkers for new precision therapeutic strategies ( Johansen et al. 2023 ).
We anticipate that EAD will be applied to improve the results of SNP-based and barcode-based multiplexing assays, provide insight into challenges presented by variation in therapeutic response, and consider the effect of these stratification strategies on proposed candidate biomarker genes ( Stoeckius et al. 2018 , Li et al. 2020 ).
Isar Nassiri (Conceptualization [lead], Formal analysis [lead], Methodology [lead], Project administration [lead], Resources [lead], Software [lead], Validation [lead], Visualization [lead], Writing—original draft [lead], Writing—review & editing [lead]), Andrew J. Kwok (Data curation [supporting], Resources [supporting], Validation [supporting], Writing—original draft [supporting]), Aneesha Bhandari (Data curation [supporting], Resources [supporting], Validation [supporting], Writing—original draft [supporting]), Katherine R. Bull (Data curation [equal], Resources [equal], Validation [equal], Writing—original draft [equal]), Lucy C. Garner (Data curation [equal], Formal analysis [equal], Resources [equal], Validation [equal], Writing—original draft [equal], Writing—review & editing [equal]), Paul Klenerman (Data curation [equal], Resources [equal], Writing—review & editing [equal]), Caleb Webber (Data curation [equal], Resources [equal], Writing—review & editing [equal]), Laura Parkkinen (Conceptualization [equal], Data curation [equal], Resources [equal], Writing—review & editing [equal]), Angela W. Lee (Data curation [equal], Resources [equal], Writing—original draft [equal]), Yanxia Wu (Conceptualization [equal], Data curation [equal], Investigation [equal], Writing—original draft [supporting], Writing—review & editing [supporting]), Benjamin Fairfax (Data curation [equal], Methodology [supporting], Resources [equal], Writing—original draft [equal], Writing—review & editing [supporting]), Julian C. Knight (Investigation [equal], Resources [equal], Writing—original draft [equal], Writing—review & editing [equal]), David Buck (Conceptualization [equal], Resources [equal], Writing—original draft [equal]), and Paolo Piazza (Funding acquisition [lead], Methodology [equal], Project administration [supporting], Resources [equal], Validation [equal], Writing—original draft [equal], Writing—review & editing [equal])
Supplementary data are available at Bioinformatics Advances online.
None declared.
The research was supported by the Wellcome Trust Core Award [203141/Z/16/Z]. I.N. was supported by the National Institute for Health Research (NIHR) Oxford Health Biomedical Research Centre (BRC). The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health.
Agarwal D , Sandor C , Volpato V et al. A single-cell atlas of the human substantia nigra reveals cell-specific pathways associated with neurological disorders . Nat Commun 2020 ; 11 : 4183 .
Google Scholar
Almeida A , Loy A , Hofmann H. ggplot2 compatible quantile-quantile plots in R . R J 2018 ; 10 : 248 – 61 .
Apostolidou S , Harbauer T , Lasch P et al. Fatal COVID-19 in a child with persistence of SARS-CoV-2 despite extensive multidisciplinary treatment: a case report . Children (Basel) 2021 ; 8 : 564 .
Auerbach BJ , Hu J , Reilly MP et al. Applications of single-cell genomics and computational strategies to study common disease and population-level variation . Genome Res 2021 ; 31 : 1728 – 41 .
Badia-I-Mompel P , Wessels L , Müller-Dott S et al. Gene regulatory network inference in the era of single-cell multi-omics . Nat Rev Genet 2023 ; 24 : 739 – 54 .
Beckelman BC , Day S , Zhou X et al. Dysregulation of elongation factor 1A expression is correlated with synaptic plasticity impairments in Alzheimer’s disease . J Alzheimers Dis 2016a ; 54 : 669 – 78 .
Beckelman BC , Zhou X , Keene CD et al. Impaired eukaryotic elongation factor 1A expression in Alzheimer’s disease . Neurodegener Dis 2016b ; 16 : 39 – 43 .
Boggy GJ , McElfresh GW , Mahyari E et al. BFF and cellhashR: analysis tools for accurate demultiplexing of cell hashing data . Bioinformatics 2022 ; 38 : 2791 – 801 .
