Advertisement


About The Journal of Immunology
The Journal of Immunology publishes peer-reviewed manuscripts describing novel findings in all areas of experimental immunology, including both basic and clinical studies.
Read More About the Journal
Editor-in-Chief Gail A. Bishop, Ph.D. Full Editorial Board
Seeking EIC Candidates
ImmunoHorizons is accepting applications For its next Editor-in-Chief

Special Collection: Immunology of Zoonotic Diseases and Relevant Animal Models
Special collection curated by Guest Editor Crystal L. Loving, PhD , featuring Brief Reviews on the immunology of zoonotic diseases and relevant animal models
.jpg?versionId=611)
AAI and its journals, The Journal of Immunology and ImmunoHorizons, have become aware that a predatory publisher is sending emails to authors to solicit manuscript submissions. To better inform our valued authors, we compiled a resource on how to spot a predatory publisher. Learn more by visiting the FAQ

AAI Volunteer Portal

Pillars of Immunology
The Journal of Immunology publishes commentaries highlighting scientific reports that were published more than 15 years ago and that have made a major impact on the course of immunological research today. Read published commentaries here .

Don't miss the exciting Brief Review Collection, coming in the January 15, 2024, issue of The Journal of Immunology!

Sign Up for New Issue Alerts
Wondering when the next issues of The JI or ImmunoHorizons are coming out?
Sign up for email alerts so that you don't miss an single issue!
- Current Issue
- Next in The JI
- Collections
Submit a Manuscript
- Instructions for Authors
- About the JI
- Journal Policies
- The JI Editorial Board
- Rights and Permissions
- Follow The JI on Twitter
- Online ISSN 1550-6606
- Print ISSN 0022-1767
Publications
- The Journal of Immunology
- ImmunoHorizons
- AAI Newsletter
- Additional AAI Publications
General Information
- Advertisers
- Subscribers
- Privacy Policy
Journal Services
- Email Alerts
- ImmunoCasts
- Twitter JI
- Cookie Policy
- Accessibility
- Terms & Conditions
- Get Adobe Acrobat Reader
- © 2024 The American Association of Immunologists, Inc.
This Feature Is Available To Subscribers Only
Sign In or Create an Account
- Search by keyword
- Search by citation
Page 1 of 24
Association of interleukin-17A and chemokine/vascular endothelial growth factor-induced angiogenesis in newly diagnosed patients with bladder cancer
The human interleukin-17 (IL-17) family comprises IL-17A to IL-17 F; their receptors are IL-17RA to IL-17RE. Evidence revealed that these cytokines can have a tumor-supportive or anti-tumor impact on human mal...
- View Full Text
Causal relationship between immune cells and telomere length: mendelian randomization analysis
The causal relationship between immune cells and telomere length remains controversial.
Subcutaneous immunoglobulin replacement therapy in patients with immunodeficiencies – impact of drug packaging and administration method on patient reported outcomes
Here, the perspective of patients with primary and secondary immunodeficiency receiving subcutaneous immunoglobulin (SCIg) via introductory smaller size pre-filled syringes (PFS) or vials were compared.
Dendritic cells under allergic condition enhance the activation of pruritogen-responsive neurons via inducing itch receptors in a co-culture study
Itch sensitization has been reported in patients with chronic allergic skin diseases and observed in a mouse model of allergic contact dermatitis (ACD). There is evidence suggesting that neuroimmune interactio...
Network pharmacology-based strategy to investigate the mechanisms of artemisinin in treating primary Sjögren’s syndrome
The study aimed to explore the mechanism of artemisinin in treating primary Sjögren’s syndrome (pSS) based on network pharmacology and experimental validation.
Hyperactivation and enhanced cytotoxicity of reduced CD8 + gamma delta T cells in the intestine of patients with Crohn’s disease correlates with disease activity
We aimed to investigate the immune characteristics of intestinal CD8 + gamma delta T (CD8 + γδ T) cells in Crohn’s disease (CD) and their correlation with disease activity.
Immune responses to P falciparum antibodies in symptomatic malaria patients with variant hemoglobin genotypes in Ghana
Haemoglobin (Hb) variants such as sickle cell trait (SCT/HbAS) play a role in protecting against clinical malaria, but little is known about the development of immune responses against malaria parasite ( Plasmodiu...
Systematic evaluation of B-cell clonal family inference approaches
The reconstruction of clonal families (CFs) in B-cell receptor (BCR) repertoire analysis is a crucial step to understand the adaptive immune system and how it responds to antigens. The BCR repertoire of an ind...
Expression of the immune checkpoint molecules CD226 and TIGIT in preeclampsia patients
Imbalanced immune responses are involved in developing preeclampsia (PE). We wish to explore the expression and potential changes of immune checkpoint molecules TIGIT, CD226 and CD155 in PE patients.
Oral administration of DNA alginate nanovaccine induced immune-protection against Helicobacter pylori in Balb/C mice
Helicobacter pylori (H. Pylori), is an established causative factor for the development of gastric cancer and the induction of persistent stomach infections that may lead to peptic ulcers. In recent decades, s...
Profiling of T cell repertoire in peripheral blood of patients from type 2 diabetes with complication
More than 90% of patients with diabetes worldwide are type 2 diabetes (T2D), which is caused by insulin resistance or impaired producing insulin by pancreatic β cells. T2D and its complications, mainly large c...
Tumor microenvironment and immune system preservation in early-stage breast cancer: routes for early recurrence after mastectomy and treatment for lobular and ductal forms of disease
Intra-ductal cancer (IDC) is the most common type of breast cancer, with intra-lobular cancer (ILC) coming in second. Surgery is the primary treatment for early stage breast cancer. There are now irrefutable d...
Predictive biomarkers for immune-related adverse events in cancer patients treated with immune-checkpoint inhibitors
The objective of this study was to identify potential predictors of immune-related adverse events (irAEs) in cancer patients receiving immune checkpoint inhibitor therapy among serum indexes, case data, and li...
Negative prognostic behaviour of PD-L1 expression in tongue and larynx squamous cell carcinoma and its significant predictive power in combination with PD-1 expression on TILs
Biomarkers that can predict outcome will improve the efficacy of treatment for HNSCC patients. In this regard, we retrospectively evaluated the prognostic effect of PD1, PD-L1, and CD45RO in tongue and larynx ...
Whole blood stimulation provides preliminary evidence of altered immune function following SRC
To implement an approach combining whole blood immune stimulation and causal modelling to estimate the impact of sport-related concussion (SRC) on immune function.
IgG antibody response to SARS-CoV-2 infection and its influencing factors in lymphoma patients
The ability of generating effective humoral immune responses to SARS-CoV-2 infection has not been clarified in lymphoma patients. The study aimed to investigate the antibody (Ab) production after SARS-Cov-2 in...
Polarized Th2 cells attenuate high-fat-diet induced obesity through the suppression of lipogenesis
Immune cells, such as macrophages, B cells, neutrophils and T cell subsets, have been implicated in the context of obesity. However, the specific role of Th2 cells in adipose tissue function has remained elusi...
The predictive value of peripheral blood CD4 cells ATP concentration for immune-related adverse events in advanced non-small cell lung cancer patients
Lung cancer with the highest incidence and mortality in the world. Immune checkpoint inhibitors (ICIs), can bring long-term survival benefits to patients, but also can bring immune-related adverse events (irAE...
Evaluation of the TLR3 involvement during Schistosoma japonicum -induced pathology
Despite the functions of TLRs in the parasitic infections have been extensively reported, few studies have addressed the role of TLR3 in the immune response to Schistosoma japonicum infections. The aim of this st...
PRMT2 silencing regulates macrophage polarization through activation of STAT1 or inhibition of STAT6
Macrophages play significant roles in innate immune responses and are heterogeneous cells that can be polarized into M1 or M2 phenotypes. PRMT2 is one of the type I protein arginine methyltransferases involved...
TRPV1 + neurons alter Staphylococcus aureus skin infection outcomes by affecting macrophage polarization and neutrophil recruitment
The interaction between the nervous system and the immune system can affect the outcome of a bacterial infection. Staphylococcus aureus skin infection is a common infectious disease, and elucidating the relations...
Retraction Note: Oral supplementation of diabetic mice with propolis restores the proliferation capacity and chemotaxis of B and T lymphocytes towards CCL21 and CXCL12 by modulating the lipid profile, the pro-inflammatory cytokine levels and oxidative stress
Cd39 identifies a specific cd8 + t cell population in lung adenocarcinoma-related metastatic pleural effusion.
Malignant pleural effusion (MPE), which is a complex microenvironment that contains numerous immune and tumour signals, is common in lung cancer. Gene alterations, such as driver gene mutations, are believed t...
Dissecting cellular states of infiltrating microenvironment cells in melanoma by integrating single-cell and bulk transcriptome analysis
Cellular states of different immune cells can affect the activity of the whole immune microenvironment.
Sec1 regulates intestinal mucosal immunity in a mouse model of inflammatory bowel disease
Inflammatory bowel disease (IBD) is a common immune-mediated condition with its molecular pathogenesis remaining to be fully elucidated. This study aimed to deepen our understanding of the role of FUT2 in human I...
Screening of four lysosome-related genes in sepsis based on RNA sequencing technology
Screening of lysosome-related genes in sepsis patients to provide direction for lysosome-targeted therapy.
Dominant negative biologics normalise the tumour necrosis factor (TNF-α) induced angiogenesis which exploits the Mycobacterium tuberculosis dissemination
Tumor necrosis factor (TNF) is known to promote T cell migration and increase the expression of vascular endothelial growth factor (VEGF) and chemokines. The administration of Xpro-1595, a dominant-negative TN...
Activation dynamics of antigen presenting cells in vivo against Mycobacterium bovis BCG in different immunized route
Control of Tuberculosis (TB) infection is mainly the result of productive teamwork between T-cell populations and antigen presenting cells (APCs). However, APCs activation at the site of initiating cellular im...
Characteristics of circulating immune cells in HBV-related acute-on-chronic liver failure following artificial liver treatment
Liver failure, which is predominantly caused by hepatitis B (HBV) can be improved by an artificial liver support system (ALSS). This study investigated the phenotypic heterogeneity of immunocytes in patients w...
Putative novel outer membrane antigens multi-epitope DNA vaccine candidates identified by Immunoinformatic approaches to control Acinetobacter baumannii
Multi-epitope polypeptide vaccines, a fusion protein, often have a string-of-beads system composed of various specific peptide epitopes, potential adjuvants, and linkers. When choosing the sequence of various ...
Long-term humoral and cellular immunity after primary SARS-CoV-2 infection: a 20-month longitudinal study
SARS-CoV-2 remains a world-wide health issue. SARS-CoV-2-specific immunity is induced upon both infection and vaccination. However, defining the long-term immune trajectory, especially after infection, is limi...
Exploration of biomarkers for systemic lupus erythematosus by machine-learning analysis
In recent years, research on the pathogenesis of systemic lupus erythematosus (SLE) has made great progress. However, the prognosis of the disease remains poor, and high sensitivity and accurate biomarkers are...
Tacrolimus reverses pemphigus vulgaris serum-induced depletion of desmoglein in HaCaT cells via inhibition of heat shock protein 27 phosphorylation
Glucocorticoids are the first-line treatment for Pemphigus vulgaris (PV), but its serious side effects can be life-threatening for PV patients. Tacrolimus (FK506) has been reported to have an adjuvant treatmen...
Increased infiltration of CD4 + T cell in the complement deficient lymphedema model
Lymphedema is an intractable disease that can be caused by injury to lymphatic vessels, such as by surgical treatments for cancer. It can lead to impaired joint mobility in the extremities and reduced quality ...
Vitamin D and biomarkers of inflammation and oxidative stress among pregnant women: a systematic review of observational studies
This systematic review aimed to map the evidence evaluated the relationship between vitamin D and redox and inflammatory status during gestation.
The upregulation of peripheral CD3 - CD56 + CD16 + natural killer cells correlates with Th1/Th2 imbalance in asthma patients during acute upper respiratory viral infections
The aim of this study is to clarify the changes of peripheral CD3 − CD56 + CD16 + NK cells and their correlation with Th1/Th2 immunity profiles in asthma during the phase of acute upper respiratory viral infections (A...
Humoral immune response and changes in peritoneal cell populations in rats immunized against two Leptospira serovars; serovar patoc and serovar pyrogenes
Leptospirosis is a zoonotic disease caused by Leptospira species. Variations in lipopolysaccharide (LPS) structure in Leptospira are known to be associated with the serovar diversity and antigenicity. Development...
Methionine enkephalin(MENK) upregulated memory T cells in anti-influenza response
Novel prophylactic drugs and vaccination strategies for protection against influenza virus should induce specific effector T-cell immune responses in pulmonary airways and peripheral lymphoid organs. Designing...
Transcriptomic analysis identifies CYP27A1 as a diagnostic marker for the prognosis and immunity in lung adenocarcinoma
The association between lipid metabolism disorder and carcinogenesis is well-established, but there is limited research on the connection between lipid metabolism-related genes (LRGs) and lung adenocarcinoma (...
The digestive system and autoimmunity
Digestive autoimmune conditions are a growing challenge to global health. Risk factors associated with autoimmune digestive diseases are complex, including genetic variation, immunological dysfunction, and var...
Bcl-3 regulates T cell function through energy metabolism
Bcl-3 is a member of the IκB protein family and an essential modulator of NF-κB activity. It is well established that Bcl-3 is critical for the normal development, survival and differentiation of adaptive immu...
The development of a highly sensitive and quantitative SARS-CoV-2 rapid antigen test applying newly developed monoclonal antibodies to an automated chemiluminescent flow-through membrane immunoassay device
Rapid and accurate diagnosis of individuals with SARS-CoV-2 infection is an effective way to prevent and control the spread of COVID-19. Although the detection of SARS‐CoV‐2 viral RNA by RT‐qPCR is the gold st...
Immune cell profiles of idiopathic inflammatory myopathy patients expressed anti-aminoacyl tRNA synthetase or anti-melanoma differentiation-associated gene 5 autoantibodies
Patients with idiopathic inflammatory myopathy (IIM) often express a different type of myositis-specific autoantibodies (MSAs), each associated with different clinical symptoms. Understanding the immunopathoge...
Development and validation of a machine learning-based nomogram for predicting HLA-B27 expression
HLA-B27 positivity is normal in patients undergoing rheumatic diseases. The diagnosis of many diseases requires an HLA-B27 examination.
Function and autophagy of monocyte-derived dendritic cells is affected by hepatitis B virus infection
The role of dendritic cells and the autophagy state of dendritic cells in the immune response of hepatitis B virus (HBV) infection was still controversial. In this study, we carefully examined the phenotype, f...
Evaluation of the relationship between serum interleukin-1β levels and expression of inflammasome-related genes in patients with COVID-19
Inflammasomes are a group of molecules that are strongly involved in causing inflammation. This study aimed to evaluate the expression of NLR family pyrin domain containing 1 (NLRP1), NLRP3, and Apoptosis-asso...
Comparison of the effect of autoclaved and non-autoclaved live soil exposure on the mouse immune system
. Lack of exposure to the natural microbial diversity of the environment has been linked to dysregulation of the immune system and numerous noncommunicable diseases, such as allergies and autoimmune disorders....
The effect of serum origin on cytokines induced killer cell expansion and function
Cytokine-induced killer (CIK) cells have shown promising results in adoptive immunotherapy. However, serum may play a determining role in the large-scale expansion of these cells for clinical applications. Acc...
Efficacy and safety of immune checkpoint inhibitors in Proficient Mismatch Repair (pMMR)/ Non-Microsatellite Instability-High (non-MSI-H) metastatic colorectal cancer: a study based on 39 cohorts incorporating 1723 patients
This study was designed to investigate the efficacy and safety of immune checkpoint inhibitors (ICIs)-based therapy in proficient mismatch repair (pMMR)/non-microsatellite instability-high (non-MSI-H) metastat...
Higher plasma interleukin − 6 levels are associated with lung cavitation in drug-resistant tuberculosis
Lung cavitation is associated with heightened TB transmission and poor treatment outcomes. This study aimed to determine the relationship between systemic inflammation and lung cavitation in drug-resistant TB ...
Featured videos
View featured videos from across the BMC-series journals
Important information
Editorial board
For authors
For editorial board members
For reviewers
- Manuscript editing services
Annual Journal Metrics
2022 Citation Impact 3.0 - 2-year Impact Factor 3.6 - 5-year Impact Factor 0.808 - SNIP (Source Normalized Impact per Paper) 0.783 - SJR (SCImago Journal Rank)
2023 Speed 26 days submission to first editorial decision for all manuscripts (Median) 188 days submission to accept (Median)
2023 Usage 671,800 downloads 176 Altmetric mentions
- More about our metrics
- Follow us on Twitter
BMC Immunology
ISSN: 1471-2172
- General enquiries: [email protected]

Low C4A copy numbers and higher HERV gene insertion contributes to increased risk of SLE, with absence of association with disease phenotype and disease activity
- Christina Mary Mariaselvam
- Gaurav Seth
- Ryad Tamouza
Hormone and reproductive factors and risk of systemic lupus erythematosus: a Mendelian randomized study
- Runyu Chang
- Shate Xiang
- Xinghong Ding
Exploring the causal relationship between Takayasu arteritis and inflammatory bowel disease using Mendelian randomization
- Xiaoli Pang
- Huizhong Yang

Polyethylene glycol and immunology: aspects of allergic reactions and their mechanisms, as well as ways to prevent them in clinical practice
- Maria Zofia Lisiecka

mTOR-mediated differentiation and maintenance of suppressive T cells at the center stage of IPEX treatment
- Rafael Cardoso Maciel Costa Silva
Assessing the steroid-sparing effect of biological agents in randomized controlled trials for lupus: a scoping review
- Savino Sciascia
- Silvia Grazietta Foddai
- Dario Roccatello
Involvement of gut microbiota in multiple sclerosis—review of a new pathophysiological hypothesis and potential treatment target
- Piotr Olejnik
- Kasper Buczma
- Kaja Kasarełło
Correction to: Characteristics of splenic PD-1 + γδT cells in Plasmodium yoelii nigeriensis infection
- Dianhui Chen
- Xingfei Pan
Correction to: The immune regulatory function of B7-H3 in malignancy: spotlight on the IFN-STAT1 axis and regulation of tumor-associated macrophages
- Jong Dae Ji
Monophosphoryl lipid A as a co-adjuvant in methicillin-resistant Staphylococcus aureus vaccine development: improvement of immune responses in a mouse model of infection
- Mehdi Mirshekar
- Setareh Haghighat
- Mohammad Hossein Yazdi
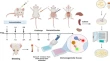
Mass cytometry revealed the circulating immune cell landscape across different Suzuki stages of Moyamoya disease
- Chenglong Liu
- Jizong Zhao
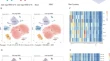
Acute stress transiently activates macrophages and chemokines in cervical lymph nodes
- Akihiro Dohi
- Tadahide Noguchi
- Yoshiyuki Mori
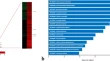

Chronic hyperglycemia impairs anti-microbial function of macrophages in response to Mycobacterium tuberculosis infection
- Gaurav Kumar Chaubey
- Radheshyam Modanwal
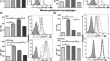
Rapid generation of an RBL cellular model to study proteins that cause allergenic reactions in vitro
- Israel Hernández-Aguilar
- Juan Carlos Vizuet-de-Rueda
- Luis M. Teran
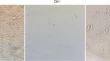

Study of pathogenic T-helper cell subsets in Asian Indian patients with Takayasu arteritis
- P. M. Punithavathy
- Ramesh Babu Telugu
- Ruchika Goel
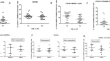

Blood and CSF anti-neuronal antibodies testing in psychotic syndromes: a retrospective analysis from a tertiary psychiatric hospital
- Joana Lopes
- Maria João Malaquias
- Ana Paula Correia
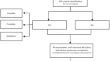
Rituximab alleviates pediatric systemic lupus erythematosus associated refractory immune thrombocytopenia: a case-based review
- Qiaoyan Guo
Evidence of concerning decline of COVID-19 vaccination in older persons
- Camilla Mattiuzzi
- Giuseppe Lippi

The immune regulatory function of B7-H3 in malignancy: spotlight on the IFN-STAT1 axis and regulation of tumor-associated macrophages

Characteristics of splenic PD-1 + γδT cells in Plasmodium yoelii nigeriensis infection
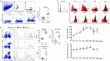

APLAID complicated with arrhythmogenic dilated cardiomyopathy caused by a novel PLCG2 variant
- Tianjiao Wang
- Xiaolin Liu
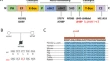
Evolutionary preservation of CpG dinucleotides in RAG1 may elucidate the relatively high rate of methylation-mediated mutagenesis of RAG1 transposase
- Mariam M. Fawzy
- Maiiada H. Nazmy
- Moustafa Fathy

DNA methylation profiling of labial salivary gland tissues revealed hypomethylation of B-cell-related genes in primary Sjögren’s syndrome
- Jayakanthan Kabeerdoss
- Prabavathi Devarajalu
- Pulukool Sandhya

Response to: regarding the significance of anti-COVID-IgA antibody response in COVID-19 breakthrough infection
- Sabiha Anis
- Mariam Ashfaq Khan
- Samreen Sarfaraz
The combination of IDO and AHR blockers reduces the migration and clonogenicity of breast cancer cells
- Maryam Soltani-asl
- Parviz Azimnasab-sorkhabi
- Jose Roberto Kfoury Jr.
Conventional Tregs in treatment-naïve rheumatoid arthritis are deficient in suppressive function with an increase in percentage of CXCR3 and CCR6 expressing Tregs
- Vallayyachari Kommoju
- Vir Singh Negi
Serum soluble LYVE1 is a promising non-invasive biomarker of renal fibrosis: a population-based retrospective cross-sectional study
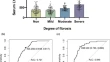

COVID-19 in Systemic Lupus Erythematosus patients treated with belimumab: a retrospective clinical study
- Martin Herrmann
Regarding the significance of anti-COVID-IgA antibody response in COVID-19 breakthrough infection
An epicutaneous therapeutic pollen-allergen extract delivery system in an allergic rhinitis mouse model: based on allergen loading on dc-specific aptamers conjugated nanogolds.
- Safoora Pordel
- Navideh Haghnavaz
- Mojtaba Sankian
Discouraging Non-ELISA antiphospholipid antibody assays in antiphospholipid syndrome classification may hinder clinical research
- Xiaochun Susan Zhang
- Nicola Bizzaro
- Jan Damoiseaux
Quantitative evaluation of citrullinated fibrinogen for detection of neutrophil extracellular traps
- Tsubasa Sue
- Tomoki Ichikawa
- Yumiko Higuchi
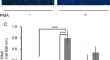

Long non-coding RNA ENSMUST00000197208 promotes a shift in the Th17/Treg ratio via the P2X7R-NLRP3 inflammasome axis in collagen-induced arthritis

Correction: Major histocompatibility complex class I molecule expression by pancreatic cancer cells is regulated by activation and inhibition of the epidermal growth factor receptor
- Shelby M. Knoche
- Alaina C. Larson
- Joyce C. Solheim
Immune cells and hypertension
A ubiquitin–proteasome system-related signature to predict prognosis, immune infiltration, and therapy efficacy for breast cancer
- Meihuan Wang
- Huawei Zhang

Fructus lycii oligosaccharide alleviates acute liver injury via PI3K/Akt/mTOR pathway
- Xingxing Zhang
- Qingtong Yu


Impact of microparticles released during murine systemic inflammation on macrophage activity and reactive nitrogen species regulation
- Weronika Ortmann
- Elzbieta Kolaczkowska
Comparative analyses of various IgE-mediated and non-IgE-mediated inducers of mast cell degranulation for in vitro study
- Sunisa Yoodee
- Chuda Rujitharanawong
- Visith Thongboonkerd

Semaphorin 3 a restores the ability of type 1 regulatory T cells to suppress food allergy
- Pengyuan Zheng

Insights into the immunological description of cryoglobulins with regard to detection and characterization in Slovenian rheumatological patients
- Manca Ogrič
- Saša Čučnik

Increased synthesis and intestinal expression of IL-39 in patients with inflammatory bowel disease
- Gabriela Fonseca-Camarillo
- Janette Furuzawa-Carballeda
- Jesús K. Yamamoto-Furusho
Role of LINC00240 on T-helper 9 differentiation in allergic rhinitis through influencing DNMT1-dependent methylation of PU.1
- JianGuo Liu
- XunShuo Jiang
- ChunPing Yang

Association of IL-23R and IL-10 variations with Behçet disease: a genetic analysis study
- Guven Yenmis
- Sema Sabancelebi
- Tumay Sadikoglu
Prevalence and sociodemographic correlates of antinuclear antibody testing by indirect immunofluorescence or solid-phase assays in a Spanish population: the Camargo Cohort
- Juan Irure-Ventura
- Daniel Martínez-Revuelta
- José Luis Hernández
Rational design of multi-epitope-based vaccine by exploring all dengue virus serotypes proteome: an immunoinformatic approach
- Ahad Amer Alsaiari
- Mohammed Ageeli Hakami
- Farzana Yasmin

Differential expression of TOB/BTG family members in patients with plaque psoriasis: cross-sectional study
- Carlos A. Barrera-Ochoa

Association study between killer immunoglobulin-like receptor polymorphisms and susceptibility to COVID-19 disease: a systematic review and meta-analysis
- Saeed Hosseini Teshnizi
- Sara Mirzazadeh
- Kurosh Kalantar

Palpitation in women with silicone breast implants: association with autoantibodies against autonomic nervous system
- Gilad Halpert
- Yehuda Shoenfeld
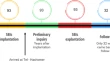
Current genetic defects in common variable immunodeficiency patients on the geography between Europe and Asia: a single-center experience
- Ezgi Topyıldız
- Necil Kutukculer
- Find a journal
- Publish with us
- Track your research
EDITORIAL article
Editorial: novel biomarkers in tumor immunity and immunotherapy.