Bongen E , Lucian H , Khatri A et al. Sex differences in the blood transcriptome identify robust changes in immune cell proportions with aging and influenza infection . Cell Rep 2019 ; 29 : 1961 – 73.e4 .
Cardiello JF , Joven Araus A , Giatrellis S et al. Evaluation of genetic demultiplexing of single-cell sequencing data from model species . Life Sci Alliance 2023 ; 6 : e202301979 .
Crowell HL , Soneson C , Germain P-L et al. Muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data . Nat Commun 2020 ; 11 : 6077 .
Dahlin JS , Hamey FK , Pijuan-Sala B et al. A single-cell hematopoietic landscape resolves 8 lineage trajectories and defects in kit mutant mice . Blood 2018 ; 131 : e1 – 11 .
Deelen P , Zhernakova DV , de Haan M et al. Calling genotypes from public RNA-sequencing data enables identification of genetic variants that affect gene-expression levels . Genome Med 2015 ; 7 : 30 .
Engebretsen S , Bohlin J. Statistical predictions with glmnet . Clin Epigenetics 2019 ; 11 : 123 .
Fairfax BP , Makino S , Radhakrishnan J et al. Genetics of gene expression in primary immune cells identifies cell type-specific master regulators and roles of HLA alleles . Nat Genet 2012 ; 44 : 502 – 10 .
Fairfax BP , Taylor CA , Watson RA et al. Peripheral CD8+ T cell characteristics associated with durable responses to immune checkpoint blockade in patients with metastatic melanoma . Nat Med 2020 ; 26 : 193 – 9 .
Fan B , Wei Z , Yao X et al. Microenvironment imbalance of spinal cord injury . Cell Transplant 2018 ; 27 : 853 – 66 .
Gandhi T , Canepa CR , Adeyelu TT et al. Neuroanatomical alterations in the CNTNAP2 mouse model of autism spectrum disorder . Brain Sci 2023 ; 13 : 891 .
Garner LC , Amini A , FitzPatrick MEB et al. Single-cell analysis of human MAIT cell transcriptional, functional and clonal diversity . Nat Immunol 2023 ; 24 : 1565 – 78 .
Gaublomme JT , Li B , McCabe C et al. Nuclei multiplexing with barcoded antibodies for single-nucleus genomics . Nat Commun 2019 ; 10 : 2907 .
Griffiths JA , Richard AC , Bach K et al. Detection and removal of barcode swapping in single-cell RNA-seq data . Nat Commun 2018 ; 9 : 2667 .
Grun B , Leisch F. FlexMix version 2: finite mixtures with concomitant variables and varying and constant parameters . J Stat Soft 2008 ; 28 : 1 – 35 .
Guo C , Kong W , Kamimoto K et al. CellTag indexing: genetic barcode-based sample multiplexing for single-cell genomics . Genome Biol 2019 ; 20 : 90 .
Guo MG , Reynolds DL , Ang CE et al. Integrative analyses highlight functional regulatory variants associated with neuropsychiatric diseases . Nat Genet 2023 ; 55 : 1876 – 91 .
Hao Y , Stuart T , Kowalski MH et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis . Nat Biotechnol 2024 ; 42 : 293 – 304 .
Heaton H , Talman AM , Knights A et al. Souporcell: robust clustering of single-cell RNA-seq data by genotype without reference genotypes . Nat Methods 2020 ; 17 : 615 – 20 .
Howitt G , Feng Y , Tobar L et al. Benchmarking single-cell hashtag oligo demultiplexing methods . NAR Genom Bioinform 2023 ; 5 : lqad086 .
Huang M , Wang J , Torre E et al. SAVER: gene expression recovery for single-cell RNA sequencing . Nat Methods 2018 ; 15 : 539 – 42 .
Huang X , Huang Y. Cellsnp-lite: an efficient tool for genotyping single cells . Bioinformatics 2021 ; 37 : 4569 – 71 .
Huang Y , McCarthy DJ , Stegle O. Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference . Genome Biol 2019 ; 20 : 273 .