- 1 Translational Research Department, Drug Development Laboratories, Kyoto R&D Center, Maruho Co., Ltd., Kyoto, Japan
- 2 Department of Pathology, University of Texas Southwestern Medical Center, Dallas, TX, United States
- 3 Department of Biological Sciences, School of Science, Xi’an Jiaotong-Liverpool University, Suzhou, Jiangsu, China
- 4 Biomedical Research Center, School of Science, Engineering and Environment, University of Salford, Manchester, United Kingdom
Editorial on the Research Topic Novel biomarkers in tumor immunity and immunotherapy
In this Research Topic, numerous researchers reported novel biomarkers and methodologies for predicting the efficacy of cancer immunotherapy across various cancers. Additionally, a wide spectrum of fundamental research has been conducted, leading to the discovery of biomarkers. Alongside traditional immunological analyses, a diverse array of methodologies such as bulk RNA-Seq, scRNA-Seq, and bacterial flora analysis have been employed. Moreover, state-of-the-art bioinformatics technologies have been effectively utilized in biomarker discovery. These investigations not only unveil intriguing new discoveries facilitated by cutting-edge technologies but also hold significant promise for shaping the future landscape of tumor immunology.
We are pleased to present this successful Research Topic to the scientific community. This Research Topic comprises six reviews and forty-one original papers. Four systemic reviews on predicting response to immune checkpoint inhibitors (ICI) were published: Qian et al. conducted a meta-analysis, affirming that plasma EBV DNA levels serve as reliable biomarkers for predicting favorable responses to ICI treatment in nasopharyngeal cancer patients, Rugambwa et al. established an association between high neutrophil-lymphocyte and platelet-lymphocyte ratios and poorer ICI treatment outcomes, and Fejza et al. presented accumulating evidence indicating extracellular matrix molecules as biomarkers identifying patients benefiting from ICI treatment. Shi et al. compared various predictive biomarker testing methods for ICI efficacy, while Wu et al. reviewed small molecule inhibitors for KRAS mutant cancers. Fonseca-Montaño et al. delved into the significance of long-non coding RNAs (lncRNAs) in breast cancer and their latest findings. These reviews furnish insights into the current status of previous studies in the realm of tumor immunology, aiding in the recognition and anticipation of forthcoming challenges.
Biomarkers encompass cancer-specific and cancer-nonspecific markers applicable across diverse cancer types. Within this purview, several intriguing factors have been identified as treatment response and prognosis markers in pan-cancer patients. Dong et al. revealed the multifaceted role of Proteasome Activator Complex Subunit 3 (PSME3) in tumors, establishing it as a pan-cancer prognostic marker. Lin et al. proposed Glioma pathogenesis related-2 (GLIPR2) as a promising novel biomarker and tumor suppressor. Liu et al. examined the functional attributes of Tubulin epsilon and delta complex 2 (TEDC2) in human tumors, identifying TEDC2 as a prognostic marker across various tumor types. Li et al. elucidated the role of disulfidoptosis-related genes (DRGs) in pan-cancer prognosis and their interplay with immunity, constructing a prognostic model utilizing various bioinformatics and machine learning techniques. Zhu et al. highlighted that high expression of Origin recognition complex 6 (ORC6) could serve as a prognostic biomarker in pan-cancer patients. Wei et al. showed the positive correlation between elevated expression levels of IFN-γ-related genes and drug sensitivity, emphasizing the pivotal role of IFN-γ in tumor immunotherapy. Pan et al. reported on the involvement of integrin-binding sialic acid protein (IBSP), a member of the small integrin-binding ligand N-linked glycoprotein (SIBLING) family, in tumorigenesis across various cancers, proposing IBSPs as prognostic biomarkers and immunotherapy targets in pan-cancer. Wu et al. delineated the prognostic potential of the Ferroptosis-related gene Hypermethylated in Cancer 1 (HIC1) in various cancers, indicative of its utility in predicting cancer prognosis, immunotherapy response, and drug sensitivity. Li et al. demonstrated the significant correlation of Thymosin beta-10 (TMSB10) with the tumor microenvironment and immune regulatory factors, advocating its role as a predictive marker for therapeutic response in cancer patients. Huang et al. identified Four Jointed Box 1 (FJX1) as a novel prognostic factor crucial in tumor immunity based on comparative expression profile analysis. Sun et al. established an association between dysregulation of the proprotein convertase subtilisin/kexin-9 (PCSK9) and poor clinical outcomes, suggesting its potential as a robust pan-cancer biomarker. These studies link these genes previously not directly linked to oncogenesis or tumor immunity to immune regulation and suggest potential role as biomarkers.
Studies focusing on specific tumors have unveiled several therapeutic and prognostic markers in hepatocellular carcinoma (HCC). Shi et al. developed the PCD Index (PCDI), comprising programmed cell death-related genes, as a prognostic and treatment response predictor in HCC. Zhang et al. observed elevated expression of DnaJ heat shock protein family member C8 (DNAJC8) in HCC tissues, correlating with poor prognosis and demonstrating its oncogenic role. Jiang et al. identified a significant correlation between CD93 expression and the prognosis of liver hepatocellular carcinoma patients. Xu et al. elucidated abnormal T follicular helper cell infiltration associated with forkhead box M1 (FOXM1) as a crucial prognostic factor in HCC patients.
Prominent biomarkers have also emerged from studies on lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). Li et al. focused on coagulation- and macrophage-associated (COMAR) genes, constructing a COMAR risk score model predictive of prognosis and clinical outcome in LUAD patients. Zhu et al. identified twelve HUB genes via Weighted Gene Coexpression Network Analysis (WGCNA), potentially implicated in LUAD progression via immune-related signaling pathways. Wu et al. derived LUSC-specific differentially expressed gene signatures (7-DEGs) with prognostic significance for LUSC patients.
A multitude of original and intensive investigations have explored valid biomarkers across a diverse array of tumors. Li et al. identified hub biomarkers closely associated with gastric cancer (GC) using microarray data and algorithmic approaches. Cai et al. delineated the multifaceted role of Fibroblast activation protein (FAP) in gastrointestinal cancer progression. Deng et al. developed a prognostic panel using hypoxia-related genes, predicting clinical prognosis and treatment efficacy in GC. Chen et al. devised a prognostic score model based on tumor microenvironment (TME)-related genes, effectively predicting breast cancer patient prognosis and chemotherapy efficacy. Wei et al. employed immune- and cancer-associated fibroblast (CAF)-associated genes (ICRGs) to prognosticate and evaluate immunotherapy efficacy in colorectal adenocarcinoma patients. Hailang et al. identified the gene encoding mitochondrial Aspartyl-tRNA synthetase 2 (DARS2) as a prognostic biomarker in bladder cancer. Dong et al. unveiled the impact of necroptosis-associated myeloid lineages on the immune landscape of pancreatic cancer through scRNA-Seq analysis. Liu et al. conducted LASSO and Cox regression analyses on angiogenesis-related genes (ARGs) in soft-tissue sarcomas (STS) to establish a novel ARG signature (ARSig). Their study demonstrated that ARSig holds promise as an independent prognostic predictor for STS. Li et al. demonstrated that C15orf48, an inflammatory response-related gene, could be a potential biomarker for tumor prognosis and a target for immunotherapy in thyroid cancer. Jiang et al. identified two immunogenic cell death (ICD) subtypes through consensus clustering analysis and constructed an ICD prognostic signature capable of predicting overall survival in patients with renal clear cell carcinoma.
Recent insights underscore the pivotal role of the gut microbiota in the cancer microenvironment and its influence on the efficacy of immunotherapies such as ICIs. Multiple studies have been dedicated to this research area. Zhao et al. reported that enrichment of the gut microbiota, particularly Lachnoclostridium, correlates with the presence of intratumoral tertiary lymphoid structures (TLS) in HCC patients. Gorgulho et al. proposed an immune-microbial score comprising the relative abundance of CD3+HLADR+, NLR, and enterobacteria, which demonstrated predictive capability for therapeutic response to ICIs. Hamada et al. identified bacteria implicated in the efficacy of ICIs and immune-related adverse events (irAEs), suggesting promise for developing a marker to predict cancer immunotherapy efficacy through gut microbiota and fecal transplantation applications.
Several novel and useful biomarkers have emerged from serological methods. Hou et al. identified serum cytokines and the neutrophil-to-lymphocyte ratio as effective biomarkers for predicting the efficacy of ICIs in gastric cancer. Liu et al. introduced an inflammatory prognostic index (InPI) based on three inflammatory markers in patients with relapsed/refractory multiple myeloma (R/R MM) treated with CAR-T therapy, demonstrating its validity as a prognostic biomarker. Raza et al. identified novel immunosuppressive/stimulatory soluble mediators as surrogate and predictive biomarkers of tissue PD-L1 (TPD-L1) status, treatment response, and progression-free survival (PFS) in NSCLC patients treated with ICI.
Many studies have shown that the development of new methods and a multifaceted approach can help in the development of new biomarkers. Ohkuma et al. developed a highly sensitive quantitative immunohistochemical method employing phosphor-integrated dots (PID) for evaluating PD-L1 expression quantitatively. Utilizing this method, they were able to detect PD-L1 expression in the tumors of a subgroup of patients with a favorable prognosis with ICI. Zhang et al. established an alternative splicing (AS) prognostic signature based on AS subtypes in clear cell carcinoma (ccRCC), emphasizing the importance of the AS-SF network, inclusive of splicing factors (SFs), in studying regulatory mechanisms. Yang et al. introduced the CRP-albumin-lymphocyte (CALLY) index, which combines C-reactive protein (CRP), albumin, and lymphocytes, demonstrating its superior prognostic value compared to classical prognostic factors in colorectal cancer patients. Liu et al. introduced a novel biomarker for breast cancer, a nectin-4-specific scFv, with diagnostic and therapeutic applications, recognizing nectin-4 expressed by breast cancer cells in vitro and ex vivo . Zhou et al. identified CD26lowPD-1+ CD8 T cells associated with acute myeloid leukemia (AML) progression and described the prognostic significance of CD26 in AML. Inaba et al. suggested amino acid polymorphisms of HLA class II molecules and HLA-DP5 as genetic predictors of ICI-T1DM in type 1 diabetes induced by ICIs. Wang et al. demonstrated the utility of 18F-fluorodeoxyglucose positron emission tomography (18F-FDG PET) as an imaging biomarker for predicting pathologic response and prognosis in patients with unresectable hepatocellular carcinoma treated with lenvatinib and PD-1 as a conversion therapy.
The compilation of studies in this Research Topic explores various facets of tumor immunology, focusing on identifying novel biomarkers and predictive methods for cancer immunotherapy across diverse cancer types. Researchers employ advanced technologies to uncover promising biomarkers with implications for treatment response and prognosis in cancer patients. Systematic reviews and original papers shed light on the multifaceted landscape of tumor immunology, exploring biomarkers ranging from traditional immunological markers to emerging candidate biomarkers. Notably, investigations extend beyond cancer-specific markers, revealing the involvement of interesting molecules in cancer progression. Moreover, studies elucidate the role of the gut microbiota in modulating the tumor microenvironment and response to immunotherapy, offering insights into potential therapeutic interventions. Serological methods offer valuable biomarkers, while technological advancements, including quantitative immunohistochemical methods, enhance prognostic accuracy. This comprehensive body of work not only highlights the current state of tumor immunology research but also paves the way for future advancements in cancer diagnosis, prognosis, and treatment.
Author contributions
TM: Writing – original draft, Writing – review & editing. EA: Writing – review & editing. EE: Writing – review & editing.
Acknowledgments
We would like to thank all the authors who participated in the Novel biomarkers in tumor immunity and immunotherapy Research Topic, each of the reviewers for their careful peer review and valuable assistance, and the editorial office of the Frontier Journal. We thank Dr. Dennis O Adeegbe for his significant contribution to the conception and editing the proposal of this Research Topic.
Conflict of interest
Author TM is employed by the company Maruho Co., Ltd.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Keywords: biomarker, ICI, bioinformatics, tumor immunology, transcriptome, microbiome
Citation: Matsutani T, Akbay E and Elkord E (2024) Editorial: Novel biomarkers in tumor immunity and immunotherapy. Front. Immunol. 15:1405082. doi: 10.3389/fimmu.2024.1405082
Received: 22 March 2024; Accepted: 27 March 2024; Published: 08 April 2024.
Edited and Reviewed by:
Copyright © 2024 Matsutani, Akbay and Elkord. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY) . The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Takaji Matsutani, [email protected] ; Eyad Elkord, [email protected]
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
- Search Menu
- Advance articles
- Editor's Choice
- Supplement Archive
- Cover Archive
- IDSA Journals
- Clinical Infectious Diseases
- Open Forum Infectious Diseases
- Author Guidelines
- Open Access
- Why Publish
- Advertising and Corporate Services
- Advertising
- Reprints and ePrints
- Sponsored Supplements
- Branded Books
- Journals Career Network
- About The Journal of Infectious Diseases
- About the Infectious Diseases Society of America
- About the HIV Medicine Association
- IDSA COI Policy
- Editorial Board
- Self-Archiving Policy
- For Reviewers
- For Press Offices
- Journals on Oxford Academic
- Books on Oxford Academic

Article Contents
Conclusions.
- < Previous
The Extended Impact of Human Immunodeficiency Virus/AIDS Research
- Article contents
- Figures & tables
- Supplementary Data
Tara A Schwetz, Anthony S Fauci, The Extended Impact of Human Immunodeficiency Virus/AIDS Research, The Journal of Infectious Diseases , Volume 219, Issue 1, 1 January 2019, Pages 6–9, https://doi.org/10.1093/infdis/jiy441
- Permissions Icon Permissions
Human immunodeficiency virus (HIV) is one of the most extensively studied viruses in history, and numerous extraordinary scientific advances, including an in-depth understanding of viral biology, pathogenesis, and life-saving antiretroviral therapies, have resulted from investments in HIV/AIDS research. While the substantial investments in HIV/AIDS research are validated solely on these advances, the collateral broader scientific progress resulting from the support of HIV/AIDS research over the past 30 years is extraordinary as well. The positive impact has ranged from innovations in basic immunology and structural biology to treatments for immune-mediated diseases and cancer and has had an enormous effect on the research and public and global health communities well beyond the field of HIV/AIDS. This article highlights a few select examples of the unanticipated and substantial positive spin-offs of HIV/AIDS research on other scientific areas.
The first cases of AIDS were reported in the United States 37 years ago. Since then, >77 million people have been infected worldwide, resulting in over 35 million deaths. Currently, there are 36.9 million people living with human immunodeficiency virus (HIV), 1.8 million new infections, and nearly 1 million AIDS-related deaths annually [ 1 ]. Billions of research dollars have been invested toward understanding, treating, and preventing HIV infection. The largest funder of HIV/AIDS research is the National Institutes of Health (NIH), investing nearly $69 billion in AIDS research from fiscal years 1982–2018. Despite the staggering disease burden, the scientific advances directly resulting from investments in AIDS research have been extraordinary. HIV is one of the most intensively studied viruses in history, leading to an in-depth understanding of viral biology and pathogenesis. However, the most impressive advances in HIV/AIDS research have come in the arena of antiretroviral therapy. Before the development of these life-saving drugs, AIDS was an almost universally fatal disease. Since the demonstration in 1987 that a single drug, zidovudine, better known as azidothymidine or AZT, could partially and temporarily suppress virus replication [ 2 ], the lives of people living with HIV have been transformed by the current availability of >30 antiretroviral drugs that, when administered in combinations of 3 drugs, now in a single daily pill, suppress the virus to undetectable levels. Today, if a person in their 20s is infected and given a combination of antiretroviral drugs that almost invariably will durably suppress virus to below detectable levels, they can anticipate living an additional 50 years, allowing them almost a normal life expectancy [ 3 ]. In addition, a person receiving antiretroviral therapy with an undetectable viral load will not transmit virus to their uninfected sexual partner. This strategy is referred to as “treatment as prevention” [ 4 ]. Also, administration of a single pill containing 2 antiretroviral drugs taken daily by an at-risk uninfected person decreases the chance of acquiring HIV by >95%. Finally, major strides are being made in the quest for a safe and effective HIV vaccine [ 5 ].
The enormous investment in HIV research is clearly justified and validated purely on the basis of advances specifically related to HIV/AIDS. However, the collateral advantages of this investment above and beyond HIV/AIDS have been profound, leading to insights and concrete advances in separate, diverse, and unrelated fields of biomedical research and medicine. In the current Perspective, we discuss a few select examples of the positive spin-offs of HIV/AIDS research on other scientific areas ( Table 1 ).
Positive Spin-offs of Human Immuno deficiency Virus/AIDS Research on Other Areas of Medicine
Abbreviation: HIV, human immunodeficiency virus.
Regulation of the Human Immune System
Congenital immunodeficiencies have been described as “experiments of nature,” whereby a specific defect in a single component of the complex immune system sheds light on the entire system. Such is the case with AIDS, an acquired defect in the immune system whereby HIV specifically and selectively infects and destroys the CD4 + subset of T lymphocytes [ 6 ]. In this respect, HIV infection functions as a natural experiment that elucidates the complexity of the human immune system. The selectivity of this defect and its resulting catastrophic effect on host defense mechanisms, as manifested by the wide range of opportunistic infections and neoplasms, underscore the critical role this cell type plays in the overall regulation of the human immune system. This has provided substantial insights into the pathogenesis of an array of other diseases characterized by aberrancies of immune regulation. Additionally, the in-depth study of immune dysfunction in HIV disease has shed light on the role of the immune system in surveillance against a variety of neoplastic diseases, such as non-Hodgkin lymphoma and Kaposi sarcoma. As a result of its association with HIV/AIDS, Kaposi sarcoma was discovered to be caused by human herpesvirus 8 [ 7 ].
Targeted Antiviral Drug Development
Targeted antiviral drug development did not begin with HIV infection. However, the enormous investments in biomedical research supported by the NIH and in drug development supported by pharmaceutical companies led to highly effective antiretroviral drugs targeting the enzymes reverse transcriptase, protease, and integrase, among other vulnerable points in the HIV replication cycle, and have transformed the field of targeted drug development, bringing it to an unprecedented level of sophistication. Building on 3 decades of experience, this HIV model has been applied in the successful development of antiviral drugs for other viral diseases, including the highly effective and curative direct-acting antivirals for hepatitis C [ 8 ].
Probing the B-Cell Repertoire
The past decade has witnessed extraordinary advances in probing the human B-cell lineage resulting from the availability of highly sophisticated technologies in cellular cloning and genomic sequencing [ 9 ]. AIDS research aimed at developing broadly reactive neutralizing antibodies against HIV and an HIV vaccine that could induce broadly neutralizing antibodies has greatly advanced the field of interrogation of human B-cell lineages, leading to greater insights into the humoral response to other infectious diseases, including Ebola [ 10 ], Zika [ 11 ], and influenza [ 12 ], as well as a range of autoimmune, neoplastic, and other noncommunicable diseases [ 13 ].
Structure-Based Vaccine Design
Although a safe and effective HIV vaccine has not yet been developed, the discipline of structure-based vaccine design using protein X-ray crystallography and cryoelectron microscopy has matured greatly in the context of HIV vaccine research. The design of immunogens based on the precise conformation of epitopes in the viral envelope as they bind to neutralizing antibodies has been perfected within the arena of HIV vaccine immunogen design. This has had immediate positive spinoffs in the design of vaccines for other viruses, such as respiratory syncytial virus, in which the prefusion glycoprotein was identified as the important immunogen for a vaccine using structure-based approaches [ 14 ].
Advances in HIV/AIDS-Related Technologies
Insights into the basic immunology of HIV drove the development and optimization of several broadly applicable technologies. Using inactivated HIV as a means of altering T lymphocytes to modulate the immune response, safe lentiviral gene therapy vectors are now US Food and Drug Administration–approved to treat certain cancers (eg, acute lymphoblastic leukemia) [ 15 ]. Additionally, it was discovered early in the epidemic that HIV is associated with the loss of CD4 + T lymphocytes [ 16 ]. While much of the initial research on CD4 + T lymphocytes was possible due to existing flow cytometry technologies, probing the complexities of immune dysregulation in HIV infection spurred the development of multicolor cytofluorometric technologies that have proven extremely useful for studying a variety of other diseases characterized by immune dysfunction [ 17 ]. The reality of utilizing these technologies in resource-poor areas accelerated the advancement of new simplified, automated, affordable, and portable point-of-care devices with broader implications for clinical medicine [ 18 ].
Role of Immune Activation in Disease Pathogenesis
Studying the pathogenesis of HIV disease has clearly demonstrated that aberrant immune activation stimulated by virus replication is the driving force of HIV replication [ 19 ]. In essence, the somewhat paradoxical situation exists whereby the very immune activation triggered by the virus in an attempt to control virus replication creates the microenvironment where the virus efficiently replicates. Even when the virus is effectively suppressed by antiretroviral drugs, a low degree of immune activation persists [ 20 ]. In this regard, the flagrant immune activation associated with uncontrolled virus replication, as well as the subtle immune activation associated with control of virus replication, are important pathogenic triggers of the increased cardiovascular and other organ system diseases associated with HIV infection. This direct association of even subtle levels of immune activation seen in HIV infection with a variety of systemic diseases has led to considerable insight into the role of immune activation and inflammation in human disease [ 21 ]. For example, recognition of the increased incidence of heart disease in the HIV population that is associated with chronic inflammation has stimulated interdisciplinary advances in understanding and treating coronary heart disease apart from HIV infection [ 22 ].
Comorbidities in HIV Disease
Antiretroviral therapy, which has transformed HIV treatment, is shifting the incidence of certain diseases in people living with HIV. Even when well-controlled by antiretrovirals, HIV disease is associated with an increased incidence of diseases, such as cardiovascular disease, kidney and liver disease, the premature appearance of pathophysiologic processes associated with aging, and several cancers [ 21–24 ]. This is especially true for non-AIDS-defining cancers, whose incidence rates are increasing while AIDS-defining cancer rates are decreasing [ 24 ]. In lower-income countries, tuberculosis is a common coinfection with HIV, and HIV coinfection was shown to be a key risk factor for progression of latent Mycobacterium tuberculosis infection to active disease [ 25 ]. There are a variety of ongoing studies [ 21 ] investigating the pathogenic bases of these conditions to shed greater insight into their causes and potential interventions that might impact these diseases apart from HIV infection and immunodeficiency.
The collateral advantages resulting from the substantial resources devoted to HIV/AIDS research over the past 30 years are extraordinary. From innovations in basic immunology and structural biology to treatments for immune-mediated diseases and cancer, the conceptual and technological advances resulting from HIV/AIDS research have had an enormous impact on the research and public and global health communities over and above the field of HIV/AIDS. The HIV/AIDS research model has proven that cross-fertilization of ideas, innovation, and research progress can lead to unforeseen and substantial advantages for a variety of other diseases.
Acknowledgments. The authors thank Carl Dieffenbach, Daniel Rotrosen, Charles Hackett, and Robert Eisinger for their helpful input in preparation of the manuscript.
Potential conflicts of interest. Both authors: No reported conflicts of interest. Both authors have submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest. Conflicts that the editors consider relevant to the content of the manuscript have been disclosed.
Joint United Nations Progamme on HIV/AIDS . Fact sheet: latest statistics on the status of the AIDS epidemic . http://www.unaids.org/en/resources/fact-sheet . Accessed 23 July 2018.
Fischl MA , Richman DD , Grieco MH , et al. The efficacy of azidothymidine (AZT) in the treatment of patients with AIDS and AIDS-related complex. A double-blind, placebo-controlled trial . N Engl J Med 1987 ; 317 : 185 – 91 .
Google Scholar
Samji H , Cescon A , Hogg RS , et al. North American AIDS Cohort Collaboration on Research and Design (NA-ACCORD) of IeDEA . Closing the gap: increases in life expectancy among treated HIV-positive individuals in the United States and Canada . PLoS One 2013 ; 8 : e81355 .
Lundgren JD , Babiker AG , Gordin F , et al. INSIGHT START Study Group . Initiation of antiretroviral therapy in early asymptomatic HIV infection . N Engl J Med 2015 ; 373 : 795 – 807 .
Trovato M , D’Apice L , Prisco A , De Berardinis P . HIV vaccination: a roadmap among advancements and concerns . Int J Mol Sci 2018 ; 19 . doi: 10.3390/ijms19041241 .
Dalgleish AG , Beverley PC , Clapham PR , Crawford DH , Greaves MF , Weiss RA . The CD4 (T4) antigen is an essential component of the receptor for the AIDS retrovirus . Nature 1984 ; 312 : 763 – 7 .
Schulz TF , Boshoff CH , Weiss RA . HIV infection and neoplasia . Lancet 1996 ; 348 : 587 – 91 .
Wyles DL . Antiviral resistance and the future landscape of hepatitis C virus infection therapy . J Infect Dis 2013 ; 207 ( Suppl 1 ): S33 – 9 .
Boyd SD , Crowe JE Jr . Deep sequencing and human antibody repertoire analysis . Curr Opin Immunol 2016 ; 40 : 103 – 9 .
Flyak AI , Kuzmina N , Murin CD , et al. Broadly neutralizing antibodies from human survivors target a conserved site in the Ebola virus glycoprotein HR2-MPER region . Nat Microbiol 2018 ; 3 : 670 – 7 .
Sapparapu G , Fernandez E , Kose N , et al. Neutralizing human antibodies prevent Zika virus replication and fetal disease in mice . Nature 2016 ; 540 : 443 – 7 .
Raymond DD , Bajic G , Ferdman J , et al. Conserved epitope on influenza-virus hemagglutinin head defined by a vaccine-induced antibody . Proc Natl Acad Sci U S A 2018 ; 115 : 168 – 73 .
Röhn TA , Bachmann MF . Vaccines against non-communicable diseases . Curr Opin Immunol 2010 ; 22 : 391 – 6 .
Tian D , Battles MB , Moin SM , et al. Structural basis of respiratory syncytial virus subtype-dependent neutralization by an antibody targeting the fusion glycoprotein . Nat Commun 2017 ; 8 : 1877 .
US Food and Drug Administration . FDA approval brings first gene therapy to the United States . Silver Spring, MD : FDA , 2017 .
Google Preview
Gottlieb MS , Schroff R , Schanker HM , et al. Pneumocystis carinii pneumonia and mucosal candidiasis in previously healthy homosexual men: evidence of a new acquired cellular immunodeficiency . N Engl J Med 1981 ; 305 : 1425 – 31 .
Chattopadhyay PK , Roederer M . Cytometry: today’s technology and tomorrow’s horizons . Methods 2012 ; 57 : 251 – 8 .
Kestens L , Mandy F . Thirty-five years of CD4 T-cell counting in HIV infection: from flow cytometry in the lab to point-of-care testing in the field . Cytometry B Clin Cytom 2017 ; 92 : 437 – 44 .
Moir S , Fauci AS . B-cell exhaustion in HIV infection: the role of immune activation . Curr Opin HIV AIDS 2014 ; 9 : 472 – 7 .
Paiardini M , Müller-Trutwin M . HIV-associated chronic immune activation . Immunol Rev 2013 ; 254 : 78 – 101 .
Lucas S , Nelson AM . HIV and the spectrum of human disease . J Pathol 2015 ; 235 : 229 – 41 .
Boccara F , Lang S , Meuleman C , et al. HIV and coronary heart disease: time for a better understanding . J Am Coll Cardiol 2013 ; 61 : 511 – 23 .
Torres RA , Lewis W . Aging and HIV/AIDS: pathogenetic role of therapeutic side effects . Lab Invest 2014 ; 94 : 120 – 8 .
Thrift AP , Chiao EY . Are non-HIV malignancies increased in the HIV-infected population ? Curr Infect Dis Rep 2018 ; 20 : 22 .
Getahun H , Gunneberg C , Granich R , Nunn P . HIV infection-associated tuberculosis: the epidemiology and the response . Clin Infect Dis 2010 ; 50 ( Suppl 3 ): S201 – 7 .
- acquired immunodeficiency syndrome
Email alerts
Related articles in pubmed, citing articles via, looking for your next opportunity.
- Recommend to your Library
Affiliations
- Online ISSN 1537-6613
- Print ISSN 0022-1899
- Copyright © 2024 Infectious Diseases Society of America
- About Oxford Academic
- Publish journals with us
- University press partners
- What we publish
- New features
- Open access
- Institutional account management
- Rights and permissions
- Get help with access
- Accessibility
- Media enquiries
- Oxford University Press
- Oxford Languages
- University of Oxford
Oxford University Press is a department of the University of Oxford. It furthers the University's objective of excellence in research, scholarship, and education by publishing worldwide
- Copyright © 2024 Oxford University Press
- Cookie settings
- Cookie policy
- Privacy policy
- Legal notice
This Feature Is Available To Subscribers Only
Sign In or Create an Account
This PDF is available to Subscribers Only
For full access to this pdf, sign in to an existing account, or purchase an annual subscription.
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- Immunotargets Ther
Immunoinformatics and Vaccine Development: An Overview
Angus nnamdi oli.
1 Department of Pharmaceutical Microbiology and Biotechnology, Faculty of Pharmaceutical Sciences, Nnamdi Azikiwe University, Awka, Nigeria
Wilson Okechukwu Obialor
Martins ositadimma ifeanyichukwu.
2 Department of Immunology, College of Health Sciences, Faculty of Medicine, Nnamdi Azikiwe University, Anambra, Nigeria
3 Department of Medical Laboratory Science,Faculty of Health Science and Technology, College of Health Sciences, Nnamdi Azikiwe University,Nnewi Campus, Nnewi, Nigeria
Damian Chukwu Odimegwu
4 Department of Pharmaceutical Microbiology and Biotechnology, Faculty of Pharmaceutical Sciences, University of Nigeria Nsukka, Enugu, Nigeria
Jude Nnaemeka Okoyeh
5 Department of Biology and Clinical Laboratory Science, Division of Arts and Sciences, Neumann University, Aston, PA, 19014-1298, USA
George Ogonna Emechebe
6 Department of Pediatrics, Faculty of Clinical Medicine, Chukwuemeka Odumegwu Ojukwu University, Awka, Nigeria
Samson Adedeji Adejumo
Gordon c ibeanu.
7 Department of Pharmaceutical Science, North Carolina Central University, Durham, NC, 27707, USA
Associated Data
All the needed data are included in this manuscript.
The use of vaccines have resulted in a remarkable improvement in global health. It has saved several lives, reduced treatment costs and raised the quality of animal and human lives. Current traditional vaccines came empirically with either vague or completely no knowledge of how they modulate our immune system. Even at the face of potential vaccine design advance, immune-related concerns (as seen with specific vulnerable populations, cases of emerging/re-emerging infectious disease, pathogens with complex lifecycle and antigenic variability, need for personalized vaccinations, and concerns for vaccines' immunological safety -specifically vaccine likelihood to trigger non-antigen-specific responses that may cause autoimmunity and vaccine allergy) are being raised. And these concerns have driven immunologists toward research for a better approach to vaccine design that will consider these challenges. Currently, immunoinformatics has paved the way for a better understanding of some infectious disease pathogenesis, diagnosis, immune system response and computational vaccinology. The importance of this immunoinformatics in the study of infectious diseases is diverse in terms of computational approaches used, but is united by common qualities related to host–pathogen relationship. Bioinformatics methods are also used to assign functions to uncharacterized genes which can be targeted as a candidate in vaccine design and can be a better approach toward the inclusion of women that are pregnant into vaccine trials and programs. The essence of this review is to give insight into the need to focus on novel computational, experimental and computation-driven experimental approaches for studying of host–pathogen interactions and thus making a case for its use in vaccine development.
Introduction
Vaccination has been undeniably very helpful in promoting a healthy global population. It has severally saved lives, reduced healthcare costs and raised man’s quality of life. 1 It greatly reduces disease burden, disability and death. However, newly emerging and re-emerging infectious diseases (ERID), infectious agents with complex lifecycle and antigenic variability and the need for personalized vaccination present additional challenges in vaccine development. 2 , 3
For many pathogens (especially the emerging and those with antigenic variability), their genomes are known but their immune correlates of protection remain unclear. 1 , 4 Some of these reasons are why vaccine development for ERID and multi-lifecycle pathogenic diseases is a tall order.
Serendipitous discoveries in immunology coupled with knowledge of bioinformatics tools for epitope predictions have resulted in the emergence of new pattern of vaccine design. 5 , 6 The art and science of efficient and comprehensive information extraction and analysis of data deposited in relevant databases is now increasingly essential in researches related to immunology. 7 Even with this capacity (efficient information extraction), some challenges in the application of bioinformatics in immunology include structure and/or function analysis and immune process analyses as concern the immune interaction specificity. Fortunately, although researches in immunology are experimentally costly and very intensive, colossal amounts of data are usually generated. Such data can only be analyzed with high precision and speed using bioinformatics tools. For instance, genome sequencing as well as in vitro T-cell confirmation is done in few months as opposed to years using the conventional vaccine design. 8 Also, computational immunological methods drastically reduce both time and labor needs in epitopes screening. 5 , 9 With computational immunology techniques, it is possible to discover vaccine candidate epitopes simply by scanning the protein sequences in a pathogen of interest. 5 Many of these proteins are yet to be isolated or at least cloned. Being pathogens specific and unique, they present ready candidates in vaccine construct.
This review describes the need to use immunoinformatics-based techniques to unveil vital determinants of immunity made available in the genome sequence database and design vaccines. Also, this review gives insight into the need to focus on novel computational, experimental and computation-driven experimental approaches for studying of host–pathogen interactions and thus making a case for its use in vaccine development. This review will further show the need for new approaches for effective drugs or vaccine design so as to combat the antigenic variability of some pathogens.
Vaccine Immunology and Design
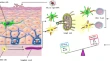
The process of generating vaccine-induced immunity is somewhat challenging in immunology. Current conventional vaccines came empirically when there were vague or no knowledge of vaccine immune system activation. A lot of research 10 – 12 has been geared toward the understanding of this challenge, but the complexity of it requires a different dimension of approach. 13 An approach that must accommodate many factors affecting vaccine development like pathogen antigenic variability, the emergence of infectious disease, human genetic variation is the goal of immunoinformatics [ Figure 1 ].