Inoue H , Lin L , Lee X et al. Inhibition of the leucine-rich repeat protein LINGO-1 enhances survival, structure, and function of dopaminergic neurons in Parkinson’s disease models . Proc Natl Acad Sci USA 2007 ; 104 : 14430 – 5 .
Johansen N , Somasundaram S , Travaglini KJ et al. Interindividual variation in human cortical cell type abundance and expression . Science 2023 ; 382 : eadf2359 .
Kang E , Kang M , Ju Y et al. Association between ARID2 and RAS-MAPK pathway in intellectual disability and short stature . J Med Genet 2021 ; 58 : 767 – 77 .
Kang HM , Subramaniam M , Targ S et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation . Nat Biotechnol 2018 ; 36 : 89 – 94 .
Kim H-J , Booth G , Saunders L et al. Nuclear oligo hashing improves differential analysis of single-cell RNA-seq . Nat Commun 2022 ; 13 : 2666 .
Kumasaka N , Rostom R , Huang N et al. Mapping interindividual dynamics of innate immune response at single-cell resolution . Nat Genet 2023 ; 55 : 1066 – 75 .
Kwok AJ , Allcock A , Ferreira RC et al. ; Emergency Medicine Research Oxford (EMROx) . Neutrophils and emergency granulopoiesis drive immune suppression and an extreme response endotype during sepsis . Nat Immunol 2023 ; 24 : 767 – 79 .
Li B , Gould J , Yang Y et al. Cumulus provides cloud-based data analysis for large-scale single-cell and single-nucleus RNA-seq . Nat Methods 2020 ; 17 : 793 – 8 .
Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data . Bioinformatics 2011 ; 27 : 2987 – 93 .
Li H , Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform . Bioinformatics 2009 ; 25 : 1754 – 60 .
Lotta LA , Pietzner M , Stewart ID et al. ; MacTel Consortium . A cross-platform approach identifies genetic regulators of human metabolism and health . Nat Genet 2021 ; 53 : 54 – 64 .
Manach C , Milenkovic D , Van de Wiele T et al. Addressing the inter-individual variation in response to consumption of plant food bioactives: towards a better understanding of their role in healthy aging and cardiometabolic risk reduction . Mol Nutr Food Res 2017 ; 61 : 1600557 .
McCarthy DJ , Campbell KR , Lun ATL et al. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R . Bioinformatics 2017 ; 33 : 1179 – 86 .
McGinnis CS , Patterson DM , Winkler J et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices . Nat Methods 2019 ; 16 : 619 – 26 .
McKenzie AT , Katsyv I , Song W-M et al. DGCA: a comprehensive R package for differential gene correlation analysis . BMC Syst Biol 2016 ; 10 : 106 .
Moffat JJ , Smith AL , Jung E-M et al. Neurobiology of ARID1B haploinsufficiency related to neurodevelopmental and psychiatric disorders . Mol Psychiatry 2022 ; 27 : 476 – 89 .
Murdock MH , Tsai LH. Insights into Alzheimer’s disease from single-cell genomic approaches . Nat Neurosci 2023 ; 26 : 181 – 95 .
Nassiri I , Fairfax B , Lee A et al. scQCEA: a framework for annotation and quality control report of single-cell RNA-sequencing data . BMC Genomics 2023 ; 24 : 381 .
Nassiri I , McCall MN. Systematic exploration of cell morphological phenotypes associated with a transcriptomic query . Nucleic Acids Res 2018 ; 46 : e116 .
Neavin D , Senabouth A , Arora H et al. Demuxafy: improvement in droplet assignment by integrating multiple single-cell demultiplexing and doublet detection methods. Genome Biol 2024 ; 25 :94.
Oelen R , de Vries DH , Brugge H et al. ; BIOS Consortium . Single-cell RNA-sequencing of peripheral blood mononuclear cells reveals widespread, context-specific gene expression regulation upon pathogenic exposure . Nat Commun 2022 ; 13 : 3267 .
Ranjan B , Sun W , Park J et al. DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data . Nat Commun 2021 ; 12 : 5849 .
Reus LM , Pasaniuc B , Posthuma D et al. ; International FTD-Genomics Consortium . Gene expression imputation across multiple tissue types provides insight into the genetic architecture of frontotemporal dementia and its clinical subtypes . Biol Psychiatry 2021 ; 89 : 825 – 35 .