Schematic illustration of the cases stemming the need for immunoinformatics vaccine development approach.
Activation of the immune system involves, among many processes, induction of the immune memory. The strength of this induction determines the efficacy of a vaccine. Hence, vaccine efficacy in the long run is influenced by the determinants of immunological memory stimulation, persisting antibodies and kind and type of immune memory cells induced. 14
The primary vaccine-mediated immunological effectors ( Table 1 ) are mainly the antibodies (from B lymphocytes/cell) 15 , 16 and sometimes CD8+ and CD4+ T cells. These antibodies bind specifically to a particular kind of toxin or pathogen. Vaccines and most antigens evoke humoral as well as cell-mediated immune responses. 17 Vaccines that mediate immune responses through these systems (B and T cell responses) are said to be more effective. Although B cells are regarded as the primary vaccine immune effectors, T cells induce immune memory cells and high-affinity antibodies. Studies in reverse vaccinology and immunomics had also proved T cells as prime immune effectors following the discovery of novel vaccine targets with EpiMatrix. 18 – 20 This change of immune target has led to successful advances in vaccine design.
Functions of the Immunological Memories
Even at the face of potential vaccine design advances, immune-related questions are now focused on specific vulnerable populations such as the young, elderly and immunocompromised. 21 , 22 These concerns have propelled a better understanding of the efficacy of current vaccines on this vulnerable population and have also paved way for the application of new approaches that can put into consideration the differences of population and better targets that can generate optimum immune induction 23 – 25 with the exception of type II T-cell-independent (TI-2) antigens (i.e., polysaccharide antigens).
Antigens that could provoke the B lymphocytes as well as the T lymphocyte responses stimulate the germinal centers causing antigen-specific highly efficient B-cell multiplication and eventual differentiation into antibody-forming plasma cells and memory B cells. All existing protein and DNA antigens induce immunological memory B cells unlike type II T-cell-independent (TI-2) antigens (i.e., polysaccharide antigens). These polysaccharide antigens do not generate memory B cells but can induce long-lasting humoral immunity even when recall responses are lacking. 26 Vaccine efficacy may be short term 27 if only the B cells are activated.
The traditional approach for developing vaccines for infectious disease threats has shifted to include other vaccine design techniques like cloning and expressing major surface antigens 28 although this frequently resulted in the formulation of vaccines with poor immunogenicity, requiring strong adjuvantation. 29 This approach is particularly likely to be less specific for pathogens with complex lifecycles (e.g., parasites) or very high mutability (e.g., RNA viruses). These pathogens do not depend on one route for their virulence of pathogenesis in human and thus to alter this process, increasing the specificity of the vaccine should be the aim and not just the effectiveness as seen in the current conventional vaccines. 28 , 29
Vaccines for several neglected tropical diseases are in various stages of development, 30 thanks to mega drug companies that have continued to demonstrate a willingness to invest money in the research and development as regards to diseases plaguing the developing nations. 16 , 30 - 32 It is very pertinent to invest in researches that have an interest in vaccine specificity on the pathogen antigens than totally on the efficacy. Fortunately, with global research efforts, the genomes for several pathogens causing neglected tropical diseases and several emerging pathogens are becoming available. 33 Computational vaccinology may now be applied to screen these genomes for possible vaccine target. With these tools, many proteins of virulence interest can be sequenced and the most essential gene of interest modeled for a potential vaccine candidate specific for that pathogen. Immunoinformatics is the way forward in the identification of vaccine candidates for these tropical ERID, for pathogens with varying antigens and for individualized therapy.
Immunoinformatics and Infectious Disease
Immunology studies produce data in colossal quantities. Also, with proteomics and genomics projects, extensive screening of pathogens and/or pathogen–host interaction, it has become increasingly necessary to store, manage and analyze these data, hence the birth of immunoinformatics. Immunoinformatics deals with computational techniques and resources used to study the immune functions. Statistical, computational, mathematical and biological knowledge and tools are applied in immunoinformatics in order to accurately and specifically store, and analyze data concerning the immune system and its functions.
To handle evidence diversity, immunoinformatics uses tools that cut across several aspects of bioinformatics such as creation and management of databases, 34 , 35 use and definition of both structural and functional signatures and the formation and application of predictive tools. 35 – 37 These strategies can synergize toward a better understanding of the immune system of both man and animals and fight against some less predictable pathogenesis. The complex nature of vertebrates’ immune system, the variable nature of pathogens and environmental antigens coupled with the multi-regulatory pathways show that colossal quantities of data will be needed to unveil how the human immune systems work. Conventionally, much cannot be achieved based on the complexity of the immune system and the virulent antigen but with the application of computational vaccinology, researches on vaccine design have been made easier, accurate and specific. Applying immunoinformatics in disease study ( Table 2 ) requires the knowledge of disease pathogenesis, the immune system dynamics, and computational vaccinology, painstaking searches of the database, sequence comparison, structural modeling as well as motif analysis. 35 , 38 These methods can go a long way in analyzing the pathogenesis of a disease and identification of vaccine candidates.
The Importance of Bioinformatics in the Research on Infectious Diseases
In order to help understand complex pathogen-related processes, computational models were developed for viral 46 , 47 bacterial, 48 parasitic 49 and fungal pathogens. 50
The bioinformatics tools ( Table 3 ) are used to identify possible epitopes for vaccine formulation. Each tool can screen protein sequences and identify aggregates of MHC binding and supertype motifs for possible use in epitope-based vaccine development and for use among human populations with genetic variability.
Bioinformatics Tools for MHC Cluster Binding and Super-Type Motifs and Protein Sequences
There are several databases ( Table 4 ) that can provide a wide range of information for all forms of immunological studies. Generated data from the studies are further organized and stored in the databases ( Table 4 ) to provide a means for the development and advance in immunological research. A tour on these databases will actually stimulate some interest in the vaccinology of emerging and re-surging disorders attributable to pathogen including cancer.
List of Some Immunological Databases
Case of Emerging and Re-Surging Disorders/Diseases Attributable to Pathogens
Emerging infections (EIs) include infections that are entirely new in a population or that may have existed before in the population but are now gaining rapid and continued spread and/or wide geographical range. Re-emerging or re-surging infections represent the infections that were previously of historical relevance but are now quickly becoming relevant because of either increasing incidence or increasing geographical and/or human host range while emerging infections represent the infections that were not originally observed in man. 66 Several factors such as human behavioral changes, environmental changes, and host/intermediate factors, animal-human switching and microbial genetic changes all affect infectious disease emergence and spread. 67 These factors interact to promote the evolution of pathogens into new ecosystems, infect, spread and thrive in their new hosts. 68 The overall consequences of these are continued infectious disease emergence and re-emergence, epidemics and public health challenges. Emerging infections and multi-antibiotic-resistant strains of pathogenic bacteria usually surge from one geographic location from where it spreads to other places due to immigration of people. 67 , 69 Most emerging infections originate from a specific population and can spread to a new population or become selectively advantaged that it can lead to the emergence of new strains of the pathogen. 67 , 70 , 71 Also, there could be microbial traffic, in which case, an infectious agent transfers from animals to humans or spreads from isolated groups to new populations. 67 , 71 , 72
Several factors, including ecology, are known to be associated with infectious disease outbreaks. These factors bring man into close contact with a natural disease reservoir/host. 70 With an increasing world population and poor infection control, the emergence of infection and increased microbial populations are sure. The human growth population will only increase the spread of the infection across populations. The information provided in Table 5 is the list of remerging infections and current emerging diseases put forward during the WHO 2018 annual review. 73 The review noted that these infections, if not well controlled, can cause disease outbreaks, bioterrorism and similar occurrences requiring urgent public health attention and that with the dearth of efficacious medicines or vaccines, there is a compelling demand for continuous as well as accelerated research and development in those areas.
Current Emerging and Re-Surging Infectious Disease
Advances in Genomics, proteomics, immunomics, vaccinomics and nanotechnology are being continually exploited in diagnostic, therapeutic and in rational drug and vaccine development. These advances have also served in the control of the afore-mentioned emergences. 74 , 75 The knowledge of the emerging pathogen’s genome, protein make-up, pathogen-immune system interactions and researching the possible therapeutics will go a long way in directing the optimum path to containing the infection spread and controlling potential re-emergence or emergence in a different population. Approaches in direct and computer-based structural determinations, 76 protein–protein interactions predicting, and bioinformatics tools now exist and are used in modern-day development of drug and biologics. 77
Vaccine development has been sped up through the advance in the knowledge of the immune system of man. Researches in the traditional targets of vaccines (adaptive immune response) and the less specific and fast-acting innate immune responses have been clear evidences for this advance. 78 – 80 As our understanding of the intercourse between innate and adaptive immunity increases, reasons and opportunities for more effective vaccine adjuvants will open up. This can be a step forward in solving a critical world’s health challenge per population. Following the conventional approach of vaccine design, much cannot be achieved but when the knowledge of immunoinformatics is applied, population safety and disease control can be achieved through pathogen’s genome sequencing leading to optimum new vaccine design or development of a novel vaccine for the infection.
Pathogens with Variable Antigens
Antigenic variability is an important mechanism pathogens use to evade their host immunity. The surface proteins of pathogens are normally variable. This assists them to escape recognition by the immune system. A successful infectious agent presents to the host immune system information that differs from that of its virulence. Pathogenic organisms have organized systems of escaping destruction by the immune system of their hosts. For instance, Toxoplasma invades and appropriates the host cells thereby circumventing phagocytosis and then spread within their host to establish infection. 81 Vertebrates on their own are endowed with immune system robust enough to efficiently and effectively surmount the non-self-attacks. Yet the more the host’s immune system elaborates, the better the organisms in their evasion of immune effector cells.
Antigenic variation refers to a pathogen’s ability to modify its surface proteins such that it can circumvent the host’s immunological attacks. It involves several mechanisms including the varying of surface protein’s phase, shifting and drifting of surface protein antigens and/or any other form of alteration of antigenic protein. 82 Antigenic variation plays significant roles in the pathogenicity of microorganisms by evasion of the host immune responses and establishment of re-infection. When a pathogen alters its surface antigens, it can evade the host’s adaptive immunity and so re-establishes infection. The immune system may battle to generate new immunoglobulins against the new antigen. Certain bacteria like Neisseria gonorrhea, Neisseria meningitides, Mycoplasma and species of the genus Streptococcus show antigenic diversity. 83 In eukaryotic pathogens, antigenic variation is shown by Trypanosoma brucei and Plasmodium falciparum . 81 , 84
Another vital cause of antigenic variation in bacteria is horizontal gene transfer (more important than point mutation) through plasmid acquisition and transduction via bacteriophages. Virulence genes are normally acquired by non-virulent organisms via these routes. Once this occurs, the new bacteria may quickly get established and cause fresh epidemic outbreak.
Species of the genus Neisseria are champions in the rapid change of surface antigens amongst bacterial pathogens. Pathogenic forms exhibit an amount of phenotypic variability not found in the commensal species. The pathogenic forms are implicated in STD and meningitis. They employ amazing varieties of antigenic variability mechanisms.
- They can recombine their pilin genes in a similar manner that eukaryotes recombine their own genes, such that they can express variable surface protein. 85
- Some cell-surface proteins and enzymes synthesizing bacterial cell-surface carbohydrates are expressed in a variety of ways. This is as a result of replication slipping or slippage errors and repairs of simple tandem nucleotide repeats involving either the di-, or tri- or tetra-nucleotides. 86
- Neisseria is able to take up and incorporate environmental DNA into its genomes. 83 , 87
- Again, the chances of Neisseria acquiring new mutations by replication errors are high compared to other bacteria like E. coli . 88
These are why an effective vaccine against Neisseria infections is not yet developed. Neisseria may be considered as an extreme example. However, many other bacterial pathogens like Streptococcus and Mycoplasma in promoting their antigenic variation tend to utilize one or more of these techniques. Additionally, there are reports that DNA-related defects have a much greater association with bacterial pathogen from symptomatic patients than samples of the same bacterial species isolated from environmental sources. 87 – 89
Pneumococcus
Streptococcus pneumoniae , Gram-positive cocci bacteria that cause otitis media, bacteremia and pneumonia, are a public health concern, causing morbidity and mortality in adults and children. 90 Two forms of vaccines (polysaccharide and conjugate vaccines) are currently marketed for the prevention of pneumococcal infections. While the polysaccharide vaccines are for vaccination in the adult population, conjugate vaccines have an added immunogenic non-pneumococcal protein conjugated to the pneumococcal polysaccharides for enhanced immunogenicity in children. It is not yet known that these vaccines can evoke complete immunity against the infection.
A polysaccharide capsule is a major virulence factor in the bacteria. Several of these capsule types have been identified, and these form the basis of pathogen’s antigenic serotyping. 91 , 92 Current pneumococcal vaccines are combinations of various capsular (polysaccharide) antigens from the serotypes most common in a particular population. Currently, over 100 different serotypes are known but are not all covered in the available vaccines. 92 The discovery of a common antigen(s) will produce an effective vaccine. Knowledge of the genome of the organism and the different strains has led to a possible advance in driving the pneumococcal potential vaccine search through a different approach. And this consideration will help solve a lot of concerns about the current vaccines. With this knowledge, many methods are been tried to determine whether they can be a source of effective vaccine design that can accommodate all the serotypes of the organism. Search for antigen that is common to all the serotypes can be achieved with the knowledge of Genomics and immunoinformatics. The introduction of genomic and computational technologies has given new directions in the study of bacterial pathogenesis and vaccine design. 93 , 94
Plasmodium falciparum undergoes two life cycles: one in humans and the other in mosquitoes. The human host’s erythrocytes and hepatocytes usually display modified parasite proteins called Plasmodium falciparum erythrocyte membrane protein 1 (PfEMP1) and Plasmodium falciparum hepatocyte membrane protein 1 (PfHMP1), respectively. These proteins function to assist the parasite to evade destruction by the host immune systems. 95 , 96 The PfEMP1 proteins were identified as the prime ligands responsible for cytoadherence and resetting. 97 They cause the infected RBCs of host tissues to sequester thus helping the parasite to circumvent clearance by the host’s spleen. 98 The membrane proteins also shield infected host cells from destruction by the spleen by adhering to the endothelium. Luckily, if the PfEMP1 protein is expressed for a long while; it comes under attack by the naturally acquired immunity. 98 , 99 In defence to this, the parasite has expanded the var genes coding for PfEMP1 such that the genes can exist as a polymorphic family of as much as over 50 members in every genomic haploid. Antigenic switches work well here in that members of the polymorphic family (also called antigenic-variant-protein family) can be interchanged and cannot express their proteins at the same time. In this way, only one particular protein at the surface of the infected RBC is expressed at any given time. 97 , 100
When studying antigenic-variant-protein families, it is pertinent to understand if grouping them into single-family results in any meaningful antigenic activity. Studies have tried to understand the “languages” of the antigenic variant of PfEMP1 proteins. 97 , 101 , 102 They sought to know the PfEMP1 proteins binding properties or search to understand the correlation between motifs and infection severity.
The varDB database is a repository for protein sequences involved in antigenic variation and their associated functionalities. 103 Antigenic variant data obtained from several pathogens may be regrouped into a unified database. This will enable researches from several multi-copy gene families to be accessed and compared swiftly in a single moment. Updated varDB database contains close to 10,000 DNA sequences, several protein translations, tens of infectious diseases and pathogens with their gene families. With a novel sequencing-based approach, PacBio, the different PfEMP1 proteins can be sequenced and the related sequences used as potential vaccine targets. 104 , 105
Trypanosoma
For many pathogens, antigenic variability occurs during the infection pathogenesis and is to enable them to escape destruction by the host antibodies. For instance, some eukaryotic parasites take to genetic assortment and re-arrangement thereby changing their surface antigens. A ready example is seen in Trypanosoma brucei , the causative organism for sleeping sickness. Trypanosoma brucei replicates in the bloodstream (outside the cells) of their host, but at maturity, it crosses the blood–brain barrier to cause several fatal complications. During replication in the bloodstream, the parasites are subjected to humoral as well as cellular immune responses. It evades the host defenses by encasing itself in homogeneous coat of glycoprotein called the variant surface glycoprotein (VSG). 106 , 107 Though at initial invasion, the protein coat tends to protect the microbe from the immune system but on constant exposure, the coat will be identified as a foreign matter, and at this point the immune effectors can launch an attack against it. In a particular Trypanosoma brucei , there are diversities of the VSG protein being coded by more than a thousand genes in the parasite’s genome. Unfortunately for the host, the expression of these genes is mutually exclusive. Expressed VSG gene is normally due to genetic re-assortments causing new alleles to be copied into the sites of expression. Some trypanosomes with these abnormal VSG genes evade humoral immunity and multiply thereby causing re-infection and chronic recurring infections. 107
Influenza Virus
Influenza is a viral infectious disease due to infection by any of the three types of RNA viruses, namely influenza Types A, B and C. Current vaccines contain double Type A and single Type B strains and induce strong antibody responses to neuraminidase and the surface glycoprotein hemagglutinin. These vaccines, however, cannot effectively protect against newly emerging viruses with antigenic shift and drift. 108 , 109
Antigenic drift results in changes in the antigenic site (a minor change) while antigenic shift results in a new virus subtype. Hemagglutinin and neuraminidase are the two enzymes dictating the antigenic properties of the viruses. While inside its host, defined host proteases break the peptide bonds in the hemagglutinin molecule to form hemagglutinin 1 and 2 subunits. Virulence tendencies are decreased when the amino acids at the cleavage sites are lipophobic, the virus exhibits high virulence tendencies. 110 The surface glycoprotein can be regarded as antigen and hence can serve as a target for the immune system which if sequenced, using the new immunoinformatics approach and a common site for the varying proteins identified, a potent vaccine can be developed which can accommodate the antigenic drift/shifts of the virus.
Influenza viruses are able to thrive for a long while in a given human population. 111 , 112 The virus has a high mutation rate such that a once effective vaccine can easily lose efficacy. Antigenic variability is only one of the evidences of phenotypic variation in the biology of the Influenza virus.
Multiepitope Vaccines
The use of immunoinformatics in vaccine development has been accelerated toward the design of a multiepitope vaccine construct which has and will fully address the challenges faced with pathogens with mutagenic antigens. Previous vaccines developed by conventional approaches consist of several proteins or a whole pathogen. This constitutes unwarranted antigenic load and increases the chances of inducing allergy. The use of peptide-based vaccines surmounts these challenges. The vaccines are made from short peptide fragments capable of eliciting highly specific immune responses, precision targeting and multiepitope constructs, in the case of varying antigenic peptides, which has been made feasible with the advancements in the field of computational biology. 113 – 116
Vaccines for pathogens with immune escape potentials can basically be constructed by using most, if not all, of their immunogenic peptides 116 , 117 because such vaccines prove to be better than single-epitope and classical vaccines. Multiepitope vaccines enjoy the following advantages over single-epitope and classical vaccines: a) they are an assemblage of several epitopes obtained from distinct protein targets/antigens of an intended infection; b) the multiple T-cell receptors (TCRs) in the vaccine recipient can easily recognize vaccines with multiple HLA epitopes; c); they can be easily adjuvanted to improve their immunogenicity; d) they can activate antibody-mediated and cell-mediated immunological responses because of their overlapping helper T lymphocytes (HTL), CD8+ T-cell and B-cell epitopes; and e) unwanted protein antigens are excluded in such construct thereby reducing the chances of untoward effects and/or immune responses likely to cause disease(s). 118 – 123 Thus producing a vaccine with these qualities can provide chances of combating most infections such as Streptococcus pneumoniae and HIV infections.
Immunoinformatics can be employed in the docking of single and multiepitope vaccines and subsequently to predict their properties (physicochemical, allergenic and antigenic). This approach has seen the use of diverse tools and database in the analysis of ligands with their targets and has greatly helped to predict the binding score of antigenic peptides with the immune proteins like HLA. Peptides and HLA allele modeling can be done by the 3D structural designing of the epitopes using PEPFOLD3 (an online server), 124 retrieving from Protein Data Bank (PDB) the x-ray crystallographic structure of human population most occurring HLA alleles (HLA-DRB1 01:01, 15:01 and HLA-A 02:01) followed by filtration of previously bound ligands.
The following is a step-wise detail on how to construct a single or multiepitope-based vaccine and its property prediction;
- Molecular Docking Analysis: to determine the interaction pattern of the screened out epitopes with the HLA alleles by employing ClusPro v.2 (a protein-protein docking web server). This server performs this task by energy minimization, calculation of both the binding energy scores of the docked complex and electrostatic/shape complementarity.
- Target-Protein Comparative Modeling and associated Structure Validation: the sequence of the amino acid in the target protein (e.g., TLR-9) can be retrieved from UniProt and the tertiary structure with Raptor-X and I tasser (online comparative modeling tools). The server constructs and creates a 3D model (mathematical representation) of the target protein using hierarchical algorithms. 118 , 125
- Designing of Final Single/Multiepitope Vaccine Construct: in the case of multiepitope-based vaccine; the different epitopes can be linked to each other using AAY and GPGPG linkers and to the adjuvant using EAAAK linker.
- Predicting the antigenicity and allergenicity together with other physicochemical information: allergenicity of the final vaccine construct can be analyzed by Vaxijen v.2 tool, antigenicity of the vaccine by the use of AlgPred server and the physicochemical properties (isoelectric point, half-life, solubility, molecular weight, aliphatic index, and average hydropathicity) with the ProtParam server.
- Structure modeling, Refinement and Validation of the final Vaccine-these can be investigated using the SOPMA server/Raptor-x, Galaxy Refine Server and ProSA-web, respectively. This process is being followed by molecular docking and molecular dynamics of receptor and the epitope vaccine using ClusPro and Molecular Dynamic Simulation (Desmond) 126 respectively.
Case of Personalized Vaccination
Personalized vaccine refers to vaccines “targeted” toward an optimized outcome. Immunogenicity is maximized while either the risk of vaccine failure or reactogenicity and side effect is minimized. Personalized are developed in the following cases;
The Individual Level
Vaccines are developed to take care of haplotype and polymorphism knowing that they can retard the formation of a protective immune response or become pointers to the risk of an adverse vaccine reaction.
The Gender Level
This is needed when it is clear that females produce a higher antibody titre against a particular vaccine antigen than do their male counterparts.
The Racial/Ethnic Level
Where it is clear that a particular human race or ethnic group has a higher or lower immune response to a particular vaccine antigen.
The Subpopulation Level
Personalized vaccines arise due to known complex interactions between host environmental, genetic and some other factors that may be influencing the vaccine immune responses. The associations between the immune response gene polymorphisms and variations in immune responses to a particular gene must be pine-pointed when it is clear that a particular drug either suppresses or augments the transcription of an immune response gene. 127 , 128 This could help in designing vaccines or vaccine adjuvants that can circumvent restrictions due to immunological differences arising from varying genetic compositions. 129 , 130
Personalized vaccines stem from our understanding of how, within the human leukocyte antigen (HLA) system – also referred to as the major histocompatibility complex (MHC), the T cells are able to recognize peptides of pathogenic origin. 131 , 132 HLA molecules enjoy the double advantages of having stable polymorphisms and being fully characterized. 133 These advantages make good candidates for personalized vaccine design. HLA polymorphism, although stable, is complex. For instance, more than 12,000 alleles of HLA class I molecules and greater than 4000 class II molecules have been identified among human populations. 133 , 134 HLA class I and II molecules have heterodimeric character comprising of α and β chains, three highly variable extracellular domains (α1, α2, and α3) and then transmembrane and intracytoplasmic domains that are less variable. 133 , 135 HLA genes contain eight exons. Exon 1 is responsible for producing the leader peptide; exons 2,3,4 produce α1, α2, and α3 extracellular domains, respectively, for MHC class 1 or α1, β1and α3, respectively, for MHC class II; exon 5 encodes transmembrane anchor; exons 6 and 7 encodes the cytoplasmic tail while exon 8 encodes the 3ʹ-untranslated region. 135 Most of the several forms associated with the class I and II genes are seen in α-1 and α-2 (as known as class I) and in α-1 and β-1 (as known as class II) domains. 133 MHC I and II bind and present the peptide to T cells.
T cell responses to viral pathogens differ from one patient to the other, basically because of the expression of differing HLA (MHC) alleles which determine the several viral amino acid sequence brought to the T cells to read. 136 , 137 It is most likely that during an infection, diverse epitopes are usually presented to the T cells to read owing to the several forms of HLA alleles and also because each human person expresses six HLA Class I and six HLA Class II alleles. 138 Now, antibody-binding sites in a given HLA (MHC) molecule are mostly prediction-servers pre-determined on the basis of particular binding motifs and the anchor residues. 139 , 140 These residues refer to known amino acids located at defined locations in the peptide chain and which are peculiar to each MHC molecule. 141 , 142 Prediction-server database of peptide motifs and/or of MHC ligands may be obtained from web-based and/or from prediction-servers dedicated to NetMHC family. 143 , 144
In another example, sequence analysis of Lassa fever virus (the LASV) and other viruses’ immunoproteomic was used to identify the best immunogenic protein predicting T-cell as well as B-cell epitopes and also target sequence and binding sites. 145 , 146 The SSNLYKGVY peptide sequence at AA41-49 of glycoprotein 1 (produced by the L segment) was the best candidate epitope for the induction of humoral as well as the cell-mediated immunity for Lassa fever vaccine construct. 17 HLA-I and 16 HLA-II molecules have been proven in sizable African populations and their combination with the SSNLYKGVY peptide sequence may prove useful in such Lassa fever virus endemic areas. 145 This approach will strongly improve individualized vaccination and help combat emerging infections. The HLA region is suspected to contribute, to a large extent, to genetic propensity to infections and differences in vaccine expected immune responses. 132 , 147
Studies show that females exhibit stronger immune responses to immunization compared to males. 148 , 149 There are differential antibody responses to rubella and measles viral protein between males and females and that both hormonal and genetic difference may be influencing the immune responses. 148 , 150 , 151
Practical issues may stand in the way of achieving this new development (personalized vaccinology). Having to use different vaccines for different persons based because of personal genetic composition requires more time and labor during the vaccination process. Also, screening for individual factors for targeted vaccination can significantly increase vaccination cost. But with all these challenges, personalized vaccination is the new age approach in achieving an optimum immunization that takes into consideration the individual immune differences in a particular population and it is a new dawn for vaccine development.
Personalized vaccine development is strongly improved by vaccinomics. The field of vaccinomics looks at how immune response gene polymorphisms affect the cell-mediated, humoral and innate immune responses to vaccine antigens at population and specific individual levels. “Vaccinomics” encompasses both immunogenomics and immunogenetics as it concerns immune responses to vaccine antigen. 152 The fields of personalized vaccinology and vaccinomics were the products of Phase I of the international HapMap and that of the Human Genome Project. Also, modern molecular assay techniques permitting high-throughput detection of variations at gene level, in particular linkage disequilibrium maps and single nucleotide polymorphism (SNP), played significant roles in the development of personalized vaccinology and vaccinomics. It has also been shown that polymorphisms at vital immune response genes can bring about differing immune responses to biopharmaceutical products including vaccines. 152 – 154
Newer, accurate, cheap and reproducible sequencing technologies; validated databases containing genotype-phenotype data; statistical and bioinformatics tools are needed in order to analyze and interpret data that will help and improve vaccine adverse and immune response quantifiability and predictability. 155 The information will enhance clinical practice and accelerate rational and directed vaccine development.
The Pregnant Women
Safe vaccines are a critical requirement for any immunization program. 156 Conventional vaccination has been an approach targeted at all groups and individuals but has failed toward the enrolment of pregnant women into vaccination programs because of presumed fetal and maternal harms. 157 , 158 Evidence on the safety of vaccination in pregnancy is very small because pregnant women were usually excluded from participating in vaccine trials. 159 Pregnancy can alter the maternal as well as fetal immunological responses. 160 It is pertinent to explore research opportunities presented in advanced vaccine designs such as immunoinformatics (multiepitope vaccine docking) by studying human immune system functions and responses specific to pregnant women and their unborn children. 157
According to a report 161 from the Dominican Republic of Congo, during the 2016–2017 Zika virus outbreaks, over a thousand pregnant women were suspected of being infected with the virus and a sizable number were at their first trimester. The report further stated that fetal loss was approximately one-tenth of the pregnancies and that there were up to 3 cases of fetus with head circumferences smaller than normal. The widespread morbidity during the epidemic showed that Zika virus infection adversely affects pregnancy outcome. 160 , 161
Currently, there is no proof that pregnancy predisposes to Ebola virus infections in comparison with the non-pregnant population, but there is some evidence suggesting pregnancy to worsen the disease prognosis including fetal loss. Also, evidence showed that the virus can pass the placental barriers to establish infection in the unborn child. 162 Designing, implementing and enrolling pregnant women as well as perspective pregnant women into vaccine trials and programs is imperative in order to protect this group and ensure good vaccine uptake by them during infection outbreaks and epidemics. 157 , 163
These recommendations will give an informed decision to be investigated using the immunoinformatic tools to determine the immunogenic responses worthy of safe vaccine development for the pregnant women and perspective pregnant women group.
Maternal immunization offers palpable benefits in several ways. Some vaccines are primarily administered to shield these pregnant women from morbid conditions and/or death including fetal death. 164 , 165 Pregnant women stand the risk of being exposed to virulent pathogens and may be at a higher risk of morbidity and/or mortality when compared to the general population. 166
Conclusion and Future Research Directions
There has been an explosion of new immunological data ( Table 4 ) due to an increase in research to understand the immune system pathway in infectious disease pathogenesis and the application of the knowledge of bioinformatics has led to a better exposition of the immune system importance through immunoinformatics. The knowledge of immune system and the cost-effective, specific and effective approach like immunoinformatics, the concerns for emerging and re-surging diseases caused by pathogenic organisms, antigenic variability/complex lifecycle of pathogens and the need of personalized vaccination can be combated on a molecular level.
The future of immunological research is sharpened by the ability to make discoveries in biologics (e.g., vaccines) more effectively and efficiently. This will mean reduction and better targeting of wet laboratory experiments and will only be possible if wet laboratory experimentation is combined with bioinformatics techniques.
Limitation of Immunoinformatics Predictions in Vaccine Design
- Immunoinformatics depends on experimental science (wet laboratory) to produce raw data for analysis. The predictions are not formal proofs of any concepts. They do not replace the traditional experimental research methods of actually testing hypotheses.
- The quality of immunoinformatics predictions depends on the quality of data and the sophistication of the algorithms being used. Sequence data from high-throughput analysis often contain errors. If the sequences are wrong, or annotations incorrect, the results from the downstream analysis are misleading as well. 167
Acknowledgments
While this review has not been funded directly by them, we gratefully acknowledge the Drug Design Laboratory of Faculty of Pharmaceutical sciences, Nnamdi Azikiwe University, Nigeria, and Drug Discovery Africa.
Availability of Data
Author contributions.
All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
The authors report no funding and no conflicts of interest in this work.
Bibliographic and Citation Tools
Code, data and media associated with this article, recommenders and search tools.
- Institution
arXivLabs: experimental projects with community collaborators
arXivLabs is a framework that allows collaborators to develop and share new arXiv features directly on our website.
Both individuals and organizations that work with arXivLabs have embraced and accepted our values of openness, community, excellence, and user data privacy. arXiv is committed to these values and only works with partners that adhere to them.
Have an idea for a project that will add value for arXiv's community? Learn more about arXivLabs .
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- My Account Login
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 08 April 2024
Large-scale phenotyping of patients with long COVID post-hospitalization reveals mechanistic subtypes of disease
- Felicity Liew 1 na1 ,
- Claudia Efstathiou ORCID: orcid.org/0000-0001-6125-8126 1 na1 ,
- Sara Fontanella 1 ,
- Matthew Richardson 2 ,
- Ruth Saunders 2 ,
- Dawid Swieboda 1 ,
- Jasmin K. Sidhu 1 ,
- Stephanie Ascough 1 ,
- Shona C. Moore ORCID: orcid.org/0000-0001-8610-2806 3 ,
- Noura Mohamed 4 ,
- Jose Nunag ORCID: orcid.org/0000-0002-4218-0500 5 ,
- Clara King 5 ,
- Olivia C. Leavy 2 , 6 ,
- Omer Elneima 2 ,
- Hamish J. C. McAuley 2 ,
- Aarti Shikotra 7 ,
- Amisha Singapuri ORCID: orcid.org/0009-0002-4711-7516 2 ,
- Marco Sereno ORCID: orcid.org/0000-0003-4573-9303 2 ,
- Victoria C. Harris 2 ,
- Linzy Houchen-Wolloff ORCID: orcid.org/0000-0003-4940-8835 8 ,
- Neil J. Greening ORCID: orcid.org/0000-0003-0453-7529 2 ,
- Nazir I. Lone ORCID: orcid.org/0000-0003-2707-2779 9 ,
- Matthew Thorpe 10 ,
- A. A. Roger Thompson ORCID: orcid.org/0000-0002-0717-4551 11 ,
- Sarah L. Rowland-Jones 11 ,
- Annemarie B. Docherty ORCID: orcid.org/0000-0001-8277-420X 10 ,
- James D. Chalmers 12 ,
- Ling-Pei Ho ORCID: orcid.org/0000-0001-8319-301X 13 ,
- Alexander Horsley ORCID: orcid.org/0000-0003-1828-0058 14 ,
- Betty Raman 15 ,
- Krisnah Poinasamy 16 ,
- Michael Marks 17 , 18 , 19 ,
- Onn Min Kon 1 ,
- Luke S. Howard ORCID: orcid.org/0000-0003-2822-210X 1 ,
- Daniel G. Wootton 3 ,
- Jennifer K. Quint 1 ,
- Thushan I. de Silva ORCID: orcid.org/0000-0002-6498-9212 11 ,
- Antonia Ho 20 ,
- Christopher Chiu ORCID: orcid.org/0000-0003-0914-920X 1 ,
- Ewen M. Harrison ORCID: orcid.org/0000-0002-5018-3066 10 ,
- William Greenhalf 21 ,
- J. Kenneth Baillie ORCID: orcid.org/0000-0001-5258-793X 10 , 22 , 23 ,
- Malcolm G. Semple ORCID: orcid.org/0000-0001-9700-0418 3 , 24 ,
- Lance Turtle 3 , 24 ,
- Rachael A. Evans ORCID: orcid.org/0000-0002-1667-868X 2 ,
- Louise V. Wain 2 , 6 ,
- Christopher Brightling 2 ,
- Ryan S. Thwaites ORCID: orcid.org/0000-0003-3052-2793 1 na1 ,
- Peter J. M. Openshaw ORCID: orcid.org/0000-0002-7220-2555 1 na1 ,
- PHOSP-COVID collaborative group &
ISARIC investigators
Nature Immunology volume 25 , pages 607–621 ( 2024 ) Cite this article
5227 Accesses
1 Citations
625 Altmetric
Metrics details
- Inflammasome
- Inflammation
- Innate immunity
One in ten severe acute respiratory syndrome coronavirus 2 infections result in prolonged symptoms termed long coronavirus disease (COVID), yet disease phenotypes and mechanisms are poorly understood 1 . Here we profiled 368 plasma proteins in 657 participants ≥3 months following hospitalization. Of these, 426 had at least one long COVID symptom and 233 had fully recovered. Elevated markers of myeloid inflammation and complement activation were associated with long COVID. IL-1R2, MATN2 and COLEC12 were associated with cardiorespiratory symptoms, fatigue and anxiety/depression; MATN2, CSF3 and C1QA were elevated in gastrointestinal symptoms and C1QA was elevated in cognitive impairment. Additional markers of alterations in nerve tissue repair (SPON-1 and NFASC) were elevated in those with cognitive impairment and SCG3, suggestive of brain–gut axis disturbance, was elevated in gastrointestinal symptoms. Severe acute respiratory syndrome coronavirus 2-specific immunoglobulin G (IgG) was persistently elevated in some individuals with long COVID, but virus was not detected in sputum. Analysis of inflammatory markers in nasal fluids showed no association with symptoms. Our study aimed to understand inflammatory processes that underlie long COVID and was not designed for biomarker discovery. Our findings suggest that specific inflammatory pathways related to tissue damage are implicated in subtypes of long COVID, which might be targeted in future therapeutic trials.
Similar content being viewed by others
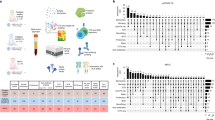
Immunopathological signatures in multisystem inflammatory syndrome in children and pediatric COVID-19
Keith Sacco, Riccardo Castagnoli, … Luigi D. Notarangelo
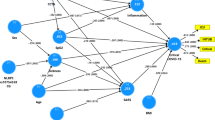
In COVID-19, NLRP3 inflammasome genetic variants are associated with critical disease and these effects are partly mediated by the sickness symptom complex: a nomothetic network approach
Michael Maes, Walton Luiz Del Tedesco Junior, … Andréa Name Colado Simão
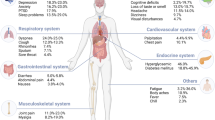
Epidemiology, clinical presentation, pathophysiology, and management of long COVID: an update
Sizhen Su, Yimiao Zhao, … Lin Lu
One in ten severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infections results in post-acute sequelae of coronavirus disease 2019 (PASC) or long coronavirus disease (COVID), which affects 65 million people worldwide 1 . Long COVID (LC) remains common, even after mild acute infection with recent variants 2 , and it is likely LC will continue to cause substantial long-term ill health, requiring targeted management based on an understanding of how disease phenotypes relate to underlying mechanisms. Persistent inflammation has been reported in adults with LC 1 , 3 , but studies have been limited in size, timing of samples or breadth of immune mediators measured, leading to inconsistent or absent associations with symptoms. Markers of oxidative stress, metabolic disturbance, vasculoproliferative processes and IFN-, NF-κB- or monocyte-related inflammation have been suggested 3 , 4 , 5 , 6 .
The PHOSP-COVID study, a multicenter United Kingdom study of patients previously hospitalized with COVID-19, has reported inflammatory profiles in 626 adults with health impairment after COVID-19, identified through clustering. Elevated IL-6 and markers of mucosal inflammation were observed in those with severe impairment compared with individuals with milder impairment 7 . However, LC is a heterogeneous condition that may be a distinct form of health impairment after COVID-19, and it remains unclear whether there are inflammatory changes specific to LC symptom subtypes. Determining whether activated inflammatory pathways underlie all cases of LC or if mechanisms differ according to clinical presentation is essential for developing effective therapies and has been highlighted as a top research priority by patients and clinicians 8 .
In this Letter, in a prospective multicenter study, we measured 368 plasma proteins in 657 adults previously hospitalized for COVID-19 (Fig. 1a and Table 1 ). Individuals in our cohort experienced a range of acute COVID-19 severities based on World Health Organization (WHO) progression scores 9 ; WHO 3–4 (no oxygen support, n = 133 and median age of 55 years), WHO 5–6 (oxygen support, n = 353 and median age of 59 years) and WHO 7–9 (critical care, n = 171 and median age of 57 years). Participants were hospitalized for COVID-19 ≥3 months before sample collection (median 6.1 months, interquartile range (IQR) 5.1–6.8 months and range 3.0–8.3 months) and confirmed clinically ( n = 36/657) or by PCR ( n = 621/657). Symptom data indicated 233/657 (35%) felt fully recovered at 6 months (hereafter ‘recovered’) and the remaining 424 (65%) reported symptoms consistent with the WHO definition for LC (symptoms ≥3 months post infection 10 ). Given the diversity of LC presentations, patients were grouped according to symptom type (Fig. 1b ). Groups were defined using symptoms and health deficits that have been commonly reported in the literature 1 ( Methods ). A multivariate penalized logistic regression model (PLR) was used to explore associations of clinical covariates and immune mediators at 6 months between recovered patients ( n = 233) and each LC group (cardiorespiratory symptoms, cardioresp, n = 398, Fig. 1c ; fatigue, n = 384, Fig. 1d ; affective symptoms, anxiety/depression, n = 202, Fig. 1e ; gastrointestinal symptoms, GI, n = 132, Fig. 1f ; and cognitive impairment, cognitive, n = 61, Fig. 1g ). Women ( n = 239) were more likely to experience CardioResp (odds ratio (OR 1.14), Fatigue (OR 1.22), GI (OR 1.13) and Cognitive (OR 1.03) outcomes (Fig. 1c,d,f,g ). Repeated cross-validation was used to optimize and assess model performance ( Methods and Extended Data Fig. 1 ). Pre-existing conditions, such as chronic lung disease, neurological disease and cardiovascular disease (Supplementary Table 1 ), were associated with all LC groups (Fig. 1c–g ). Age, C-reactive protein (CRP) and acute disease severity were not associated with any LC group (Table 1 ).
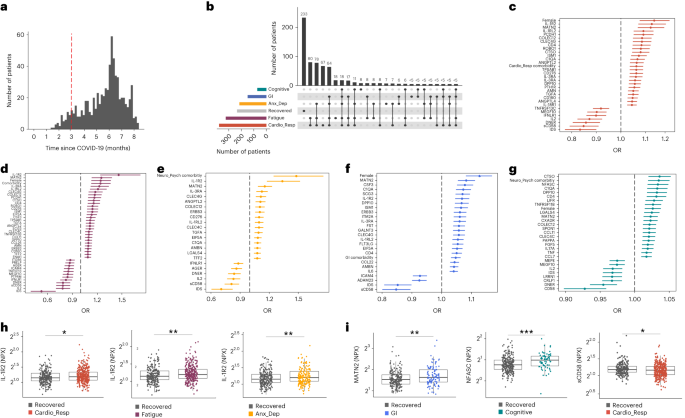
a , Distribution of time from COVID-19 hospitalization at sample collection. All samples were cross-sectional. The vertical red line indicates the 3 month cutoff used to define our final cohort and samples collected before 3 months were excluded. b , An UpSet plot describing pooled LC groups. The horizontal colored bars represent the number of patients in each symptom group: cardiorespiratory (Cardio_Resp), fatigue, cognitive, GI and anxiety/depression (Anx_Dep). Vertical black bars represent the number of patients in each symptom combination group. To prevent patient identification, where less than five patients belong to a combination group, this has been represented as ‘<5’. The recovered group ( n = 233) were used as controls. c – g , Forest plots of Olink protein concentrations (NPX) associated with Cardio_Resp ( n = 365) ( c ), fatigue (n = 314) ( d ), Anx_Dep ( n = 202) ( e ), GI ( n = 124) ( f ) and cognitive ( n = 60) ( g ). Neuro_Psych, neuropsychiatric. The error bars represent the median accuracy of the model. h , i , Distribution of Olink values (NPX) for IL-1R2 ( h ) and MATN2, neurofascin and sCD58 ( i ) measured between symptomatic and recovered individuals in recovered ( n = 233), Cardio_Resp ( n = 365), fatigue ( n = 314) and Anx_Dep ( n = 202) groups ( h ) and MATN2 in GI ( n = 124), neurofascin in cognitive ( n = 60) and sCD58 in Cardio_Resp and recovered groups ( i ). The box plot center line represents the median, the boundaries represent IQR and the whisker length represents 1.5× IQR. The median values were compared between groups using two-sided Wilcoxon signed-rank test, * P < 0.05, ** P < 0.01, *** P < 0.001 and **** P < 0.0001.
To study the association of peripheral inflammation with symptoms, we analyzed cross-sectional data collected approximately 6 months after hospitalizations. We measured 368 immune mediators from plasma collected contemporaneously with symptom data. Mediators suggestive of myeloid inflammation were associated with all symptoms (Fig. 1c–h ). Elevated IL-1R2, an IL-1 receptor expressed by monocytes and macrophages modulating inflammation 11 and MATN2, an extracellular matrix protein that modulates tissue inflammation through recruitment of innate immune cells 12 , were associated with cardioresp (IL-1R2 OR 1.14, Fig. 1c,h ), fatigue (IL-1R2 OR 1.45, Fig. 1d,h ), anxiety/depression (IL-1R2 OR 1.34. Fig. 1e,h ) and GI (MATN2 OR 1.08, Fig. 1f ). IL-3RA, an IL-3 receptor, was associated with cardioresp (OR 1.07, Fig. 1c ), fatigue (OR 1.21, Fig. 1d ), anxiety/depression (OR 1.12, Fig. 1e ) and GI (OR 1.06, Fig. 1f ) groups, while CSF3, a cytokine promoting neutrophilic inflammation 13 , was elevated in cardioresp (OR 1.06, Fig. 1c ), fatigue (OR 1.12, Fig. 1d ) and GI (OR 1.08, Fig. 1f ).