Rizzo AN , Yuan JXJ. NEDD9 provides mechanistic insight into the coagulopathy of COVID-19 . Pulm Circ 2022 ; 12 : e12087 .
St George-Hyslop F , Haneklaus M , Kivisild T et al. Loss of CNTNAP2 alters human cortical excitatory neuron differentiation and neural network development . Biol Psychiatry 2023 ; 94 : 780 – 91 .
Stoeckius M , Zheng S , Houck-Loomis B et al. Cell hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics . Genome Biol 2018 ; 19 : 224 .
Tang L , Zhang D , Han P et al. Structural basis of SARS-CoV-2 and its variants binding to intermediate horseshoe bat ACE2 . Int J Biol Sci 2022 ; 18 : 4658 – 68 .
Van der Auwera GA , Carneiro MO , Hartl C et al. From FastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline . Curr Protoc Bioinformatics 2013 ; 43 : 11.10.11 – .10.33 .
van der Wijst MGP , Brugge H , de Vries DH et al. ; BIOS Consortium . Single-cell RNA sequencing identifies celltype-specific cis-eQTLs and co-expression QTLs . Nat Genet 2018 ; 50 : 493 – 7 .
Wang L , Li S , Stone SS et al. The role of the lncRNA MALAT1 in neuroprotection against hypoxic/ischemic injury . Biomolecules 2022 ; 12 : 146 .
Wang W , Min L , Qiu X et al. Biological function of long non-coding RNA (LncRNA) xist . Front Cell Dev Biol 2021 ; 9 : 645647 .
Wolock SL , Lopez R , Klein AM. Scrublet: computational identification of cell doublets in single-cell transcriptomic data . Cell Syst 2019 ; 8 : 281 – 91.e9 .
Wu TZ et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data . Innovation (Camb) 2021 ; 2 : 100141 .
Xin H , Lian Q , Jiang Y et al. GMM-Demux: sample demultiplexing, multiplet detection, experiment planning, and novel cell-type verification in single cell sequencing . Genome Biol 2020 ; 21 : 188 .
Xu J , Falconer C , Nguyen Q et al. Genotype-free demultiplexing of pooled single-cell RNA-seq . Genome Biol 2019 ; 20 : 290 .
Yang PY , Huang H , Liu CL. Feature selection revisited in the single-cell era . Genome Biol 2021 ; 22 : 321 .
Yazar S , Alquicira-Hernandez J , Wing K et al. Single-cell eQTL mapping identifies cell type-specific genetic control of autoimmune disease . Science 2022 ; 376 : eabf3041 .
| Month: | Total Views: |
|---|---|
| June 2024 | 234 |
Email alerts
Citing articles via.
- Recommend to your Library
- Advertising & Corporate Services
- Journals Career Network
Affiliations
- Online ISSN 2635-0041
- Copyright © 2024 Oxford University Press
- About Oxford Academic
- Publish journals with us
- University press partners
- What we publish
- New features
- Open access
- Institutional account management
- Rights and permissions
- Get help with access
- Accessibility
- Advertising
- Media enquiries
- Oxford University Press
- Oxford Languages
- University of Oxford
Oxford University Press is a department of the University of Oxford. It furthers the University's objective of excellence in research, scholarship, and education by publishing worldwide
- Copyright © 2024 Oxford University Press
- Cookie settings
- Cookie policy
- Privacy policy
- Legal notice
This Feature Is Available To Subscribers Only
Sign In or Create an Account
This PDF is available to Subscribers Only
For full access to this pdf, sign in to an existing account, or purchase an annual subscription.

- Advanced Search
Accurate multi-view clustering to seek the cross-viewed yet uniform sample assignment via tensor feature matching
New citation alert added.
This alert has been successfully added and will be sent to:
You will be notified whenever a record that you have chosen has been cited.
To manage your alert preferences, click on the button below.
New Citation Alert!
Please log in to your account
Information & Contributors
Bibliometrics & citations, view options, recommendations, multi-view clustering via spectral partitioning and local refinement.