Elevated COLEC12, which initiates inflammation in tissues by activating the alternative complement pathway 14 , associated with cardioresp (OR 1.09, Fig. 1c ), fatigue (OR 1.19, Fig. 1d ) and anxiety/depression (OR 1.11, Fig. 1e ), but not with GI (Fig. 1f ) and only weakly with cognitive (OR 1.02, Fig. 1g ). C1QA, a degradation product released by complement activation 15 was associated with GI (OR 1.08, Fig. 1f ) and cognitive (OR 1.03, Fig. 1g ). C1QA, which is known to mediate dementia-related neuroinflammation 16 , had the third strongest association with cognitive (Fig. 1g ). These observations indicated that myeloid inflammation and complement activation were associated with LC.
Increased expression of DPP10 and SCG3 was observed in the GI group compared with recovered (DPP10 OR 1.07 and SCG3 OR 1.08, Fig. 1f ). DPP10 is a membrane protein that modulates tissue inflammation, and increased DPP10 expression is associated with inflammatory bowel disease 17 , 18 , suggesting that GI symptoms may result from enteric inflammation. Elevated SCG3, a multifunctional protein that has been associated with irritable bowel syndrome 19 , suggested that noninflammatory disturbance of the brain–gut axis or dysbiosis, may occur in the GI group. The cognitive group was associated with elevated CTSO (OR 1.04), NFASC (OR 1.03) and SPON-1 (OR 1.02, Fig. 1g,i ). NFASC and SPON-1 regulate neural growth 20 , 21 , while CTSO is a cysteine proteinase supporting tissue turnover 22 . The increased expression of these three proteins as well as C1QA and DPP10 in the cognitive group (Fig. 1g ) suggested neuroinflammation and alterations in nerve tissue repair, possibly resulting in neurodegeneration. Together, our findings indicated that complement activation and myeloid inflammation were common to all LC groups, but subtle differences were observed in the GI and cognitive groups, which may have mechanistic importance. Acutely elevated fibrinogen during hospitalization has been reported to be predictive of LC cognitive deficits 23 . We found elevated fibrinogen in LC relative to recovered (Extended Data Fig. 2a ; P = 0.0077), although this was not significant when restricted to the cognitive group ( P = 0.074), supporting our observation of complement pathway activation in LC and in keeping with reports that complement dysregulation and thrombosis drive severe COVID-19 (ref. 24 ).
Elevated sCD58 was associated with lower odds of all LC symptoms and was most pronounced in cardioresp (OR 0.85, Fig. 1c,i ), fatigue (OR 0.80, Fig. 1d ) and anxiety/depression (OR 0.83, Fig. 1e ). IL-2 was negatively associated with the cardioresp (Fig. 1c , OR 0.87), fatigue (Fig. 1d , OR 0.80), anxiety/depression (Fig. 1e , OR 0.84) and cognitive (Fig. 1g , OR 0.96) groups. Both IL-2 and sCD58 have immunoregulatory functions 25 , 26 . Specifically, sCD58 suppresses IL-1- or IL-6-dependent interactions between CD2 + monocytes and CD58 + T or natural killer cells 26 . The association of sCD58 with recovered suggests a central role of dysregulated myeloid inflammation in LC. Elevated markers of tissue repair, IDS and DNER 27 , 28 , were also associated with recovered relative to all LC groups (Fig. 1c–g ). Taken together, our data suggest that suppression of myeloid inflammation and enhanced tissue repair were associated with recovered, supporting the use of immunomodulatory agents in therapeutic trials 29 (Supplementary Table 2 ).
We next sought to validate the experimental and analytical approaches used. Although Olink has been validated against other immunoassay platforms, showing superior sensitivity and specificity 30 , 31 , we confirmed the performance of Olink against chemiluminescent immunoassays within our cohort. We performed chemiluminescent immunoassays on plasma from a subgroup of 58 participants (recovered n = 13 and LC n = 45). There were good correlations between results from Olink (normalized protein expression (NPX)) and chemiluminescent immunoassays (pg ml −1 ) for CSF3, IL-1R2, IL-3RA, TNF and TFF2 (Extended Data Fig. 3 ). Most samples did not have concentrations of IL-2 detectable using a mesoscale discovery chemiluminescent assay, limiting this analysis to 14 samples (recovered n = 4, LC n = 10, R = 0.55 and P = 0.053, Extended Data Fig. 3 ). We next repeated our analysis using alternative definitions of LC. The Centers for Disease Control and Prevention and National Institute for Health and Care Excellence definitions for LC include symptoms occurring 1 month post infection 32 , 33 . Using the 1 month post-infection definition included 62 additional participants to our analysis (recovered n = 21, 3 females and median age 61 years and LC n = 41, 15 females and median age 60 years, Extended Data Fig. 2c ) and found that inflammatory associations with each LC group were consistent with our analysis based on the WHO definition (Extended Data Fig. 2d–h ). Finally, to validate the analytical approach (PLR) we examined the distribution of data, prioritizing proteins that were most strongly associated with each LC/recovered group (IL-1R2, MATN2, NFASC and sCD58). Each protein was significantly elevated in the LC group compared with recovered (Fig. 1h,i and Extended Data Fig. 4 ), consistent with the PLR. Alternative regression approaches (unadjusted regression models and partial least squares, PLS) reported results consistent with the original analysis of protein associations and LC outcome in the WHO-defined cohort (Fig. 1c–g , Supplementary Table 3 and Extended Data Figs. 5 and 6 ). The standard errors of PLS estimates were wide (Extended Data Fig. 6 ), consistent with previous demonstrations that PLR is the optimal method to analyze high-dimensional data where variables may have combined effects 34 . As inflammatory proteins are often colinear, working in-tandem to mediate effects, we prioritized PLR results to draw conclusions.
To explore the relationship between inflammatory mediators associated with different LC symptoms, we performed a network analysis of Olink mediators highlighted by PLR within each LC group. COLEC12 and markers of endothelial and mucosal inflammation (MATN2, PCDH1, ROBO1, ISM1, ANGPTL2, TGF-α and TFF2) were highly correlated within the cardioresp, fatigue and anxiety/depression groups (Fig. 2 and Extended Data Fig. 7 ). Elevated PCDH1, an adhesion protein modulating airway inflammation 35 , was highly correlated with other inflammatory proteins associated with the cardioresp group (Fig. 2 ), suggesting that systemic inflammation may arise from the lung in these individuals. This was supported by increased expression of IL-3RA, which regulates innate immune responses in the lung through interactions with circulating IL-3 (ref. 36 ), in fatigue (Figs. 1d and 2 ), which correlated with markers of tissue inflammation, including PCDH1 (Fig. 2 ). MATN2 and ISM1, mucosal proteins that enhance inflammation 37 , 38 , were highly correlated in the GI group (Fig. 2 ), highlighting the role of tissue-specific inflammation in different LC groups. SCG3 correlated less closely with mediators in the GI group (Fig. 2 ), suggesting that the brain–gut axis may contribute separately to some GI symptoms. SPON-1, which regulates neural growth 21 , was the most highly correlated mediator in the cognitive group (Fig. 2 and Extended Data Fig. 7 ), highlighting that processes within nerve tissue may underlie this group. These observations suggested that inflammation might arise from mucosal tissues and that additional mechanisms may contribute to pathophysiology underlying the GI and cognitive groups.
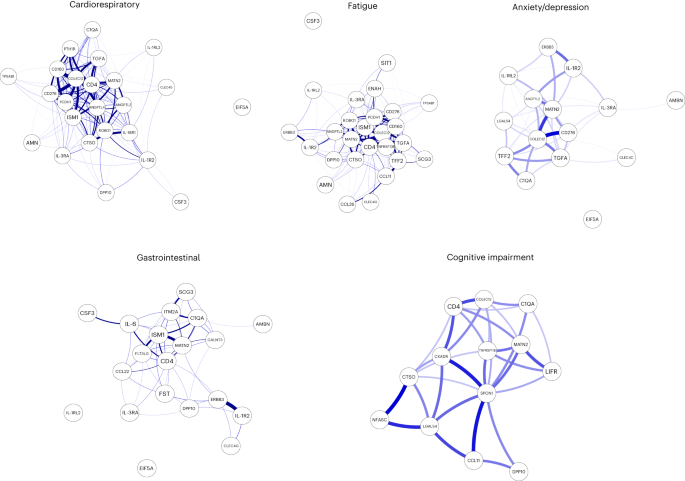
Network analysis of Olink mediators associated with cardioresp ( n = 365), fatigue ( n = 314), anxiety/depression ( n = 202), GI ( n = 124) and cognitive groups ( n = 60). Each node corresponds to a protein mediator identified by PLR. The edges (blue lines) were weighted according to the size of Spearman’s rank correlation coefficient between proteins. All edges represent positive and significant correlations ( P < 0.05) after FDR adjustment.
Women were more likely to experience LC (Table 1 ), as found in previous studies 1 . As estrogen can influence immunological responses 39 , we investigated whether hormonal differences between men and women with LC in our cohort explained this trend. We grouped men and women with LC symptoms into two age groups (those younger than 50 years and those 50 years and older, using age as a proxy for menopause status in women) and compared mediator levels between men and women in each age group, prioritizing those identified by PLR to be higher in LC compared with recovered. As we aimed to understand whether women with LC had stronger inflammatory responses than men with LC, we did not assess differences in men and women in the recovered group. IL-1R2 and MATN2 were significantly higher in women ≥50 years than men ≥50 years in the cardioresp group (Fig. 3a , IL-1R2 and MATN2) and the fatigue group (Fig. 3b ). In the GI group, CSF3 was higher in women ≥50 years compared with men ≥50 years (Fig. 3c ), indicating that the inflammatory markers observed in women were not likely to be estrogen-dependent. Women have been reported to have stronger innate immune responses to infection and to be at greater risk of autoimmunity 39 , possibly explaining why some women in the ≥50 years group had higher inflammatory proteins than men the same group. Proteins associated with the anxiety/depression (IL-1R2 P = 0.11 and MATN2 P = 0.61, Extended Data Fig. 8a ) and cognitive groups (CTSO P = 0.64 and NFASC P = 0.41, Extended Data Fig. 8b ) were not different between men and women in either age group, consistent with the absent/weak association between sex and these outcomes identified by PLR (Fig. 1e,g ). Though our findings suggested that nonhormonal differences in inflammatory responses may explain why some women are more likely to have LC, they require confirmation in adequately powered studies.
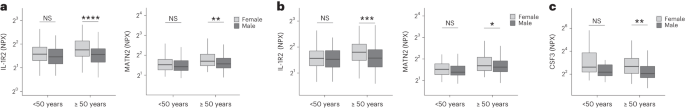
a – c , Olink-measured plasma protein levels (NPX) of IL-1R2 and MATN2 ( a and b ) and CSF3 ( c ) between LC men and LC women divided by age (<50 or ≥50 years) in the cardiorespiratory group (<50 years n = 8 and ≥50 years n = 270) ( a ), fatigue group (<50 years n = 81 and ≥50 years n = 227) ( b ) and GI group (<50 years n = 34 and ≥50 years n = 82) ( c ). the median values were compared between men and women using two-sided Wilcoxon signed-rank test, * P < 0.05, ** P < 0.01, *** P < 0.001 and **** P < 0.0001. The box plot center line represents the median, the boundaries represent IQR and the whisker length represents 1.5× IQR.
To test whether local respiratory tract inflammation persisted after COVID-19, we compared nasosorption samples from 89 participants (recovered, n = 31; LC, n = 33; and healthy SARS-CoV-2 naive controls, n = 25, Supplementary Tables 4 and 5 ). Several inflammatory markers were elevated in the upper respiratory tract post COVID (including IL-1α, CXCL10, CXCL11, TNF, VEGF and TFF2) when compared with naive controls, but similar between recovered and LC (Fig. 4a ). In the cardioresp group ( n = 29), inflammatory mediators elevated in plasma (for example, IL-6, APO-2, TGF-α and TFF2) were not elevated in the upper respiratory tract (Extended Data Fig. 9a ) and there was no correlation between plasma and nasal mediator levels (Extended Data Fig. 9b ). This exploratory analysis suggested upper respiratory tract inflammation post COVID was not specifically associated with cardiorespiratory symptoms.
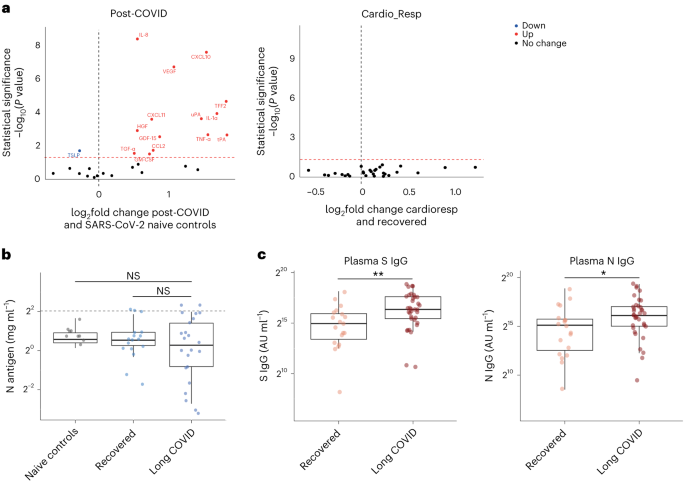
a , Nasal cytokines measured by immunoassay in post-COVID participants ( n = 64) compared with healthy SARS-CoV-2 naive controls ( n = 25), and between the the cardioresp group ( n = 29) and the recovered group ( n = 31). The red values indicate significantly increased cytokine levels after FDR adjustment ( P < 0.05) using two-tailed Wilcoxon signed-rank test. b , SARS-CoV-2 N antigen measured in sputum by electrochemiluminescence from recovered ( n = 17) and pooled LC ( n = 23) groups, compared with BALF from SARS-CoV-2 naive controls ( n = 9). The horizontal dashed line indicates the lower limit of detection of the assay. c , Plasma S- and N-specific IgG responses measured by electrochemiluminescence in the LC ( n = 35) and recovered ( n = 19) groups. The median values were compared using two-sided Wilcoxon signed-rank tests, NS P > 0.05, * P < 0.05, ** P < 0.01, *** P < 0.001 and **** P < 0.0001. The box plot center lines represent the median, the boundaries represent IQR and the whisker length represents 1.5× IQR.
To explore whether SARS-CoV-2 persistence might explain the inflammatory profiles observed in the cardioresp group, we measured SARS-CoV-2 nucleocapsid (N) antigen in sputum from 40 participants (recovered n = 17 and LC n = 23) collected approximately 6 months post hospitalization (Supplementary Table 6 ). All samples were compared with prepandemic bronchoalveolar lavage fluid ( n = 9, Supplementary Table 4 ). Only four samples (recovered n = 2 and LC n = 2) had N antigen above the assay’s lower limit of detection, and there was no difference in N antigen concentrations between LC and recovered (Fig. 4b , P = 0.78). These observations did not exclude viral persistence, which might require tissues samples for detection 40 , 41 . On the basis of the hypothesis that persistent viral antigen might prevent a decline in antibody levels over time, we examined the titers of SARS-CoV-2-specific antibodies in unvaccinated individuals (recovered n = 19 and LC n = 35). SARS-CoV-2 N-specific ( P = 0.023) and spike (S)-specific ( P = 0.0040) immunoglobulin G (IgG) levels were elevated in LC compared with recovered (Fig. 4c ).
Overall, we identified myeloid inflammation and complement activation in the cardioresp, fatigue, anxiety/depression, cognitive and GI groups 6 months after hospitalization (Extended Data Fig. 10 ). Our findings build on results of smaller studies 5 , 6 , 42 and are consistent with a genome-wide association study that identified an independent association between LC and FOXP4 , which modulates neutrophilic inflammation and immune cell function 43 , 44 . In addition, we identified tissue-specific inflammatory elements, indicating that myeloid disturbance in different tissues may result in distinct symptoms. Multiple mechanisms for LC have been suggested, including autoimmunity, thrombosis, vascular dysfunction, SARS-CoV-2 persistence and latent virus reactivation 1 . All these processes involve myeloid inflammation and complement activation 45 . Complement activation in LC has been suggested in a proteomic study in 97 mostly nonhospitalized COVID-19 cases 42 and a study of 48 LC patients, of which one-third experienced severe acute disease 46 . As components of the complement system are known to have a short half-life 47 , ongoing complement activation suggests active inflammation rather than past tissue damage from acute infection.
Despite the heterogeneity of LC and the likelihood of coexisting or multiple etiologies, our work suggests some common pathways that might be targeted therapeutically and supports the rationale for several drugs currently under trial. Our finding of increased sCD58 levels (associated with suppression of monocyte–lymphocyte interactions 26 ) in the recovered group, strengthens our conclusion that myeloid inflammation is central to the biology of LC and that trials of steroids, IL-1 antagonists, JAK inhibitors, naltrexone and colchicine are justified. Although anticoagulants such as apixaban might prevent thrombosis downstream of complement dysregulation, they can also increase the risk of serious bleeding when given after COVID-19 hospitalization 48 . Thus, clinical trials, already underway, need to carefully assess the risks and benefits of anticoagulants (Supplementary Table 2 ).
Our finding of elevated S- and N-specific IgG in LC could suggest viral persistence, as found in other studies 6 , 42 , 49 . Our network analysis indicated that inflammatory proteins in the cardioresp group interacted strongly with ISM1 and ROBO1, which are expressed during respiratory tract infection and regulate lung inflammation 50 , 51 . Although we were unable to find SARS-CoV-2 antigen in sputum from our LC cases, we did not test for viral persistence in GI tract and lung tissue 40 , 41 or in plasma 52 . Evidence of SARS-CoV-2 persistence would justify trials of antiviral drugs (singly or in combination) in LC. It is also possible that autoimmune processes could result in an innate inflammatory profile in LC. Autoreactive B cells have been identified in LC patients with higher SARS-CoV-2-specific antibody titers in a study of mostly mild acute COVID cases (59% WHO 2–3) 42 , a different population from our study of hospitalized cases.
Our observations of distinct protein profiles in GI and cognitive groups support previous reports on distinct associations between Epstein–Barr virus reactivation and neurological symptoms, or autoantibodies and GI symptoms relative to other forms of LC 49 , 53 . We did not assess autoantibody induction but found evidence of brain–gut axis disturbance (SCG3) in the GI group, which occurs in many autoimmune diseases 54 . We found signatures suggestive of neuroinflammation (C1QA) in the cognitive group, consistent with findings of brain abnormalities on magnetic resonance imaging after COVID-19 hospitalization 55 , as well as findings of microglial activation in mice after COVID-19 (ref. 56 ). Proinflammatory signatures dominated in the cardioresp, fatigue and anxiety/depression groups and were consistent with those seen in non-COVID depression, suggesting shared mechanisms 57 . The association between markers of myeloid inflammation, including IL-3RA, and symptoms was greatest for fatigue. Whilst membrane-bound IL-3RA facilitates IL-3 signaling upstream of myelopoesis 36 its soluble form (measured in plasma) can bind IL-3 and can act as a decoy receptor, preventing monocyte maturation and enhancing immunopathology 58 . Monocytes from individuals with post-COVID fatigue are reported to have abnormal expression profiles (including reduced CXCR2), suggestive of altered maturation and migration 5 , 59 . Lung-specific inflammation was suggested by the association between PCDH1 (an airway epithelial adhesion molecule 35 ) and cardioresp symptoms.
Our observations do not align with all published observations on LC. One proteomic study of 55 LC cases after generally mild (WHO 2–3) acute disease found that TNF and IFN signatures were elevated in LC 3 . Vasculoproliferative processes and metabolic disturbance have been reported in LC 4 , 60 , but these studies used uninfected healthy individuals for comparison and cannot distinguish between LC-specific phenomena and residual post-COVID inflammation. A study of 63 adults (LC, n = 50 and recovered, n = 13) reported no association between immune cell activation and LC 3 months after infection 61 , though myeloid inflammation was not directly measured, and 3 months post infection may be too early to detect subtle differences between LC and recovered cases due to residual acute inflammation.
Our study has limitations. We designed the study to identify inflammatory markers identifying pathways underlying LC subgroups rather than diagnostic biomarkers. The ORs we report are small, but associations were consistent across alternative methods of analysis and when using different LC definitions. Small effect sizes can be expected when using PLR, which shrinks correlated mediator coefficients to reflect combined effects and prevent colinear inflation 62 , and could also result from measurement of plasma mediators that may underestimate tissue inflammation. Although our LC cohort is large compared with most other published studies, some of our subgroups are small (only 60 cases were designated cognitive). Though the performance of the cognitive PLR model was adequate, our findings should be validated in larger studies. It should be noted that our cohort of hospitalized cases may not represent all types of LC, especially those occurring after mild infection. We looked for an effect of acute disease severity within our study and did not find it, and are reassured that the inflammatory profiles we observed were consistent with those seen in smaller studies including nonhospitalized cases 42 , 46 . Studies of posthospital LC may be confounded by ‘posthospital syndrome’, which encompasses general and nonspecific effects of hospitalization (particularly intensive care) 63 .
In conclusion, we found markers of myeloid inflammation and complement activation in our large prospective posthospital cohort of patients with LC, in addition to distinct inflammatory patterns in patients with cognitive impairment or gastrointestinal symptoms. These findings show the need to consider subphenotypes in managing patients with LC and support the use of antiviral or immunomodulatory agents in controlled therapeutic trials.
Study design and ethics
After hospitalization for COVID-19, adults who had no comorbidity resulting in a prognosis of less than 6 months were recruited to the PHOSP-COVID study ( n = 719). Patients hospitalized between February 2020 and January 2021 were recruited. Both sexes were recruited and gender was self-reported (female, n = 257 and male, n = 462). Written informed consent was obtained from all patients. Ethical approvals for the PHOSP-COVID study were given by Leeds West Research Ethics Committee (20/YH/0225).
Symptom data and samples were prospectively collected from individuals approximately 6 months (IQR 5.1–6.8 months and range 3.0–8.3 months) post hospitalization (Fig. 1a ), via the PHOSP-COVID multicenter United Kingdom study 64 . Data relating to patient demographics and acute admission were collected via the International Severe Acute Respiratory and Emerging Infection Consortium World Health Organization Clinical Characterisation Protocol United Kingdom (ISARIC4C study; IRAS260007/IRAS126600) (ref. 65 ). Adults hospitalized during the SARS-CoV-2 pandemic were systematically recruited into ISARIC4C. Written informed consent was obtained from all patients. Ethical approval was given by the South Central–Oxford C Research Ethics Committee in England (reference 13:/SC/0149), Scotland A Research Ethics Committee (20/SS/0028) and WHO Ethics Review Committee (RPC571 and RPC572l, 25 April 2013).
Data were collected to account for variables affecting symptom outcome, via hospital records and self-reporting. Acute disease severity was classified according to the WHO clinical progression score: WHO class 3–4: no oxygen therapy; class 5: oxygen therapy; class 6: noninvasive ventilation or high-flow nasal oxygen; and class 7–9: managed in critical care 9 . Clinical data were used to place patients into six categories: ‘recovered’, ‘GI’, ‘cardiorespiratory’, ‘fatigue’, ‘cognitive impairment’ and ‘anxiety/depression’ (Supplementary Table 7 ). Patient-reported symptoms and validated clinical scores were used when feasible, including Medical Research Council (MRC) breathlessness score, dyspnea-12 score, Functional Assessment of Chronic Illness Therapy (FACIT) score, Patient Health Questionnaire (PHQ)-9 and Generalized Anxiety Disorder (GAD)-7. Cognitive impairment was defined as a Montreal Cognitive Assessment score <26. GI symptoms were defined as answering ‘Yes’ to the presence of at least two of the listed symptoms. ‘Recovered’ was defined by self-reporting. Patients were placed in multiple groups if they experienced a combination of symptoms.
Matched nasal fluid and sputum samples were prospectively collected from a subgroup of convalescent patients approximately 6 months after hospitalization via the PHOSP-COVID study. Nasal and bronchoalveolar lavage fluid (BALF) collected from healthy volunteers before the COVID-19 pandemic were used as controls (Supplementary Table 4 ). Written consent was obtained for all individuals and ethical approvals were given by London–Harrow Research Ethics Committee (13/LO/1899) for the collection of nasal samples and the Health Research Authority London–Fulham Research Ethics Committee (IRAS project ID 154109; references 14/LO/1023, 10/H0711/94 and 11/LO/1826) for BALF samples.
Ethylenediaminetetraacetic acid plasma was collected from whole blood taken by venepuncture and frozen at −80 °C as previously described 7 , 66 . Nasal fluid was collected using a NasosorptionTM FX·I device (Hunt Developments), which uses a synthetic absorptive matrix to collect concentrated nasal fluid. Samples were eluted and stored as previously described 67 . Sputum samples were collected via passive expectoration and frozen at −80 °C without the addition of buffers. Sputum samples from convalescent individuals were compared with BALF from healthy SARS-CoV-2-naive controls, collected before the pandemic. BALF samples were used to act as a comparison for lower respiratory tract samples since passively expectorated sputum from healthy SARS-CoV-2-naive individuals was not available. BALF samples were obtained by instillation and recovery of up to 240 ml of normal saline via a fiberoptic bronchoscope. BALF was filtered through 100 µM strainers into sterile 50 ml Falcon tubes, then centrifuged for 10 min at 400 g at 4 °C. The resulting supernatant was transferred into sterile 50 ml Falcon tubes and frozen at −80 °C until use. The full methods for BALF collection and processing have been described previously 68 , 69 .
Immunoassays
To determine inflammatory signatures that associated with symptom outcomes, plasma samples were analyzed on an Olink Explore 384 Inflammation panel 70 . Supplementary Table 8 (Appendix 1 ) lists all the analytes measured. To ensure the validity of results, samples were run in a single batch with the use of negative controls, plate controls in triplicate and repeated measurement of patient samples between plates in duplicate. Samples were randomized between plates according to site and sample collection date. Randomization between plates was blind to LC/recovered outcome. Data were first normalized to an internal extension control that was included in each sample well. Plates were standardized by normalizing to interplate controls, run in triplicate on each plate. Each plate contained a minimum of four patient samples, which were duplicates on another plate; these duplicate pairs allowed any plate to be linked to any other through the duplicates. Data were then intensity normalized across all cohort samples. Finally, Olink results underwent quality control processing and samples or analytes that did not reach quality control standards were excluded. Final normalized relative protein quantities were reported as log 2 NPX values.
To further validate our findings, we performed conventional electrochemiluminescence (ECL) assays and enzyme-linked immunosorbent assay for Olink mediators that were associated with symptom outcome ( Supplementary Methods ). Contemporaneously collected plasma samples were available from 58 individuals. Like most omics platforms, Olink measures relative quantities, so perfect agreement with conventional assays that measure absolute concentrations is not expected.
Sputum samples were thawed before analysis and sputum plugs were extracted with the addition of 0.1% dithiothreitol creating a one in two sample dilution, as previously described 71 . SARS-CoV-2 S and N proteins were measured by ECL S-plex assay at a fixed dilution of one in two (Mesoscale Diagnostics), as per the manufacturers protocol 72 . Control BALF samples were thawed and measured on the same plate, neat. The S-plex assay is highly sensitive in detecting viral antigen in respiratory tract samples 73 .
Nasal cytokines were measured by ECL (mesoscale discovery) and Luminex bead multiplex assays (Biotechne). The full methods and list of analytes are detailed in Supplementary Methods .
Statistics and reproducibility
Clinical data was collected via the PHOSP REDCap database, to which access is available under reasonable request as per the data sharing statement in the manuscript. All analyses were performed within the Outbreak Data Analysis Platform (ODAP). All data and code can be accessed using information in the ‘Data sharing’ and ‘Code sharing’ statements at the end of the manuscript. No statistical method was used to predetermine sample size. Data distribution was assumed to be normal but this was not formally tested. Olink assays and immunoassays were randomized and investigators were blinded to outcomes.
To determine protein signatures that associated with each symptom outcome, a ridge PLR was used. PLR shrinks coefficients to account for combined effects within high-dimensional data, preventing false discovery while managing multicollinearity 34 . Thus, PLR was chosen a priori as the most appropriate model to assess associations between a large number of explanatory variables (that may work together to mediate effects) and symptom outcome 34 , 62 , 70 , 74 . In keeping with our aim to perform an unbiased exploration of inflammatory process, the model alpha was set to zero, facilitating regularization without complete penalization of any mediator. This enabled review of all possible mediators that might associate with LC 62 .
A 50 repeats tenfold nested cross-validation was used to select the optimal lambda for each model and assess its accuracy (Extended Data Fig. 1 ). The performance of the cognitive impairment model was influenced by the imbalance in size of the symptom group ( n = 60) relative to recovered ( n = 250). The model was weighted to account for this imbalance resulting in a sensitivity of 0.98, indicating its validity. We have expanded on the model performance and validation approaches in Supplementary Information .
Age, sex, acute disease severity and preexisting comorbidities were included as covariates in the PLR analysis (Supplementary Tables 1 and 3 ). Covariates were selected a priori using features reported to influence the risk of LC and inflammatory responses 1 , 39 , 64 , 75 . Ethnicity was not included since it has been shown not to predict symptom outcome in this cohort 64 . Individuals with missing data were excluded from the regression analysis. Each symptom group was compared with the ‘recovered’ group. The model coefficients of each covariate were converted into ORs for each outcome and visualized in a forest plot, after removing variables associated with regularized OR between 0.98 and 1.02 or in cases where most variables fell outside of this range, using mediators associated with the highest decile of coefficients either side of this range. This enabled exclusion of mediators with effect sizes that were unlikely to have clinical or mechanistic importance since the ridge PLR shrinks and orders coefficients according to their relative importance rather than making estimates with standard error. Thus, confidence intervals cannot be appropriately derived from PLR, and forest plot error bars were calculated using the median accuracy of the model generated by the nested cross-validation. To verify observations made through PLR analysis, we also performed an unadjusted PLR, an unadjusted logistic regression and a PLS analysis. Univariate analyses using Wilcoxon signed-rank test was also performed (Supplementary Table 8 , Appendix 1 ). Analyses were performed in R version 4.2.0 using ‘data.table v1.14.2’, ‘EnvStats v2.7.0’ ‘tidyverse v1.3.2’, ‘lme4 v1.1-32’, ‘caret v6.0-93’, ‘glmnet v4.1-6’, ‘mdatools v0.14.0’, ‘ggpubbr v0.4.0’ and ‘ggplot2 v3.3.6’ packages.
To further investigate the relationship between proteins elevated in each symptom group, we performed a correlation network analysis using Spearman’s rank correlation coefficient and false discovery rate (FDR) thresholding. The mediators visualized in the PLR forest plots, which were associated with cardiorespiratory symptoms, fatigue, anxiety/depression GI symptoms and cognitive impairment were used, respectively. Analyses were performed in R version 4.2.0 using ‘bootnet v1.5.6 ’ and ‘qgraph v1.9.8 ’ packages.
To determine whether differences in protein levels between men and women related to hormonal differences, we divided each symptom group into premenopausal and postmenopausal groups using an age cutoff of 50 years old. Differences between sexes in each group were determined using the Wilcoxon signed-rank test. To understand whether antigen persistence contributed to inflammation in adults with LC, the median viral antigen concentration from sputum/BALF samples and cytokine concentrations from nasal samples were compared using the Wilcoxon signed-rank test. All tests were two-tailed and statistical significance was defined as a P value < 0.05 after adjustment for FDR ( q -value of 0.05). Analyses were performed in R version 4.2.0 using ‘bootnet v1.5.6’ and ‘qgraph v1.9.8’ packages.
Extended Data Fig. 10 was made using Biorender, accessed at www.biorender.com .
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
This is an open access article under the CC BY 4.0 license.
The PHOSP-COVID protocol, consent form, definition and derivation of clinical characteristics and outcomes, training materials, regulatory documents, information about requests for data access, and other relevant study materials are available online at ref. 76 . Access to these materials can be granted by contacting [email protected] and [email protected].
The ISARIC4C protocol, data sharing and publication policy are available at https://isaric4c.net . ISARIC4C’s Independent Data and Material Access Committee welcomes applications for access to data and materials ( https://isaric4c.net ).
The datasets used in the study contain extensive clinical information at an individual level that prevent them from being deposited in an public depository due to data protection policies of the study. Study data can only be accessed via the ODAP, a protected research environment. All data used in this study are available within ODAP and accessible under reasonable request. Data access criteria and information about how to request access is available online at ref. 76 . If criteria are met and a request is made, access can be gained by signing the eDRIS user agreement.
Code availability
Code was written within the ODAP, using R v4.2.0 and publicly available packages (‘data.table v1.14.2’, ‘EnvStats v2.7.0’, ‘tidyverse v1.3.2’, ‘lme4 v1.1-32’, ‘caret v6.0-93’, ‘glmnet v4.1-6’, ‘mdatools v0.14.0’, ‘ggpubbr v0.4.0’, ‘ggplot2 v3.3.6’, ‘bootnet v1.5.6’ and ‘qgraph v1.9.8’ packages). No new algorithms or functions were created and code used in-built functions in listed packages available on CRAN. The code used to generate data and to analyze data is publicly available at https://github.com/isaric4c/wiki/wiki/ISARIC ; https://github.com/SurgicalInformatics/cocin_cc and https://github.com/ClaudiaEfstath/PHOSP_Olink_NatImm .
Davis, H. E., McCorkell, L., Vogel, J. M. & Topol, E. J. Long COVID: major findings, mechanisms and recommendations. Nat. Rev. Microbiol. 21 , 133–146 (2023).
Article CAS PubMed PubMed Central Google Scholar
Antonelli, M., Pujol, J. C., Spector, T. D., Ourselin, S. & Steves, C. J. Risk of long COVID associated with delta versus omicron variants of SARS-CoV-2. Lancet 399 , 2263–2264 (2022).
Talla, A. et al. Persistent serum protein signatures define an inflammatory subcategory of long COVID. Nat. Commun. 14 , 3417 (2023).
Captur, G. et al. Plasma proteomic signature predicts who will get persistent symptoms following SARS-CoV-2 infection. EBioMedicine 85 , 104293 (2022).
Scott, N. A. et al. Monocyte migration profiles define disease severity in acute COVID-19 and unique features of long COVID. Eur. Respir. J. https://doi.org/10.1183/13993003.02226-2022 (2023).
Klein, J. et al. Distinguishing features of Long COVID identified through immune profiling. Nature https://doi.org/10.1038/s41586-023-06651-y (2023).
Article PubMed PubMed Central Google Scholar
Evans, R. A. et al. Clinical characteristics with inflammation profiling of long COVID and association with 1-year recovery following hospitalisation in the UK: a prospective observational study. Lancet Respir. Med . 10 , 761–775 (2022).
Article CAS Google Scholar
Houchen-Wolloff, L. et al. Joint patient and clinician priority setting to identify 10 key research questions regarding the long-term sequelae of COVID-19. Thorax 77 , 717–720 (2022).
Article PubMed Google Scholar
Marshall, J. C. et al. A minimal common outcome measure set for COVID-19 clinical research. Lancet Infect. Dis. 20 , e192–e197 (2020).
Post COVID-19 condition (long COVID). World Health Organization https://www.who.int/europe/news-room/fact-sheets/item/post-covid-19-condition#:~:text=Definition,months%20with%20no%20other%20explanation (2022).
Peters, V. A., Joesting, J. J. & Freund, G. G. IL-1 receptor 2 (IL-1R2) and its role in immune regulation. Brain Behav. Immun. 32 , 1–8 (2013).
Article CAS PubMed Google Scholar
Luo, Z. et al. Monocytes augment inflammatory responses in human aortic valve interstitial cells via β2-integrin/ICAM-1-mediated signaling. Inflamm. Res. 71 , 681–694 (2022).
Bendall, L. J. & Bradstock, K. F. G-CSF: from granulopoietic stimulant to bone marrow stem cell mobilizing agent. Cytokine Growth Factor Rev. 25 , 355–367 (2014).
Ma, Y. J. et al. Soluble collectin-12 (CL-12) is a pattern recognition molecule initiating complement activation via the alternative pathway. J. Immunol. 195 , 3365–3373 (2015).
Laursen, N. S. et al. Functional and structural characterization of a potent C1q inhibitor targeting the classical pathway of the complement system. Front. Immunol. 11 , 1504 (2020).
Dejanovic, B. et al. Complement C1q-dependent excitatory and inhibitory synapse elimination by astrocytes and microglia in Alzheimer’s disease mouse models. Nat. Aging 2 , 837–850 (2022).
Xue, G., Hua, L., Zhou, N. & Li, J. Characteristics of immune cell infiltration and associated diagnostic biomarkers in ulcerative colitis: results from bioinformatics analysis. Bioengineered 12 , 252–265 (2021).
He, T. et al. Integrative computational approach identifies immune‐relevant biomarkers in ulcerative colitis. FEBS Open Bio. 12 , 500–515 (2022).
Sundin, J. et al. Fecal chromogranins and secretogranins are linked to the fecal and mucosal intestinal bacterial composition of IBS patients and healthy subjects. Sci. Rep. 8 , 16821 (2018).
Kriebel, M., Wuchter, J., Trinks, S. & Volkmer, H. Neurofascin: a switch between neuronal plasticity and stability. Int. J. Biochem. Cell Biol. 44 , 694–697 (2012).
Woo, W.-M. et al. The C. elegans F-spondin family protein SPON-1 maintains cell adhesion in neural and non-neural tissues. Development 135 , 2747–2756 (2008).
Yadati, T., Houben, T., Bitorina, A. & Shiri-Sverdlov, R. The ins and outs of cathepsins: physiological function and role in disease management. Cells 9 , 1679 (2020).
Taquet, M. et al. Acute blood biomarker profiles predict cognitive deficits 6 and 12 months after COVID-19 hospitalization. Nat. Med. https://doi.org/10.1038/s41591-023-02525-y (2023).
Siggins, M. K. et al. Alternative pathway dysregulation in tissues drives sustained complement activation and predicts outcome across the disease course in COVID‐19. Immunology 168 , 473–492 (2023).
Pol, J. G., Caudana, P., Paillet, J., Piaggio, E. & Kroemer, G. Effects of interleukin-2 in immunostimulation and immunosuppression. J. Exp. Med. 217 , e20191247 (2020).
Zhang, Y., Liu, Q., Yang, S. & Liao, Q. CD58 immunobiology at a glance. Front. Immunol. 12 , 705260 (2021).
Demydchuk, M. et al. Insights into Hunter syndrome from the structure of iduronate-2-sulfatase. Nat. Commun. 8 , 15786 (2017).
Wang, Z. et al. DNER promotes epithelial–mesenchymal transition and prevents chemosensitivity through the Wnt/β-catenin pathway in breast cancer. Cell Death Dis. 11 , 642 (2020).
Bonilla, H. et al. Therapeutic trials for long COVID-19: a call to action from the interventions taskforce of the RECOVER initiative. Front. Immunol. 14 , 1129459 (2023).
Wik, L. et al. Proximity extension assay in combination with next-generation sequencing for high-throughput proteome-wide analysis. Mol. Cell. Proteomics 20 , 100168 (2021).
Measuring protein biomarkers with Olink—technical comparisons and orthogonal validation. Olink Proteomics https://www.olink.com/content/uploads/2021/09/olink-technical-comparisons-and-orthogonal-validation-1118-v2.0.pdf (2021).
COVID-19 rapid guideline: managing the long-term effects of COVID-19. National Institute for Health and Care Excellence (NICE), Scottish Intercollegiate Guidelines Network (SIGN) and Royal College of General Practitioners (RCGP) https://www.nice.org.uk/guidance/ng188/resources/covid19-rapid-guideline-managing-the-longterm-effects-of-covid19-pdf-51035515742 (2022).
Long COVID or post-COVID conditions. Centers for Disease Control and Prevention https://www.cdc.gov/coronavirus/2019-ncov/long-term-effects/index.html#:~:text=Long%20COVID%20is%20broadly%20defined,after%20acute%20COVID%2D19%20infection (2023).
Firinguetti, L., Kibria, G. & Araya, R. Study of partial least squares and ridge regression methods. Commun. Stat. Simul. Comput 46 , 6631–6644 (2017).
Article Google Scholar
Mortensen, L. J., Kreiner-Moller, E., Hakonarson, H., Bonnelykke, K. & Bisgaard, H. The PCDH1 gene and asthma in early childhood. Eur. Respir. J. 43 , 792–800 (2014).
Tong, Y. et al. The RNFT2/IL-3Rα axis regulates IL-3 signaling and innate immunity. JCI Insight 5 , e133652 (2020).
Wu, Y. et al. Effect of ISM1 on the immune microenvironment and epithelial-mesenchymal transition in colorectal cancer. Front. Cell Dev. Biol. 9 , 681240 (2021).
Luo, G. G. & Ou, J. J. Oncogenic viruses and cancer. Virol. Sin. 30 , 83–84 (2015).
Klein, S. L. & Flanagan, K. L. Sex differences in immune responses. Nat. Rev. Immunol. 16 , 626–638 (2016).
Gaebler, C. et al. Evolution of antibody immunity to SARS-CoV-2. Nature 591 , 639–644 (2021).
Bussani, R. et al. Persistent SARS‐CoV‐2 infection in patients seemingly recovered from COVID‐19. J. Pathol. 259 , 254–263 (2023).
Woodruff, M. C. et al. Chronic inflammation, neutrophil activity, and autoreactivity splits long COVID. Nat. Commun. 14 , 4201 (2023).
Lammi, V. et al. Genome-wide association study of long COVID. Preprint at medRxiv https://doi.org/10.1101/2023.06.29.23292056 (2023).
Ismailova, A. et al. Identification of a forkhead box protein transcriptional network induced in human neutrophils in response to inflammatory stimuli. Front. Immunol. 14 , 1123344 (2023).
Beurskens, F. J., van Schaarenburg, R. A. & Trouw, L. A. C1q, antibodies and anti-C1q autoantibodies. Mol. Immunol. 68 , 6–13 (2015).
Cervia-Hasler, C. et al. Persistent complement dysregulation with signs of thromboinflammation in active long Covid. Science 383 , eadg7942 (2024).
Morgan, B. P. & Harris, C. L. Complement, a target for therapy in inflammatory and degenerative diseases. Nat. Rev. Drug Discov. 14 , 857–877 (2015).
Toshner, M. R. et al. Apixaban following discharge in hospitalised adults with COVID-19: preliminary results from a multicentre, open-label, randomised controlled platform clinical trial. Preprint at medRxiv , https://doi.org/10.1101/2022.12.07.22283175 (2022).
Su, Y. et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell 185 , 881–895.e20 (2022).
Branchfield, K. et al. Pulmonary neuroendocrine cells function as airway sensors to control lung immune response. Science 351 , 707–710 (2016).
Rivera-Torruco, G. et al. Isthmin 1 identifies a subset of lung hematopoietic stem cells and it is associated with systemic inflammation. J. Immunol. 202 , 118.18 (2019).
Swank, Z. et al. Persistent circulating severe acute respiratory syndrome coronavirus 2 spike is associated with post-acute coronavirus disease 2019 sequelae. Clin. Infect. Dis. 76 , e487–e490 (2023).
Peluso, M. J. et al. Chronic viral coinfections differentially affect the likelihood of developing long COVID. J. Clin. Invest. 133 , e163669 (2023).
Bellocchi, C. et al. The interplay between autonomic nervous system and inflammation across systemic autoimmune diseases. Int. J. Mol. Sci. 23 , 2449 (2022).
Raman, B. et al. Multiorgan MRI findings after hospitalisation with COVID-19 in the UK (C-MORE): a prospective, multicentre, observational cohort study. Lancet Respir. Med 11 , 1003–1019 (2023).
Fernández-Castañeda, A. et al. Mild respiratory COVID can cause multi-lineage neural cell and myelin dysregulation. Cell 185 , 2452–2468.e16 (2022).
Dantzer, R., O’Connor, J. C., Freund, G. G., Johnson, R. W. & Kelley, K. W. From inflammation to sickness and depression: when the immune system subjugates the brain. Nat. Rev. Neurosci. 9 , 46–56 (2008).
Broughton, S. E. et al. Dual mechanism of interleukin-3 receptor blockade by an anti-cancer antibody. Cell Rep. 8 , 410–419 (2014).
Ley, K., Miller, Y. I. & Hedrick, C. C. Monocyte and macrophage dynamics during atherogenesis. Arterioscler. Thromb. Vasc. Biol. 31 , 1506–1516 (2011).
Iosef, C. et al. Plasma proteome of long-COVID patients indicates HIF-mediated vasculo-proliferative disease with impact on brain and heart function. J. Transl. Med. 21 , 377 (2023).
Santopaolo, M. et al. Prolonged T-cell activation and long COVID symptoms independently associate with severe COVID-19 at 3 months. eLife 12 , e85009 (2023).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33 , 1–22 (2010).
Voiriot, G. et al. Chronic critical illness and post-intensive care syndrome: from pathophysiology to clinical challenges. Ann. Intensive Care 12 , 58 (2022).
Evans, R. A. et al. Physical, cognitive, and mental health impacts of COVID-19 after hospitalisation (PHOSP-COVID): a UK multicentre, prospective cohort study. Lancet Respir. Med 9 , 1275–1287 (2021).
Docherty, A. B. et al. Features of 20,133 UK patients in hospital with covid-19 using the ISARIC WHO clinical characterisation protocol: prospective observational cohort study. BMJ https://doi.org/10.1136/bmj.m1985 (2020).
Elneima, O. et al. Cohort profile: post-hospitalisation COVID-19 study (PHOSP-COVID). Preprint at medRxiv https://doi.org/10.1101/2023.05.08.23289442 (2023).
Liew, F. et al. SARS-CoV-2-specific nasal IgA wanes 9 months after hospitalisation with COVID-19 and is not induced by subsequent vaccination. EBioMedicine 87 , 104402 (2023).
Ascough, S. et al. Divergent age-related humoral correlates of protection against respiratory syncytial virus infection in older and young adults: a pilot, controlled, human infection challenge model. Lancet Healthy Longev. 3 , e405–e416 (2022).
Guvenel, A. et al. Epitope-specific airway-resident CD4+ T cell dynamics during experimental human RSV infection. J. Clin. Invest. 130 , 523–538 (2019).
Article PubMed Central Google Scholar
Greenwood, C. J. et al. A comparison of penalised regression methods for informing the selection of predictive markers. PLoS ONE 15 , e0242730 (2020).
Higham, A. et al. Leukotriene B4 levels in sputum from asthma patients. ERJ Open Res. 2 , 00088–02015 (2016).
SARS-CoV-2 spike kit. MSD https://www.mesoscale.com/~/media/files/product%20inserts/s-plex%20sars-cov-2%20spike%20kit%20product%20insert.pdf (2023).
Ren, A. et al. Ultrasensitive assay for saliva-based SARS-CoV-2 antigen detection. Clin. Chem. Lab. Med. 60 , 771–777 (2022).
Breheny, P. & Huang, J. Penalized methods for bi-level variable selection. Stat. Interface 2 , 369–380 (2009).
Thwaites, R. S. et al. Inflammatory profiles across the spectrum of disease reveal a distinct role for GM-CSF in severe COVID-19. Sci. Immunol. 6 , eabg9873 (2021).
Resources. PHOSP-COVID https://phosp.org/resource/ (2022).
Download references
Acknowledgements
This research used data assets made available by ODAP as part of the Data and Connectivity National Core Study, led by Health Data Research UK in partnership with the Office for National Statistics and funded by UK Research and Innovation (grant ref. MC_PC_20058). This work is supported by the following grants: the PHOSP-COVD study is jointly funded by UK Research and Innovation and National Institute of Health and Care Research (NIHR; grant references MR/V027859/1 and COV0319). ISARIC4C is supported by grants from the National Institute for Health and Care Research (award CO-CIN-01) and the MRC (grant MC_PC_19059) Liverpool Experimental Cancer Medicine Centre provided infrastructure support for this research (grant reference C18616/A25153). Other grants that have supported this work include the UK Coronavirus Immunology Consortium (funder reference 1257927), the Imperial Biomedical Research Centre (NIHR Imperial BRC, grant IS-BRC-1215-20013), the Health Protection Research Unit in Respiratory Infections at Imperial College London and NIHR Health Protection Research Unit in Emerging and Zoonotic Infections at University of Liverpool, both in partnership with Public Health England, (NIHR award 200907), Wellcome Trust and Department for International Development (215091/Z/18/Z), Health Data Research UK (grant code 2021.0155), MRC (grant code MC_UU_12014/12) and NIHR Clinical Research Network for providing infrastructure support for this research. We also acknowledge the support of the MRC EMINENT Network (MR/R502121/1), which is cofunded by GSK, the Comprehensive Local Research Networks, the MRC HIC-Vac network (MR/R005982/1) and the RSV Consortium in Europe Horizon 2020 Framework Grant 116019. F.L. is supported by an MRC clinical training fellowship (award MR/W000970/1). C.E. is funded by NIHR (grant P91258-4). L.-P.H. is supported by Oxford NIHR Biomedical Research Centre. A.A.R.T. is supported by a British Heart Foundation (BHF) Intermediate Clinical Fellowship (FS/18/13/33281). S.L.R.-J. receives support from UK Research and Innovation (UKRI), Global Challenges Research Fund (GCRF), Rosetrees Trust, British HIV association (BHIVA), European & Developing Countries Clinical Trials Partnership (EDCTP) and Globvac. J.D.C. has grants from AstraZeneca, Boehringer Ingelheim, GSK, Gilead Sciences, Grifols, Novartis and Insmed. R.A.E. holds a NIHR Clinician Scientist Fellowship (CS-2016-16-020). A. Horsley is currently supported by UK Research and Innovation, NIHR and NIHR Manchester BRC. B.R. receives support from BHF Oxford Centre of Research Excellence, NIHR Oxford BRC and MRC. D.G.W. is supported by an NIHR Advanced Fellowship. A. Ho has received support from MRC and for the Coronavirus Immunology Consortium (MR/V028448/1). L.T. is supported by the US Food and Drug Administration Medical Countermeasures Initiative contract 75F40120C00085 and the National Institute for Health Research Health Protection Research Unit in Emerging and Zoonotic Infections (NIHR200907) at the University of Liverpool in partnership with UK Health Security Agency (UK-HSA), in collaboration with Liverpool School of Tropical Medicine and the University of Oxford. L.V.W. has received support from UKRI, GSK/Asthma and Lung UK and NIHR for this study. M.G.S. has received support from NIHR UK, MRC UK and Health Protection Research Unit in Emerging and Zoonotic Infections, University of Liverpool. J.K.B. is supported by the Wellcome Trust (223164/Z/21/Z) and UKRI (MC_PC_20004, MC_PC_19025, MC_PC_1905, MRNO2995X/1 and MC_PC_20029). The funders were not involved in the study design, interpretation of data or writing of this manuscript. The views expressed are those of the authors and not necessarily those of the Department of Health and Social Care (DHSC), the Department for International Development (DID), NIHR, MRC, the Wellcome Trust, UK-HSA, the National Health Service or the Department of Health. P.J.M.O. is supported by a NIHR Senior Investigator Award (award 201385). We thank all the participants and their families. We thank the many research administrators, health-care and social-care professionals who contributed to setting up and delivering the PHOSP-COVID study at all of the 65 NHS trusts/health boards and 25 research institutions across the United Kingdom, as well as those who contributed to setting up and delivering the ISARIC4C study at 305 NHS trusts/health boards. We also thank all the supporting staff at the NIHR Clinical Research Network, Health Research Authority, Research Ethics Committee, Department of Health and Social Care, Public Health Scotland and Public Health England. We thank K. Holmes at the NIHR Office for Clinical Research Infrastructure for her support in coordinating the charities group. The PHOSP-COVID industry framework was formed to provide advice and support in commercial discussions, and we thank the Association of the British Pharmaceutical Industry as well the NIHR Office for Clinical Research Infrastructure for coordinating this. We are very grateful to all the charities that have provided insight to the study: Action Pulmonary Fibrosis, Alzheimer’s Research UK, Asthma and Lung UK, British Heart Foundation, Diabetes UK, Cystic Fibrosis Trust, Kidney Research UK, MQ Mental Health, Muscular Dystrophy UK, Stroke Association Blood Cancer UK, McPin Foundations and Versus Arthritis. We thank the NIHR Leicester Biomedical Research Centre patient and public involvement group and Long Covid Support. We also thank G. Khandaker and D. C. Newcomb who provided valuable feedback on this work. Extended Data Fig. 10 was created using Biorender.