A new multi-view clustering algorithm is proposed.The proposed MVNC algorithm uses spectral partitioning and local refinement.MVNC is compared to state-of-the-art algorithms using three real-world datasets.MVNC significantly outperforms the other ...
Multi-view Clustering with Graph Embedding for Connectome Analysis
Multi-view clustering has become a widely studied problem in the area of unsupervised learning. It aims to integrate multiple views by taking advantages of the consensus and complimentary information from multiple views. Most of the existing works in ...
Multi-view clustering with dual tensors
Multi-view clustering methods based on tensor have achieved favorable performance thanks to the powerful capacity of capturing the high-order correlation hidden in multi-view data. However, many existing works only pay attention to exploring the ...
Information
Published in.
Elsevier Science Inc.
United States
Publication History
Author tags.
- Multi-view clustering
- Tensor feature matching
- Uniform sample assignment
- Multimedia computing
- Data mining
- Research-article
Contributors
Other metrics, bibliometrics, article metrics.
- 0 Total Citations
- 0 Total Downloads
- Downloads (Last 12 months) 0
- Downloads (Last 6 weeks) 0
View options
Login options.
Check if you have access through your login credentials or your institution to get full access on this article.
Full Access
Share this publication link.
Copying failed.
Share on social media
Affiliations, export citations.
- Please download or close your previous search result export first before starting a new bulk export. Preview is not available. By clicking download, a status dialog will open to start the export process. The process may take a few minutes but once it finishes a file will be downloadable from your browser. You may continue to browse the DL while the export process is in progress. Download
- Download citation
- Copy citation
We are preparing your search results for download ...
We will inform you here when the file is ready.
Your file of search results citations is now ready.
Your search export query has expired. Please try again.

IMAGES
VIDEO
COMMENTS
Browse various samples of academic assignments written by professional writers for different disciplines and levels. Learn how to plan, structure, and write your own assignment with our tips and guidance.
Find sample papers formatted in seventh edition APA Style for different types of professional and student papers. Download Word files to use as templates and edit them as needed for your own papers.
Assignment is a task given to students by a teacher or professor, usually as a means of assessing their understanding and application of course material. Assignments can take various forms, including essays, research papers, presentations, problem sets, lab reports, and more. Assignments are typically designed to be completed outside of class ...
Learn how to read and interpret college writing assignments by identifying the task, audience, evidence, style, and format. Find tips and examples for different types of assignments and key terms.
This handout provides information about annotated bibliographies in MLA, APA, and CMS. These OWL resources will help you understand and complete specific types of writing assignments, such as annotated bibliographies, book reports, and research papers. This section also includes resources on writing academic proposals for conference ...
Media Files: APA Sample Student Paper , APA Sample Professional Paper This resource is enhanced by Acrobat PDF files. Download the free Acrobat Reader. Note: The APA Publication Manual, 7 th Edition specifies different formatting conventions for student and professional papers (i.e., papers written for credit in a course and papers intended for scholarly publication).
In this sample paper, we've put four blank lines above the title. Commented [AF3]: Authors' names are written below the title, with one double-spaced blank line between them. Names should be written as follows: First name, middle initial(s), last name. Commented [AF4]: Authors' affiliations follow immediately after their names.
Come up with a thesis. Create an essay outline. Write the introduction. Write the main body, organized into paragraphs. Write the conclusion. Evaluate the overall organization. Revise the content of each paragraph. Proofread your essay or use a Grammar Checker for language errors. Use a plagiarism checker.
To construct an assignment structure, use outlines. These are pieces of text that relate to your topic. It can be ideas, quotes, all your thoughts, or disparate arguments. Type in everything that you think about. Separate thoughts scattered across the sheets of Word will help in the next step. Then it is time to form the text.
Popular essay samples. Though we have an extensive range of samples and are always adding and writing more, you'll find . a selection of the topics that students search for most often down below. Search. Social Issues: Animal Rights. Teenage Pregnancy. Сyber Bullying. Racism.
View or download authentic sample assignments for academic writing skills. Find annotated bibliography, essay, literature review, reflective writing and report examples.
Learn how to write effective assignments with clear and simple tips on outline, introduction, structure, referencing, conclusion and more. See an example of an assignment on obesity epidemic and its causes and solutions.