Author information
These authors contributed equally: Felicity Liew, Claudia Efstathiou, Ryan S. Thwaites, Peter J. M. Openshaw.
Authors and Affiliations
National Heart and Lung Institute, Imperial College London, London, UK
Felicity Liew, Claudia Efstathiou, Sara Fontanella, Dawid Swieboda, Jasmin K. Sidhu, Stephanie Ascough, Onn Min Kon, Luke S. Howard, Jennifer K. Quint, Christopher Chiu, Ryan S. Thwaites, Peter J. M. Openshaw, Jake Dunning & Peter J. M. Openshaw
Institute for Lung Health, Leicester NIHR Biomedical Research Centre, University of Leicester, Leicester, UK
Matthew Richardson, Ruth Saunders, Olivia C. Leavy, Omer Elneima, Hamish J. C. McAuley, Amisha Singapuri, Marco Sereno, Victoria C. Harris, Neil J. Greening, Rachael A. Evans, Louise V. Wain, Christopher Brightling & Ananga Singapuri
NIHR Health Protection Research Unit in Emerging and Zoonotic Infections, Institute of Infection, Veterinary and Ecological Sciences, University of Liverpool, Liverpool, UK
Shona C. Moore, Daniel G. Wootton, Malcolm G. Semple, Lance Turtle, William A. Paxton & Georgios Pollakis
The Imperial Clinical Respiratory Research Unit, Imperial College NHS Trust, London, UK
Noura Mohamed
Cardiovascular Research Team, Imperial College Healthcare NHS Trust, London, UK
Jose Nunag & Clara King
Department of Population Health Sciences, University of Leicester, Leicester, UK
Olivia C. Leavy, Louise V. Wain & Beatriz Guillen-Guio
NIHR Leicester Biomedical Research Centre, University of Leicester, Leicester, UK
Aarti Shikotra
Centre for Exercise and Rehabilitation Science, NIHR Leicester Biomedical Research Centre-Respiratory, University of Leicester, Leicester, UK
Linzy Houchen-Wolloff
Usher Institute, University of Edinburgh, Edinburgh, UK
Nazir I. Lone, Luke Daines, Annemarie B. Docherty, Nazir I. Lone, Matthew Thorpe, Annemarie B. Docherty, Thomas M. Drake, Cameron J. Fairfield, Ewen M. Harrison, Stephen R. Knight, Kenneth A. Mclean, Derek Murphy, Lisa Norman, Riinu Pius & Catherine A. Shaw
Centre for Medical Informatics, The Usher Institute, University of Edinburgh, Edinburgh, UK
Matthew Thorpe, Annemarie B. Docherty, Ewen M. Harrison, J. Kenneth Baillie, Sarah L. Rowland-Jones, A. A. Roger Thompson & Thushan de Silva
Department of Infection, Immunity and Cardiovascular Disease, University of Sheffield, Sheffield, UK
A. A. Roger Thompson, Sarah L. Rowland-Jones, Thushan I. de Silva & James D. Chalmers
University of Dundee, Ninewells Hospital and Medical School, Dundee, UK
James D. Chalmers & Ling-Pei Ho
MRC Human Immunology Unit, University of Oxford, Oxford, UK
Ling-Pei Ho & Alexander Horsley
Division of Infection, Immunity and Respiratory Medicine, Faculty of Biology, Medicine and Health, University of Manchester, Manchester, UK
Alexander Horsley & Betty Raman
Radcliffe Department of Medicine, University of Oxford, Oxford, UK
Betty Raman & Krisnah Poinasamy
Asthma + Lung UK, London, UK
Krisnah Poinasamy & Michael Marks
Department of Clinical Research, London School of Hygiene and Tropical Medicine, London, UK
Michael Marks
Hospital for Tropical Diseases, University College London Hospital, London, UK
Division of Infection and Immunity, University College London, London, UK
Michael Marks & Mahdad Noursadeghi
MRC Centre for Virus Research, School of Infection and Immunity, University of Glasgow, Glasgow, UK
Antonia Ho & William Greenhalf
Institute of Systems, Molecular and Integrative Biology, University of Liverpool, Liverpool, UK
William Greenhalf & J. Kenneth Baillie
The Roslin Institute, University of Edinburgh, Edinburgh, UK
J. Kenneth Baillie, J. Kenneth Baillie, Sara Clohisey, Fiona Griffiths, Ross Hendry, Andrew Law & Wilna Oosthuyzen
Pandemic Science Hub, University of Edinburgh, Edinburgh, UK
J. Kenneth Baillie
The Pandemic Institute, University of Liverpool, Liverpool, UK
Malcolm G. Semple & Lance Turtle
University of Manchester, Manchester, UK
Kathryn Abel, Perdita Barran, H. Chinoy, Bill Deakin, M. Harvie, C. A. Miller, Stefan Stanel & Drupad Trivedi
Intensive Care Unit, Royal Infirmary of Edinburgh, Edinburgh, UK
Kathryn Abel & J. Kenneth Baillie
North Bristol NHS Trust and University of Bristol, Bristol, UK
H. Adamali, David Arnold, Shaney Barratt, A. Dipper, Sarah Dunn, Nick Maskell, Anna Morley, Leigh Morrison, Louise Stadon, Samuel Waterson & H. Welch
University of Edinburgh, Manchester, UK
Davies Adeloye, D. E. Newby, Riinu Pius, Igor Rudan, Manu Shankar-Hari, Catherine Sudlow, Sarah Walmsley & Bang Zheng
King’s College Hospital NHS Foundation Trust and King’s College London, London, UK
Oluwaseun Adeyemi, Rita Adrego, Hosanna Assefa-Kebede, Jonathon Breeze, S. Byrne, Pearl Dulawan, Amy Hoare, Caroline Jolley, Abigail Knighton, M. Malim, Sheetal Patale, Ida Peralta, Natassia Powell, Albert Ramos, K. Shevket, Fabio Speranza & Amelie Te
Guy’s and St Thomas’ NHS Foundation Trust, London, UK
Laura Aguilar Jimenez, Gill Arbane, Sarah Betts, Karen Bisnauthsing, A. Dewar, Nicholas Hart, G. Kaltsakas, Helen Kerslake, Murphy Magtoto, Philip Marino, L. M. Martinez, Marlies Ostermann, Jennifer Rossdale & Teresa Solano
Royal Free London NHS Foundation Trust, London, UK
Shanaz Ahmad, Simon Brill, John Hurst, Hannah Jarvis, C. Laing, Lai Lim, S. Mandal, Darwin Matila, Olaoluwa Olaosebikan & Claire Singh
University Hospital Birmingham NHS Foundation Trust and University of Birmingham, Birmingham, UK
N. Ahmad Haider, Catherine Atkin, Rhiannon Baggott, Michelle Bates, A. Botkai, Anna Casey, B. Cooper, Joanne Dasgin, Camilla Dawson, Katharine Draxlbauer, N. Gautam, J. Hazeldine, T. Hiwot, Sophie Holden, Karen Isaacs, T. Jackson, Vicky Kamwa, D. Lewis, Janet Lord, S. Madathil, C. McGee, K. Mcgee, Aoife Neal, Alex Newton-Cox, Joseph Nyaboko, Dhruv Parekh, Z. Peterkin, H. Qureshi, Liz Ratcliffe, Elizabeth Sapey, J. Short, Tracy Soulsby, J. Stockley, Zehra Suleiman, Tamika Thompson, Maximina Ventura, Sinead Walder, Carly Welch, Daisy Wilson, S. Yasmin & Kay Por Yip
Stroke Association, London, UK
Rubina Ahmed & Richard Francis
University College London Hospital and University College London, London, UK
Nyarko Ahwireng, Dongchun Bang, Donna Basire, Jeremy Brown, Rachel Chambers, A. Checkley, R. Evans, M. Heightman, T. Hillman, Joseph Jacob, Roman Jastrub, M. Lipman, S. Logan, D. Lomas, Marta Merida Morillas, Hannah Plant, Joanna Porter, K. Roy & E. Wall
Oxford University Hospitals NHS Foundation Trust and University of Oxford, Oxford, UK
Mark Ainsworth, Asma Alamoudi, Angela Bloss, Penny Carter, M. Cassar, Jin Chen, Florence Conneh, T. Dong, Ranuromanana Evans, V. Ferreira, Emily Fraser, John Geddes, F. Gleeson, Paul Harrison, May Havinden-Williams, P. Jezzard, Ivan Koychev, Prathiba Kurupati, H. McShane, Clare Megson, Stefan Neubauer, Debby Nicoll, C. Nikolaidou, G. Ogg, Edmund Pacpaco, M. Pavlides, Yanchun Peng, Nayia Petousi, John Pimm, Najib Rahman, M. J. Rowland, Kathryn Saunders, Michael Sharpe, Nick Talbot, E. M. Tunnicliffe & C. Xie
St George’s University Hospitals NHS Foundation Trust, London, UK
Mariam Ali, Raminder Aul, A. Dunleavy, D. Forton, Mark Mencias, N. Msimanga, T. Samakomva, Sulman Siddique, Vera Tavoukjian & J. Teixeira
University Hospitals of Leicester NHS Trust and University of Leicester, Leicester, UK
M. Aljaroof, Natalie Armstrong, H. Arnold, Hnin Aung, Majda Bakali, M. Bakau, E. Baldry, Molly Baldwin, Charlotte Bourne, Michelle Bourne, Nigel Brunskill, P. Cairns, Liesel Carr, Amanda Charalambou, C. Christie, Melanie Davies, Enya Daynes, Sarah Diver, Rachael Dowling, Sarah Edwards, C. Edwardson, H. Evans, J. Finch, Sarah Glover, Nicola Goodman, Bibek Gooptu, Kate Hadley, Pranab Haldar, Beverley Hargadon, W. Ibrahim, L. Ingram, Kamlesh Khunti, A. Lea, D. Lee, Gerry McCann, P. McCourt, Teresa Mcnally, George Mills, Will Monteiro, Manish Pareek, S. Parker, Anne Prickett, I. N. Qureshi, A. Rowland, Richard Russell, Salman Siddiqui, Sally Singh, J. Skeemer, M. Soares, E. Stringer, T. Thornton, Martin Tobin, T. J. C. Ward, F. Woodhead, Tom Yates & A. J. Yousuf
University of Exeter, Exeter, UK
Louise Allan, Clive Ballard & Andrew McGovern
University of Leicester, Leicester, UK
Richard Allen, Michelle Bingham, Terry Brugha, Selina Finney, Rob Free, Don Jones, Claire Lawson, Daniel Lozano-Rojas, Gardiner Lucy, Alistair Moss, Elizabeta Mukaetova-Ladinska, Petr Novotny, Kimon Ntotsis, Charlotte Overton, John Pearl, Tatiana Plekhanova, M. Richardson, Nilesh Samani, Jack Sargant, Ruth Saunders, M. Sharma, Mike Steiner, Chris Taylor, Sarah Terry, C. Tong, E. Turner, J. Wormleighton & Bang Zhao
Liverpool University Hospitals NHS Foundation Trust and University of Liverpool, Liverpool, UK
Lisa Allerton, Ann Marie Allt, M. Beadsworth, Anthony Berridge, Jo Brown, Shirley Cooper, Andy Cross, Sylviane Defres, S. L. Dobson, Joanne Earley, N. French, Kera Hainey, Hayley Hardwick, Jenny Hawkes, Victoria Highett, Sabina Kaprowska, Angela Key, Lara Lavelle-Langham, N. Lewis-Burke, Gladys Madzamba, Flora Malein, Sophie Marsh, Chloe Mears, Lucy Melling, Matthew Noonan, L. Poll, James Pratt, Emma Richardson, Anna Rowe, Victoria Shaw, K. A. Tripp, Lilian Wajero, S. A. Williams-Howard, Dan Wootton & J. Wyles
Sherwood Forest Hospitals NHS Foundation Trust, Nottingham, UK
Lynne Allsop, Kaytie Bennett, Phil Buckley, Margaret Flynn, Mandy Gill, Camelia Goodwin, M. Greatorex, Heidi Gregory, Cheryl Heeley, Leah Holloway, Megan Holmes, John Hutchinson, Jill Kirk, Wayne Lovegrove, Terri Ann Sewell, Sarah Shelton, D. Sissons, Katie Slack, Susan Smith, D. Sowter, Sarah Turner, V. Whitworth & Inez Wynter
Nottingham University Hospitals NHS Trust and University of Nottingham, London, UK
Paula Almeida, Akram Hosseini, Robert Needham & Karen Shaw
Manchester University NHS Foundation Trust and University of Manchester, London, UK
Bashar Al-Sheklly, Cristina Avram, John Blaikely, M. Buch, N. Choudhury, David Faluyi, T. Felton, T. Gorsuch, Neil Hanley, Tracy Hussell, Zunaira Kausar, Natasha Odell, Rebecca Osbourne, Karen Piper Hanley, K. Radhakrishnan & Sue Stockdale
Imperial College London, London, UK
Danny Altmann, Anew Frankel, Luke S. Howard, Desmond Johnston, Liz Lightstone, Anne Lingford-Hughes, William Man, Steve McAdoo, Jane Mitchell, Philip Molyneaux, Christos Nicolaou, D. P. O’Regan, L. Price, Jennifer K. Quint, David Smith, Jonathon Valabhji, Simon Walsh, Martin Wilkins & Michelle Willicombe
Hampshire Hospitals NHS Foundation Trust, Basingstoke, UK
Maria Alvarez Corral, Ava Maria Arias, Emily Bevan, Denise Griffin, Jane Martin, J. Owen, Sheila Payne, A. Prabhu, Annabel Reed, Will Storrar, Nick Williams & Caroline Wrey Brown
British Heart Foundation, Birmingham, UK
Shannon Amoils
NHS Greater Glasgow and Clyde Health Board and University of Glasgow, Glasgow, UK
David Anderson, Neil Basu, Hannah Bayes, Colin Berry, Ammani Brown, Andrew Dougherty, K. Fallon, L. Gilmour, D. Grieve, K. Mangion, I. B. McInnes, A. Morrow, Kathryn Scott & R. Sykes
University of Oxford, Oxford, UK
Charalambos Antoniades, A. Bates, M. Beggs, Kamaldeep Bhui, Katie Breeze, K. M. Channon, David Clark, X. Fu, Masud Husain, Lucy Kingham, Paul Klenerman, Hanan Lamlum, X. Li, E. Lukaschuk, Celeste McCracken, K. McGlynn, R. Menke, K. Motohashi, T. E. Nichols, Godwin Ogbole, S. Piechnik, I. Propescu, J. Propescu, A. A. Samat, Z. B. Sanders, Louise Sigfrid & M. Webster
Belfast Health and Social Care Trust and Queen’s University Belfast, Belfast, UK
Cherie Armour, Vanessa Brown, John Busby, Bronwen Connolly, Thelma Craig, Stephen Drain, Liam Heaney, Bernie King, Nick Magee, E. Major, Danny McAulay, Lorcan McGarvey, Jade McGinness, Tunde Peto & Roisin Stone
Airedale NHS Foundation Trust, Keighley, UK
Lisa Armstrong, Brigid Hairsine, Helen Henson, Claire Kurasz, Alison Shaw & Liz Shenton
Wrightington Wigan and Leigh NHS Trust, Wigan, UK
A. Ashish, Josh Cooper & Emma Robinson
Leeds Teaching Hospitals and University of Leeds, Leeds, UK
Andrew Ashworth, Paul Beirne, Jude Clarke, C. Coupland, Matthhew Dalton, Clair Favager, Jodie Glossop, John Greenwood, Lucy Hall, Tim Hardy, Amy Humphries, Jennifer Murira, Dan Peckham, S. Plein, Jade Rangeley, Gwen Saalmink, Ai Lyn Tan, Elaine Wade, Beverley Whittam, Nicola Window & Janet Woods
University of Liverpool, Liverpool, UK
M. Ashworth, D. Cuthbertson, G. Kemp, Anne McArdle, Benedict Michael, Will Reynolds, Lisa Spencer, Ben Vinson, Katie A. Ahmed, Jane A. Armstrong, Milton Ashworth, Innocent G. Asiimwe, Siddharth Bakshi, Samantha L. Barlow, Laura Booth, Benjamin Brennan, Katie Bullock, Nicola Carlucci, Emily Cass, Benjamin W. A. Catterall, Jordan J. Clark, Emily A. Clarke, Sarah Cole, Louise Cooper, Helen Cox, Christopher Davis, Oslem Dincarslan, Alejandra Doce Carracedo, Chris Dunn, Philip Dyer, Angela Elliott, Anthony Evans, Lorna Finch, Lewis W. S. Fisher, Lisa Flaherty, Terry Foster, Isabel Garcia-Dorival, Philip Gunning, Catherine Hartley, Karl Holden, Anthony Holmes, Rebecca L. Jensen, Christopher B. Jones, Trevor R. Jones, Shadia Khandaker, Katharine King, Robyn T. Kiy, Chrysa Koukorava, Annette Lake, Suzannah Lant, Diane Latawiec, Lara Lavelle-Langham, Daniella Lefteri, Lauren Lett, Lucia A. Livoti, Maria Mancini, Hannah Massey, Nicole Maziere, Sarah McDonald, Laurence McEvoy, John McLauchlan, Soeren Metelmann, Nahida S. Miah, Joanna Middleton, Joyce Mitchell, Ellen G. Murphy, Rebekah Penrice-Randal, Jack Pilgrim, Tessa Prince, P. Matthew Ridley, Debby Sales, Rebecca K. Shears, Benjamin Small, Krishanthi S. Subramaniam, Agnieska Szemiel, Aislynn Taggart, Jolanta Tanianis-Hughes, Jordan Thomas, Erwan Trochu, Libby van Tonder, Eve Wilcock & J. Eunice Zhang
University College London, London, UK
Shahab Aslani, Amita Banerjee, R. Batterham, Gabrielle Baxter, Robert Bell, Anthony David, Emma Denneny, Alun Hughes, W. Lilaonitkul, P. Mehta, Ashkan Pakzad, Bojidar Rangelov, B. Williams, James Willoughby & Moucheng Xu
Hull University Teaching Hospitals NHS Trust and University of Hull, Hull, UK
Paul Atkin, K. Brindle, Michael Crooks, Katie Drury, Nicholas Easom, Rachel Flockton, L. Holdsworth, A. Richards, D. L. Sykes, Susannah Thackray-Nocera & C. Wright
East Kent Hospitals University NHS Foundation Trust, Canterbury, UK
Liam Austin, Eva Beranova, Tracey Cosier, Joanne Deery, Tracy Hazelton, Carly Price, Hazel Ramos, Reanne Solly, Sharon Turney & Heather Weston
Baillie Gifford Pandemic Science Hub, Centre for Inflammation Research, The Queen’s Medical Research Institute, University of Edinburgh, Edinburgh, UK
Nikos Avramidis, J. Kenneth Baillie, Erola Pairo-Castineira & Konrad Rawlik
Roslin Institute, University of Edinburgh, Edinburgh, UK
Nikos Avramidis, J. Kenneth Baillie & Erola Pairo-Castineira
Newcastle upon Tyne Hospitals NHS Foundation Trust and University of Newcastle, Newcastle upon Tyne, UK
A. Ayoub, J. Brown, G. Burns, Gareth Davies, Anthony De Soyza, Carlos Echevarria, Helen Fisher, C. Francis, Alan Greenhalgh, Philip Hogarth, Joan Hughes, Kasim Jiwa, G. Jones, G. MacGowan, D. Price, Avan Sayer, John Simpson, H. Tedd, S. Thomas, Sophie West, M. Witham, S. Wright & A. Young
East Cheshire NHS Trust, Macclesfield, UK
Marta Babores, Maureen Holland, Natalie Keenan, Sharlene Shashaa & Helen Wassall
Sheffield Teaching NHS Foundation Trust and University of Sheffield, Sheffield, UK
J. Bagshaw, M. Begum, K. Birchall, Robyn Butcher, H. Carborn, Flora Chan, Kerry Chapman, Yutung Cheng, Luke Chetham, Cameron Clark, Zach Coburn, Joby Cole, Myles Dixon, Alexandra Fairman, J. Finnigan, H. Foot, David Foote, Amber Ford, Rebecca Gregory, Kate Harrington, L. Haslam, L. Hesselden, J. Hockridge, Ailsa Holbourn, B. Holroyd-Hind, L. Holt, Alice Howell, E. Hurditch, F. Ilyas, Claire Jarman, Allan Lawrie, Ju Hee Lee, Elvina Lee, Rebecca Lenagh, Alison Lye, Irene Macharia, M. Marshall, Angeline Mbuyisa, J. McNeill, Sharon Megson, J. Meiring, L. Milner, S. Misra, Helen Newell, Tom Newman, C. Norman, Lorenza Nwafor, Dibya Pattenadk, Megan Plowright, Julie Porter, Phillip Ravencroft, C. Roddis, J. Rodger, Peter Saunders, J. Sidebottom, Jacqui Smith, Laurie Smith, N. Steele, G. Stephens, R. Stimpson, B. Thamu, N. Tinker, Kim Turner, Helena Turton, Phillip Wade, S. Walker, James Watson, Imogen Wilson & Amira Zawia
University of Nottingham, Nottingham, UK
David Baguley, Chris Coleman, E. Cox, Laura Fabbri, Susan Francis, Ian Hall, E. Hufton, Simon Johnson, Fasih Khan, Paaig Kitterick, Richard Morriss, Nick Selby, Iain Stewart & Louise Wright
Wirral University Teaching Hospital, Wirral, UK
Elisabeth Bailey, Anne Reddington & Andrew Wight
MRC Human Genetics Unit, Institute of Genetics and Cancer, University of Edinburgh, Western General Hospital, Edinburgh, UK
University of Swansea, Swansea, UK
University of Southampton, London, UK
David Baldwin, P. C. Calder, Nathan Huneke & Gemma Simons
Royal Brompton and Harefield Clinical Group, Guy’s and St Thomas’ NHS Foundation Trust, London, UK
R. E. Barker, Daniele Cristiano, N. Dormand, P. George, Mahitha Gummadi, S. Kon, Kamal Liyanage, C. M. Nolan, B. Patel, Suhani Patel, Oliver Polgar, L. Price, P. Shah, Suver Singh & J. A. Walsh
York and Scarborough NHS Foundation Trust, York, UK
Laura Barman, Claire Brookes, K. Elliott, L. Griffiths, Zoe Guy, Kate Howard, Diana Ionita, Heidi Redfearn, Carol Sarginson & Alison Turnbull
NHS Highland, Inverness, UK
Fiona Barrett, A. Donaldson & Beth Sage
Royal Papworth Hospital NHS Foundation Trust, Cambridge, UK
Helen Baxendale, Lucie Garner, C. Johnson, J. Mackie, Alice Michael, J. Newman, Jamie Pack, K. Paques, H. Parfrey, J. Parmar & A. Reddy
University Hospitals of Derby and Burton, Derby, UK
Paul Beckett, Caroline Dickens & Uttam Nanda
NHS Lanarkshire, Hamilton, UK
Murdina Bell, Angela Brown, M. Brown, R. Hamil, Karen Leitch, L. Macliver, Manish Patel, Jackie Quigley, Andrew Smith & B. Welsh
Cambridge University Hospitals NHS Foundation Trust, NIHR Cambridge Clinical Research Facility and University of Cambridge, Cambridge, UK
Areti Bermperi, Isabel Cruz, K. Dempsey, Anne Elmer, Jonathon Fuld, H. Jones, Sherly Jose, Stefan Marciniak, M. Parkes, Carla Ribeiro, Jessica Taylor, Mark Toshner, L. Watson & J. Worsley
Loughborough University, Loughborough, UK
Lettie Bishop & David Stensel
Betsi Cadwallader University Health Board, Bangor, UK
Annette Bolger, Ffyon Davies, Ahmed Haggar, Joanne Lewis, Arwel Lloyd, R. Manley, Emma McIvor, Daniel Menzies, K. Roberts, W. Saxon, David Southern, Christian Subbe & Victoria Whitehead
Nottingham University Hospitals NHS Trust and University of Nottingham, Nottingham, UK
Charlotte Bolton, J. Bonnington, Melanie Chrystal, Catherine Dupont, Paul Greenhaff, Ayushman Gupta, W. Jang, S. Linford, Laura Matthews, Athanasios Nikolaidis, Sabrina Prosper & Andrew Thomas
King’s College London, London, UK
Kate Bramham, M. Brown, Khalida Ismail, Tim Nicholson, Carmen Pariante, Claire Sharpe, Simon Wessely & J. Whitney
Bradford Teaching Hospitals NHS Foundation Trust, Bradford, UK
Lucy Brear, Karen Regan, Dinesh Saralaya & Kim Storton
South London and Maudsley NHS Foundation Trust and King’s College London, London, UK
G. Breen & M. Hotopf
London School of Hygiene and Tropical Medicine, London, UK
Andrew Briggs
Whittington Health NHS Trust, London, UK
E. Bright, P. Crisp, Ruvini Dharmagunawardena & M. Stern
Cardiff and Vale University Health Board, Cardiff, UK
Lauren Broad, Teriann Evans, Matthew Haynes, L. Jones, Lucy Knibbs, Alison McQueen, Catherine Oliver, Kerry Paradowski, Ramsey Sabit & Jenny Williams
Yeovil District Hospital NHS Foundation Trust, Yeovil, UK
Andrew Broadley
University of Birmingham, Birmingham, UK
Mattew Broome, Paul McArdle, Paul Moss, David Thickett, Rachel Upthegrove, Dan Wilkinson, David Wraith & Erin L. Aldera
BHF Centre for Cardiovascular Science, University of Edinburgh, Edinburgh, UK
Anda Bularga
University of Cambridge, Cambridge, UK
Ed Bullmore, Jonathon Heeney, Claudia Langenberg, William Schwaeble, Charlotte Summers & J. Weir McCall
NIHR Leicester Biomedical Research Centre–Respiratory Patient and Public Involvement Group, Leicester, UK
Jenny Bunker, Rhyan Gill & Rashmita Nathu
Imperial College Healthcare NHS Trust and Imperial College London, London, UK
L. Burden, Ellen Calvelo, Bethany Card, Caitlin Carr, Edwin Chilvers, Donna Copeland, P. Cullinan, Patrick Daly, Lynsey Evison, Tamanah Fayzan, Hussain Gordon, Sulaimaan Haq, Gisli Jenkins, Clara King, Onn Min Kon, Katherine March, Myril Mariveles, Laura McLeavey, Silvia Moriera, Unber Munawar, Uchechi Nwanguma, Lorna Orriss-Dib, Alexandra Ross, Maura Roy, Emily Russell, Katherine Samuel, J. Schronce, Neil Simpson, Lawrence Tarusan, David Thomas, Chloe Wood & Najira Yasmin
Harrogate and District NHD Foundation Trust, Harrogate, UK
Tracy Burdett, James Featherstone, Cathy Lawson, Alison Layton, Clare Mills & Lorraine Stephenson
Newcastle University/Chair of NIHR Dementia TRC, Newcastle, UK
Oxford University Hospitals NHS Foundation Trust, Oxford, UK
A. Burns & N. Kanellakis
Tameside and Glossop Integrated Care NHS Foundation Trust, Ashton-under-Lyne, UK
Al-Tahoor Butt, Martina Coulding, Heather Jones, Susan Kilroy, Jacqueline McCormick, Jerome McIntosh, Heather Savill, Victoria Turner & Joanne Vere
University of Oxford, Nuffield Department of Medicine, Oxford, UK
University of Glasgow, Glasgow, UK
Jonathon Cavanagh, S. MacDonald, Kate O’Donnell, John Petrie, Naveed Sattar & Mark Spears
United Lincolnshire Hospitals NHS Trust, Grantham, UK
Manish Chablani & Lynn Osborne
Department of Psychological Medicine, Institute of Psychiatry, Psychology and Neuroscience, King’s College London, London, UK
Trudie Chalder
University Hospital of South Manchester NHS Foundation Trust, Manchester, UK
N. Chaudhuri
University Hospital Southampton NHS Foundation Trust and University of Southampton, Southampton, UK
Caroline Childs, R. Djukanovic, S. Fletcher, Matt Harvey, Mark Jones, Elizabeth Marouzet, B. Marshall, Reena Samuel, T. Sass, Tim Wallis & Helen Wheeler
King’s College Hospital/Guy’s and St Thomas’ NHS FT, London, UK
A. Chiribiri & C. O’Brien
Barts Health NHS Trust, London, UK
K. Chong-James, C. David, W. Y. James, Paul Pfeffer & O. Zongo
NHS Lothian and University of Edinburgh, Edinburgh, UK
Gaunab Choudhury, S. Clohisey, Andrew Deans, J. Furniss, Ewen Harrison, S. Kelly & Aziz Sheikh
School of Cardiovascular Medicine and Sciences. King’s College London, London, UK
Phillip Chowienczyk
Lewisham and Greenwich NHS Trust, London, UK
Hywel Dda University Health Board, Haverfordwest, UK
S. Coetzee, Kim Davies, Rachel Ann Hughes, Ronda Loosley, Heather McGuinness, Abdelrahman Mohamed, Linda O’Brien, Zohra Omar, Emma Perkins, Janet Phipps, Gavin Ross, Abigail Taylor, Helen Tench & Rebecca Wolf-Roberts
NHS Tayside and University of Dundee, Dundee, UK
David Connell, C. Deas, Anne Elliott, J. George, S. Mohammed, J. Rowland, A. R. Solstice, Debbie Sutherland & Caroline Tee
Swansea Bay University Health Board, Port Talbot, UK
Lynda Connor, Amanda Cook, Gwyneth Davies, Tabitha Rees, Favas Thaivalappil & Caradog Thomas
Faculty of Medicine, Nursing and Health Sciences, School of Biomedical Sciences, Monash University, Melbourne, Victoria, Australia
Eamon Coughlan
Rotherham NHS Foundation Trust, Rotherham, UK
Alison Daniels, Anil Hormis, Julie Ingham & Lisa Zeidan
Salford Royal NHS Foundation Trust, Salford, UK
P. Dark, Nawar Diar-Bakerly, D. Evans, E. Hardy, Alice Harvey, D. Holgate, Sean Knight, N. Mairs, N. Majeed, L. McMorrow, J. Oxton, Jessica Pendlebury, C. Summersgill, R. Ugwuoke & S. Whittaker
Cwm Taf Morgannwg University Health Board, Mountain Ash, UK
Ellie Davies, Cerys Evenden, Alyson Hancock, Kia Hancock, Ceri Lynch, Meryl Rees, Lisa Roche, Natalie Stroud & T. Thomas-Woods
Borders General Hospital, NHS Borders, Melrose, UK
Joy Dawson, Hosni El-Taweel & Leanne Robinson
Aneurin Bevan University Health Board, Caerleon, UK
Amanda Dell, Sara Fairbairn, Nancy Hawkings, Jill Haworth, Michaela Hoare, Victoria Lewis, Alice Lucey, Georgia Mallison, Heeah Nassa, Chris Pennington, Andrea Price, Claire Price, Andrew Storrie, Gemma Willis & Susan Young
University of Exeter Medical School, Exeter, UK
London North West University Healthcare NHS Trust, London, UK
Shalin Diwanji, Sambasivarao Gurram, Padmasayee Papineni, Sheena Quaid, Gerlynn Tiongson & Ekaterina Watson