Tables and figures are only included in student papers if needed for the assignment. Tables and figures share the same elements and layout. See the website for sample tables and sample figures. Table elements. Tables include the following four elements: Number; Title; Body (rows and columns) Note (optional if needed to explain elements in the ...
hardest thinking, and feel the greatest sense of mastery and. growth, in their writing. Cour. es. and assignments should be planned with this in mi. d. Three principles are paramount:1. Name what you want and imagine students doing itHowever free students are to range and explore in a paper, the general kind of paper you're inviting has com.
nment. Some people find it easiest to write the introduction first, whereas others leave it until. the end. Neither approach is right or wrong, so write the assignment in whichever order feels best. for you. The introduction might be up to around 10% of the word count (e.g. up to 200 words for a 2000 word ass.
A sample of a business-style report with an annotated format. Investigative report sample. Sample of an investigative report [PDF 500KB]. A resource for science, engineering and technology students. How to write an investigative report, including an annotated format. Assignment topics and editing. Interpreting assignment topics [PDF 370 KB ...
This guide includes tips on writing common course assignments. Both in traditional and online classrooms, journal entries are used as tools for student reflection. By consciously thinking about and comparing issues, life experiences, and course readings, students are better able to understand links between theory and practice and to generate ...
From college assignment samples to university essay assignment samples, students can access all kinds of resources from our collection. Our top-qualified experts curate all these solved papers. So, if you check a sample of our essay writing, you can immediately notice the difference in quality between us and a fake. We also regularly update our ...
Learn how to create effective writing assignments for your course by defining the purpose, audience, mode, and criteria. Find tips on sequencing, checking, and selecting the format of your assignments.
43 Sample Writing Assignments Process Analysis. Be the expert and teach your reader how to do something. You will focus on writing the main steps to completing this process and organizing based on chronology or priority of steps. This essay could be demonstrative in nature (ex. How to bathe and groom a dog at home or how to make banana nut ...
Low stakes writing helps: Describe, apply, and retain information. Explore and personalize ideas. Focus thoughts and questions. Demonstrate the value of writing as a part of the learning process. Informally engage each student in the classroom. Improve high-stakes writing. Efficiently assess student learning.
This guide will provide research and writing tips to help students complete a literature review assignment. Home; Steps for Conducting a Lit Review; ... Chicago: Notes Bibliography This link opens in a new window; MLA Style This link opens in a new window; Sample Literature Reviews. Sample Lit Reviews from Communication Arts; Have an exemplary ...
Assignment and Course Assessment KCTL Spring Workshop 5/2/2024. Today's Objectives •Explain what rubrics are and how they differ from ... Using samples of existing work, divide into levels corresponding to the levels you set. What are the qualities of the best work? The
Lecture/text homework assignment # 6 Note: Please circle your answers when appropriate! 0) Problems 6 & 7 from last time are due; note that your recitation instructor may ask you to present ... For both the sample standard deviation is about s = 7.65 cm. (a) Suppose you take a sample of 8 women and 6 men. Construct a 99% CI for both. Do the
Cite your source automatically in MLA or APA format. Cite. Using citation machines responsibly. Powered by. Media File: APA Sample Paper. This resource is enhanced by an Acrobat PDF file. Download the free Acrobat Reader. Click this link to download the PDF handout of the APA Sample Paper.
An assignment can be submitted late, with a participation-score penalty of 3 points per 24-hour-period, or fraction thereof, after the deadline. The teaching staff is happy to discuss assignment requirements, and to discuss how you might go about completing them, but cannot confirm in advance that a draft assignment fulfills the requirements.
We utilized 30 samples from both sepsis and healthy individuals in six batches to assess the performance of classification approaches. The results indicate that combining genetic and EAD results can enhance the accuracy of assignments (Min. 0.94, Mean 0.98, Max. 1).
Finally, a sample assignment matrix is learned via jointly reconstructing the samples using the two bases further cooperating with spectral clustering. In this way, the heterogeneity gap is bridged by tensor feature matching, and the inaccurate sample-wise similarity estimation is omitted by using feature-wise matching to guide sample-wise ...
login.microsoftonline.com