Alzheimer’s Research UK, Cambridge, UK
Hannah Dobson
Health and Care Research Wales, Cardiff, UK
Yvette Ellis
University of Bristol, Bristol, UK
Jonathon Evans
University of Sheffield, Sheffield, UK
L. Finnigan, Laura Saunders & James Wild
Great Western Hospital Foundation Trust, Swindon, UK
Eva Fraile & Jacinta Ugoji
Royal Devon and Exeter NHS Trust, Barnstaple, UK
Michael Gibbons
Kettering General Hospital NHS Trust, Kettering, UK
Anne-Marie Guerdette, Melanie Hewitt, R. Reddy, Katie Warwick & Sonia White
NIHR Leicester Biomedical Research Centre, Leicester, UK
Beatriz Guillen-Guio
University of Leeds, Leeds, UK
Elspeth Guthrie & Max Henderson
Royal Surrey NHS Foundation Trust, Cranleigh, UK
Mark Halling-Brown & Katherine McCullough
Chesterfield Royal Hospital NHS Trust, Calow, UK
Edward Harris & Claire Sampson
Long Covid Support, London, UK
Claire Hastie, Natalie Rogers & Nikki Smith
King’s College Hospital, NHS Foundation Trust and King’s College London, London, UK
Department of Oncology and Metabolism, University of Sheffield, Sheffield, UK
Simon Heller
NIHR Office for Clinical Research Infrastructure, London, UK
Katie Holmes
Asthma UK and British Lung Foundation Partnership, London, UK
Ian Jarrold & Samantha Walker
North Middlesex University Hospital NHS Trust, London, UK
Bhagy Jayaraman & Tessa Light
Action for Pulmonary Fibrosis, Peterborough, UK
Cardiff University, National Centre for Mental Health, Cardiff, UK
McPin Foundation, London, UK
Thomas Kabir
Roslin Institute, The University of Edinburgh, Edinburgh, UK
Steven Kerr
The Hillingdon Hospitals NHS Foundation Trust, London, UK
Samantha Kon, G. Landers, Harpreet Lota, Mariam Nasseri & Sofiya Portukhay
Queen Mary University of London, London, UK
Ania Korszun
Swansea University, Swansea Welsh Network, Hywel Dda University Health Board, Swansea, UK
Royal Infirmary of Edinburgh, NHS Lothian, Edinburgh, UK
Nazir I. Lone
Barts Heart Centre, London, UK
Barts Health NHS Trust and Queen Mary University of London, London, UK
Adrian Martineau
Salisbury NHS Foundation Trust, Salisbury, UK
Wadzanai Matimba-Mupaya & Sophia Strong-Sheldrake
University of Newcastle, Newcastle, UK
Hamish McAllister-Williams, Stella-Maria Paddick, Anthony Rostron & John Paul Taylor
Gateshead NHS Trust, Gateshead, UK
W. McCormick, Lorraine Pearce, S. Pugmire, Wendy Stoker & Ann Wilson
Manchester Centre for Clinical Neurosciences, Salford Royal NHS Foundation Trust, Manchester, UK
Katherine McIvor
Kidney Research UK, Peterborough, UK
Aisling McMahon
NHS Dumfries and Galloway, Dumfries, UK
Michael McMahon & Paula Neill
Swansea University, Swansea, UK
MQ Mental Health Research, London, UK
Lea Milligan
BHF Centre for Cardiovascular Science, Usher Institute of Population Health Sciences and Informatics, University of Edinburgh, Edinburgh, UK
Nicholas Mills
Shropshire Community Health NHS Trust, Shropshire, UK
Sharon Painter, Johanne Tomlinson & Louise Warburton
Somerset NHS Foundation Trust, Taunton, UK
Sue Palmer, Dawn Redwood, Jo Tilley, Carinna Vickers & Tania Wainwright
Francis Crick Institute, London, UK
Markus Ralser
Manchester University NHD Foundation Trust, Manchester, UK
Pilar Rivera-Ortega
Diabetes UK, University of Glasgow, Glasgow, UK
Elizabeth Robertson
Barnsley Hospital NHS Foundation Trust, Barnsley, UK
Amy Sanderson
MRC–University of Glasgow Centre for Virus Research, Glasgow, UK
Janet Scott
Diabetes UK, London, UK
Kamini Shah
British Heart Foundation Centre, King’s College London, London, UK
King’s College Hospital NHS Foundation Trust, London, UK
University Hospitals Birmingham NHS Foundation Trust and University of Birmingham, Birmingham, UK
Institute of Cardiovascular and Medical Sciences, BHF Glasgow Cardiovascular Research Centre, University of Glasgow, Glasgow, UK
University College London NHS Foundation Trust, London and Barts Health NHS Trust, London, UK
Northumbria University, Newcastle upon Tyne, UK
Ioannis Vogiatzis
Swansea University and Swansea Welsh Network, Swansea, UK
N. Williams
DUK | NHS Digital, Salford Royal Foundation Trust, Salford, UK
Queen Alexandra Hospital, Portsmouth, UK
- Kayode Adeniji
Princess Royal Hospital, Haywards Heath, UK
Daniel Agranoff & Chi Eziefula
Bassetlaw Hospital, Bassetlaw, UK
Darent Valley Hospital, Dartford, UK
Queen Elizabeth the Queen Mother Hospital, Margate, UK
Ana Alegria
School of Informatics, University of Edinburgh, Edinburgh, UK
Beatrice Alex, Benjamin Bach & James Scott-Brown
North East and North Cumbria Ingerated, Newcastle upon Tyne, UK
Section of Biomolecular Medicine, Division of Systems Medicine, Department of Metabolism, Digestion and Reproduction, Imperial College London, London, UK
Petros Andrikopoulos, Kanta Chechi, Marc-Emmanuel Dumas, Julian Griffin, Sonia Liggi & Zoltan Takats
Section of Genomic and Environmental Medicine, Respiratory Division, National Heart and Lung Institute, Imperial College London, London, UK
Petros Andrikopoulos, Marc-Emmanuel Dumas, Michael Olanipekun & Anthonia Osagie
John Radcliffe Hospital, Oxford, UK
Brian Angus
Royal Albert Edward Infirmary, Wigan, UK
Abdul Ashish
Manchester Royal Infirmary, Manchester, UK
Dougal Atkinson
MRC Human Genetics Unit, Institute of Genetics and Cancer, University of Edinburgh, Western General Hospital, Crewe Road, Edinburgh, UK
Section of Molecular Virology, Imperial College London, London, UK
Wendy S. Barclay
Furness General Hospital, Barrow-in-Furness, UK
Shahedal Bari
Hull University Teaching Hospital Trust, Kingston upon Hull, UK
Gavin Barlow
Hillingdon Hospital, Hillingdon, UK
Stella Barnass
St Thomas’ Hospital, London, UK
Nicholas Barrett
Coventry and Warwickshire, Coventry, UK
Christopher Bassford
St Michael’s Hospital, Bristol, UK
Sneha Basude
Stepping Hill Hospital, Stockport, UK
David Baxter
Royal Liverpool University Hospital, Liverpool, UK
Michael Beadsworth
Bristol Royal Hospital Children’s, Bristol, UK
Jolanta Bernatoniene
Scarborough Hospital, Scarborough, UK
John Berridge
Golden Jubilee National Hospital, Clydebank, UK
Colin Berry
Liverpool Heart and Chest Hospital, Liverpool, UK
Nicola Best
Centre for Inflammation Research, The Queen’s Medical Research Institute, University of Edinburgh, Edinburgh, UK
Debby Bogaert & Clark D. Russell
James Paget University Hospital, Great Yarmouth, UK
Pieter Bothma & Darell Tupper-Carey
Aberdeen Royal Infirmary, Aberdeen, UK
Robin Brittain-Long
Adamson Hospital, Cupar, UK
Naomi Bulteel
Royal Devon and Exeter Hospital, Exeter, UK
Worcestershire Royal Hospital, Worcester, UK
Andrew Burtenshaw
ISARIC Global Support Centre, Centre for Tropical Medicine and Global Health, Nuffield Department of Medicine, University of Oxford, Oxford, UK
Gail Carson, Laura Merson & Louise Sigfrid
Conquest Hospital, Hastings, UK
Vikki Caruth
The James Cook University Hospital, Middlesbrough, UK
David Chadwick
Dorset County Hospital, Dorchester, UK
Duncan Chambler
Antimicrobial Resistance and Hospital Acquired Infection Department, Public Health England, London, UK
Meera Chand
Department of Epidemiology and Biostatistics, School of Public Health, Faculty of Medicine, Imperial College London, London, UK
Kanta Chechi
Royal Bournemouth General Hospital, Bournemouth, UK
Harrogate Hospital, Harrogate, UK
Jenny Child
Royal Blackburn Teaching Hospital, Blackburn, UK
Srikanth Chukkambotla
Edinburgh Clinical Research Facility, University of Edinburgh, Edinburgh, UK
Richard Clark, Audrey Coutts, Lorna Donelly, Angie Fawkes, Tammy Gilchrist, Katarzyna Hafezi, Louise MacGillivray, Alan Maclean, Sarah McCafferty, Kirstie Morrice, Lee Murphy & Nicola Wrobel
Torbay Hospital, Torquay, UK
Northern General Hospital, Sheffield, UK
Paul Collini, Cariad Evans & Gary Mills
Liverpool Clinical Trials Centre, University of Liverpool, Liverpool, UK
Marie Connor, Jo Dalton, Chloe Donohue, Carrol Gamble, Michelle Girvan, Sophie Halpin, Janet Harrison, Clare Jackson, Laura Marsh, Stephanie Roberts & Egle Saviciute
Department of Infectious Disease, Imperial College London, London, UK
Graham S. Cooke & Shiranee Sriskandan
St Georges Hospital (Tooting), London, UK
Catherine Cosgrove
Blackpool Victoria Hospital, Blackpool, UK
Jason Cupitt & Joanne Howard
The Royal London Hospital, London, UK
Maria-Teresa Cutino-Moguel
MRC-University of Glasgow Centre for Virus Research, Glasgow, UK
Ana da Silva Filipe, Antonia Y. W. Ho, Sarah E. McDonald, Massimo Palmarini, David L. Robertson, Janet T. Scott & Emma C. Thomson
Salford Royal Hospital, Salford, UK
University Hospital of North Durham, Durham, UK
Chris Dawson
Norfolk and Norwich University Hospital, Norwich, UK
Samir Dervisevic
Intensive Care Unit, Royal Infirmary Edinburgh, Edinburgh, UK
Annemarie B. Docherty & Seán Keating
Institute of Infection, Veterinary and Ecological Sciences, Faculty of Health and Life Sciences, University of Liverpool, Liverpool, UK
Cara Donegan & Rebecca G. Spencer
Salisbury District Hospital, Salisbury, UK
Phil Donnison
National Phenome Centre, Department of Metabolism, Digestion and Reproduction, Imperial College London, London, UK
Gonçalo dos Santos Correia, Matthew Lewis, Lynn Maslen, Caroline Sands, Zoltan Takats & Panteleimon Takis
Section of Bioanalytical Chemistry, Department of Metabolism, Digestion and Reproduction, Imperial College London, London, UK
Gonçalo dos Santos Correia, Matthew Lewis, Lynn Maslen, Caroline Sands & Panteleimon Takis
Guy’s and St Thomas’, NHS Foundation Trust, London, UK
Sam Douthwaite, Michael MacMahon, Marlies Ostermann & Manu Shankar-Hari
The Royal Oldham Hospital, Oldham, UK
Andrew Drummond
European Genomic Institute for Diabetes, Institut Pasteur de Lille, Lille University Hospital, University of Lille, Lille, France
Marc-Emmanuel Dumas
McGill University and Genome Quebec Innovation Centre, Montreal, Qeubec, Canada
National Infection Service, Public Health England, London, UK
Jake Dunning & Maria Zambon
Hereford Count Hospital, Hereford, UK
Ingrid DuRand
Southampton General Hospital, Southampton, UK
Ahilanadan Dushianthan
Northampton General Hospital, Northampton, UK
Tristan Dyer
University Hospital of Wales, Cardiff, UK
Chrisopher Fegan
University Hospitals Bristol NHS Foundation Trust, Bristol, UK
Liverpool School of Tropical Medicine, Liverpool, UK
Tom Fletcher
Leighton Hospital, Crewe, UK
Duncan Fullerton & Elijah Matovu
Manor Hospital, Walsall, UK
Scunthorpe Hospital, Scunthorpe, UK
Sanjeev Garg
Cambridge University Hospital, Cambridge, UK
Effrossyni Gkrania-Klotsas
West Suffolk NHS Foundation Trust, Bury St Edmunds, UK
Basingstoke and North Hampshire Hospital, Basingstoke, UK
Arthur Goldsmith
North Cumberland Infirmary, Carlisle, UK
Clive Graham
Paediatric Liver, GI and Nutrition Centre and MowatLabs, King’s College Hospital, London, UK
Tassos Grammatikopoulos
Institute of Liver Studies, King’s College London, London, UK
Institute of Microbiology and Infection, University of Birmingham, Birmingham, UK
Christopher A. Green
Department of Molecular and Clinical Cancer Medicine, University of Liverpool, Liverpool, UK
William Greenhalf
Institute for Global Health, University College London, London, UK
Rishi K. Gupta
NIHR Health Protection Research Unit, Institute of Infection, Veterinary and Ecological Sciences, Faculty of Health and Life Sciences, University of Liverpool, Liverpool, UK
Hayley Hardwick, Malcolm G. Semple, Tom Solomon & Lance C. W. Turtle
Warwick Hospital, Warwick, UK
Elaine Hardy
Birmingham Children’s Hospital, Birmingham, UK
Stuart Hartshorn
Nottingham City Hospital, Nottingham, UK
Daniel Harvey
Glangwili Hospital Child Health Section, Carmarthen, UK
Peter Havalda
Alder Hey Children’s Hospital, Liverpool, UK
Daniel B. Hawcutt
Department of Infectious Diseases, Queen Elizabeth University Hospital, Glasgow, UK
Antonia Y. W. Ho
Bronglais General Hospital, Aberystwyth, UK
Maria Hobrok
Worthing Hospital, Worthing, UK
Luke Hodgson
Centre for Tropical Medicine and Global Health, Nuffield Department of Medicine, University of Oxford, Oxford, UK
Peter W. Horby
Rotheram District General Hospital, Rotheram, UK
Anil Hormis
Virology Reference Department, National Infection Service, Public Health England, Colindale Avenue, London, UK
Samreen Ijaz
Royal Free Hospital, London, UK
Michael Jacobs & Padmasayee Papineni
Homerton Hospital, London, UK
Airedale Hospital, Airedale, UK
Paul Jennings
Basildon Hospital, Basildon, UK
Agilan Kaliappan
The Christie NHS Foundation Trust, Manchester, UK
Vidya Kasipandian
University Hospital Lewisham, London, UK
Stephen Kegg
The Whittington Hospital, London, UK
Michael Kelsey
Southmead Hospital, Bristol, UK
Jason Kendall
Sheffield Childrens Hospital, Sheffield, UK
Caroline Kerrison
Royal United Hospital, Bath, UK
Ian Kerslake
Department of Pharmacology, University of Liverpool, Liverpool, UK
Nuffield Department of Medicine, Peter Medawar Building for Pathogen Research, University of Oxford, Oxford, UK
Paul Klenerman
Translational Gastroenterology Unit, Nuffield Department of Medicine, University of Oxford, Oxford, UK
Public Health Scotland, Edinburgh, UK
Susan Knight, Eva Lahnsteiner & Sarah Tait
Western General Hospital, Edinburgh, UK
Oliver Koch
Southend University Hospital NHS Foundation Trust, Southend-on-Sea, UK
Gouri Koduri
Hinchingbrooke Hospital, Huntingdon, UK
George Koshy & Tamas Leiner
Royal Preston Hospital, Fulwood, UK
Shondipon Laha
University Hospital (Coventry), Coventry, UK
Steven Laird
The Walton Centre, Liverpool, UK
Susan Larkin
ISARIC, Global Support Centre, COVID-19 Clinical Research Resources, Epidemic diseases Research Group, Oxford (ERGO), University of Oxford, Oxford, UK
James Lee & Daniel Plotkin
Centre for Health Informatics, Division of Informatics, Imaging and Data Science, School of Health Sciences, Faculty of Biology, Medicine and Health, University of Manchester, Manchester Academic Health Science Centre, Manchester, UK
Gary Leeming
Hull Royal Infirmary, Hull, UK
Patrick Lillie
Nottingham University Hospitals NHS Trust:, Nottingham, UK
Wei Shen Lim
Darlington Memorial Hospital, Darlington, UK
Queen Elizabeth Hospital (Gateshead), Gateshead, UK
Vanessa Linnett
Warrington Hospital, Warrington, UK
Jeff Little
Bristol Royal Hospital for Children, Bristol, UK
Mark Lyttle
St Mary’s Hospital (Isle of Wight), Isle of Wight, UK
Emily MacNaughton
The Tunbridge Wells Hospital, Royal Tunbridge Wells, UK
Ravish Mankregod
Huddersfield Royal, Huddersfield, UK
Countess of Chester Hospital, Liverpool, UK
Ruth McEwen & Lawrence Wilson
Frimley Park Hospital, Frimley, UK
Manjula Meda
Nuffield Department of Medicine, John Radcliffe Hospital, Oxford, UK
Alexander J. Mentzer
Department of Microbiology/Infectious Diseases, Oxford University Hospitals NHS Foundation Trust, John Radcliffe Hospital, Oxford, UK
MRC Human Genetics Unit, MRC Institute of Genetics and Molecular Medicine, University of Edinburgh, Edinburgh, UK
Alison M. Meynert & Murray Wham
St James University Hospital, Leeds, UK
Jane Minton
Arrowe Park Hospital, Birkenhead, UK
Kavya Mohandas
Great Ormond Street Hospital, London, UK
Royal Shrewsbury Hospital, Shrewsbury, UK
Addenbrookes Hospital, Cambridge, UK
Elinoor Moore
Institute of Infection, Veterinary and Ecological Sciences, University of Liverpool, Liverpool, UK
Shona C. Moore, William A. Paxton & Georgios Pollakis
East Surrey Hospital (Redhill), Redhill, UK
Patrick Morgan
Burton Hospital, Burton, UK
Craig Morris & Tim Reynolds
Peterborough City Hospital, Peterborough, UK
Katherine Mortimore
Kent and Canterbury Hospital, Canterbury, UK
Samuel Moses
Weston Area General Trust, Bristol, UK
Mbiye Mpenge
Bedfordshire Hospital, Bedfordshire, UK
Rohinton Mulla
Glasgow Royal Infirmary, Glasgow, UK
Michael Murphy
Macclesfield General Hospital, Macclesfield, UK
Thapas Nagarajan
Derbyshire Healthcare, Derbyshire, UK
Megan Nagel
Chelsea and Westminster Hospital, London, UK
Mark Nelson & Matthew K. O’Shea
Watford General Hospital, Watford, UK
Lillian Norris & Tom Stambach
EPCC, University of Edinburgh, Edinburgh, UK
Lucy Norris
Section of Biomolecular Medicine, Division of Systems Medicine, Department of Metabolism, Digestion and Reproduction, London, UK
Michael Olanipekun
Imperial College Healthcare NHS Trust: London, London, UK
Peter J. M. Openshaw
Division of Systems Medicine, Department of Metabolism, Digestion and Reproduction, Imperial College London, London, UK
Anthonia Osagie
Prince Philip Hospital, Llanelli, UK
Igor Otahal & Andrew Workman
George Eliot Hospital – Acute Services, Nuneaton, UK
Molecular and Clinical Cancer Medicine, Institute of Systems, Molecular and Integrative Biology, University of Liverpool, Liverpool, UK
Carlo Palmieri
Clatterbridge Cancer Centre NHS Foundation Trust, Liverpool, UK
Kettering General Hospital, Kettering, UK
Selva Panchatsharam
University Hospitals of North Midlands NHS Trust, North Midlands, UK
Danai Papakonstantinou
Russells Hall Hospital, Dudley, UK
Hassan Paraiso
Harefield Hospital, Harefield, UK
Lister Hospital, Lister, UK
Natalie Pattison
Musgrove Park Hospital, Taunton, UK
Justin Pepperell
Kingston Hospital, Kingston, UK
Mark Peters
Queen’s Hospital, Romford, UK
Mandeep Phull
Southport and Formby District General Hospital, Southport, UK
Stefania Pintus
St George’s University of London, London, UK
Tim Planche
King’s College Hospital (Denmark Hill), London, UK
Centre for Clinical Infection and Diagnostics Research, Department of Infectious Diseases, School of Immunology and Microbial Sciences, King’s College London, London, UK
Nicholas Price
Department of Infectious Diseases, Guy’s and St Thomas’ NHS Foundation Trust, London, UK
The Clatterbridge Cancer Centre NHS Foundation, Bebington, UK
David Price
The Great Western Hospital, Swindon, UK
Rachel Prout
Ninewells Hospital, Dundee, UK
Nikolas Rae
Institute of Evolutionary Biology, University of Edinburgh, Edinburgh, UK
Andrew Rambaut
Poole Hospital NHS Trust, Poole, UK
Henrik Reschreiter
William Harvey Hospital, Ashford, UK
Neil Richardson
King’s Mill Hospital, Sutton-in-Ashfield, UK
Mark Roberts
Liverpool Women’s Hospital, Liverpool, UK
Devender Roberts
Pinderfields Hospital, Wakefield, UK
Alistair Rose
North Devon District Hospital, Barnstaple, UK
Guy Rousseau
Queen Elizabeth Hospital, Birmingham, UK
Tameside General Hospital, Ashton-under-Lyne, UK
Brendan Ryan
City Hospital (Birmingham), Birmingham, UK
Taranprit Saluja
Department of Pediatrics and Virology, St Mary’s Medical School Bldg, Imperial College London, London, UK
Vanessa Sancho-Shimizu
The Newcastle Upon Tyne Hospitals NHS Foundation Trust, Newcastle Upon Tyne, UK
Matthias Schmid
NHS Greater Glasgow and Clyde, Glasgow, UK
Janet T. Scott
Respiratory Medicine, Institute in The Park, University of Liverpool, Alder Hey Children’s Hospital, Liverpool, UK
Malcolm G. Semple
Broomfield Hospital, Broomfield, UK
Stoke Mandeville, UK
Prad Shanmuga
University Hospital of North Tees, Stockton-on-Tees, UK
Anil Sharma
Institute of Translational Medicine, University of, Liverpool, Merseyside, UK
Victoria E. Shaw
Royal Manchester Children’s Hospital, Manchester, UK
Anna Shawcross
New Cross Hospital, Wolverhampton, UK
Jagtur Singh Pooni
Bedford Hospital, Bedford, UK
Jeremy Sizer
Colchester General Hospital, Colchester, UK
Richard Smith
University Hospital Birmingham NHS Foundation Trust, Birmingham, UK
Catherine Snelson & Tony Whitehouse
Walton Centre NHS Foundation Trust, Liverpool, UK
Tom Solomon
Chesterfield Royal Hospital, Calow, UK
Nick Spittle
MRC Centre for Molecular Bacteriology and Infection, Imperial College London, London, UK
Shiranee Sriskandan
Princess Alexandra Hospital, Harlow, UK
Nikki Staines & Shico Visuvanathan
Milton Keynes Hospital, Eaglestone, UK
Richard Stewart
Division of Structural Biology, The Wellcome Centre for Human Genetics, University of Oxford, Oxford, UK
David Stuart
Royal Bolton Hopital, Farnworth, UK
Pradeep Subudhi
Department of Medicine, University of Cambridge, Cambridge, UK
Charlotte Summers
Department of Child Life and Health, University of Edinburgh, Edinburgh, UK
Olivia V. Swann
Royal Gwent (Newport), Newport, UK
Tamas Szakmany
The Royal Marsden Hospital (London), London, UK
Kate Tatham
Blood Borne Virus Unit, Virus Reference Department, National Infection Service, Public Health England, London, UK
Richard S. Tedder
Transfusion Microbiology, National Health Service Blood and Transplant, London, UK
Department of Medicine, Imperial College London, London, UK
Queen Victoria Hospital (East Grinstead), East Grinstead, UK
Leeds Teaching Hospitals NHS Trust, Leeds, UK
Robert Thompson
Royal Stoke University Hospital, Stoke-on-Trent, UK
Chris Thompson
Whiston Hospital, Rainhill, UK
Ascanio Tridente
Tropical and Infectious Disease Unit, Royal Liverpool University Hospital, Liverpool, UK
Lance C. W. Turtle
Croydon University Hospital, Thornton Heath, UK
Mary Twagira
Gloucester Royal, Gloucester, UK
Nick Vallotton
West Hertfordshire Teaching Hospitals NHS Trust, Hertfordshire, UK
Rama Vancheeswaran
North Middlesex Hospital, London, UK
Rachel Vincent
Medway Maritime Hospital, Gillingham, UK
Lisa Vincent-Smith
Royal Papworth Hospital Everard, Cambridge, UK
Alan Vuylsteke
Derriford (Plymouth), Plymouth, UK
St Helier Hospital, Sutton, UK
Rachel Wake
Royal Berkshire Hospital, Reading, UK
Andrew Walden
Royal Liverpool Hospital, Liverpool, UK
Ingeborg Welters
Bradford Royal infirmary, Bradford, UK
Paul Whittaker
Central Middlesex, London, UK
Ashley Whittington
Royal Cornwall Hospital (Tresliske), Truro, UK
Meme Wijesinghe
North Bristol NHS Trust, Bristol, UK
Martin Williams
St. Peter’s Hospital, Runnymede, UK
Stephen Winchester
Leicester Royal Infirmary, Leicester, UK
Martin Wiselka
Grantham and District Hospital, Grantham, UK
Adam Wolverson
Aintree University Hospital, Liverpool, UK
Daniel G. Wootton
North Tyneside General Hospital, North Shields, UK
Bryan Yates
Queen Elizabeth Hospital, King’s Lynn, UK
Peter Young
You can also search for this author in PubMed Google Scholar
PHOSP-COVID collaborative group
- Kathryn Abel
- , H. Adamali
- , Davies Adeloye
- , Oluwaseun Adeyemi
- , Rita Adrego
- , Laura Aguilar Jimenez
- , Shanaz Ahmad
- , N. Ahmad Haider
- , Rubina Ahmed
- , Nyarko Ahwireng
- , Mark Ainsworth
- , Asma Alamoudi
- , Mariam Ali
- , M. Aljaroof
- , Louise Allan
- , Richard Allen
- , Lisa Allerton
- , Lynne Allsop
- , Ann Marie Allt
- , Paula Almeida
- , Bashar Al-Sheklly
- , Danny Altmann
- , Maria Alvarez Corral
- , Shannon Amoils
- , David Anderson
- , Charalambos Antoniades
- , Gill Arbane
- , Ava Maria Arias
- , Cherie Armour
- , Lisa Armstrong
- , Natalie Armstrong
- , David Arnold
- , H. Arnold
- , A. Ashish
- , Andrew Ashworth
- , M. Ashworth
- , Shahab Aslani
- , Hosanna Assefa-Kebede
- , Paul Atkin
- , Catherine Atkin
- , Raminder Aul
- , Hnin Aung
- , Liam Austin
- , Cristina Avram
- , Nikos Avramidis
- , Marta Babores
- , Rhiannon Baggott
- , J. Bagshaw
- , David Baguley
- , Elisabeth Bailey
- , J. Kenneth Baillie
- , Steve Bain
- , Majda Bakali
- , E. Baldry
- , Molly Baldwin
- , David Baldwin
- , Clive Ballard
- , Amita Banerjee
- , Dongchun Bang
- , R. E. Barker
- , Laura Barman
- , Perdita Barran
- , Shaney Barratt
- , Fiona Barrett
- , Donna Basire
- , Neil Basu
- , Michelle Bates
- , R. Batterham
- , Helen Baxendale
- , Gabrielle Baxter
- , Hannah Bayes
- , M. Beadsworth
- , Paul Beckett
- , Paul Beirne
- , Murdina Bell
- , Robert Bell
- , Kaytie Bennett
- , Eva Beranova
- , Areti Bermperi
- , Anthony Berridge
- , Colin Berry
- , Sarah Betts
- , Emily Bevan
- , Kamaldeep Bhui
- , Michelle Bingham
- , K. Birchall
- , Lettie Bishop
- , Karen Bisnauthsing
- , John Blaikely
- , Angela Bloss
- , Annette Bolger
- , Charlotte Bolton
- , J. Bonnington
- , A. Botkai
- , Charlotte Bourne
- , Michelle Bourne
- , Kate Bramham
- , Lucy Brear
- , Jonathon Breeze
- , Katie Breeze
- , Andrew Briggs
- , E. Bright
- , Christopher Brightling
- , Simon Brill
- , K. Brindle
- , Lauren Broad
- , Andrew Broadley
- , Claire Brookes
- , Mattew Broome
- , Vanessa Brown
- , Ammani Brown
- , Angela Brown
- , Jeremy Brown
- , Terry Brugha
- , Nigel Brunskill
- , Phil Buckley
- , Anda Bularga
- , Ed Bullmore
- , Jenny Bunker
- , L. Burden
- , Tracy Burdett
- , David Burn
- , John Busby
- , Robyn Butcher
- , Al-Tahoor Butt
- , P. Cairns
- , P. C. Calder
- , Ellen Calvelo
- , H. Carborn
- , Bethany Card
- , Caitlin Carr
- , Liesel Carr
- , G. Carson
- , Penny Carter
- , Anna Casey
- , M. Cassar
- , Jonathon Cavanagh
- , Manish Chablani
- , Trudie Chalder
- , James D. Chalmers
- , Rachel Chambers
- , Flora Chan
- , K. M. Channon
- , Kerry Chapman
- , Amanda Charalambou
- , N. Chaudhuri
- , A. Checkley
- , Yutung Cheng
- , Luke Chetham
- , Caroline Childs
- , Edwin Chilvers
- , H. Chinoy
- , A. Chiribiri
- , K. Chong-James
- , N. Choudhury
- , Gaunab Choudhury
- , Phillip Chowienczyk
- , C. Christie
- , Melanie Chrystal
- , Cameron Clark
- , David Clark
- , Jude Clarke
- , S. Clohisey
- , G. Coakley
- , Zach Coburn
- , S. Coetzee
- , Joby Cole
- , Chris Coleman
- , Florence Conneh
- , David Connell
- , Bronwen Connolly
- , Lynda Connor
- , Amanda Cook
- , Shirley Cooper
- , B. Cooper
- , Josh Cooper
- , Donna Copeland
- , Tracey Cosier
- , Eamon Coughlan
- , Martina Coulding
- , C. Coupland
- , Thelma Craig
- , Daniele Cristiano
- , Michael Crooks
- , Andy Cross
- , Isabel Cruz
- , P. Cullinan
- , D. Cuthbertson
- , Luke Daines
- , Matthhew Dalton
- , Patrick Daly
- , Alison Daniels
- , Joanne Dasgin
- , Anthony David
- , Ffyon Davies
- , Ellie Davies
- , Kim Davies
- , Gareth Davies
- , Gwyneth Davies
- , Melanie Davies
- , Joy Dawson
- , Camilla Dawson
- , Enya Daynes
- , Anthony De Soyza
- , Bill Deakin
- , Andrew Deans
- , Joanne Deery
- , Sylviane Defres
- , Amanda Dell
- , K. Dempsey
- , Emma Denneny
- , J. Dennis
- , Ruvini Dharmagunawardena
- , Nawar Diar-Bakerly
- , Caroline Dickens
- , A. Dipper
- , Sarah Diver
- , Shalin Diwanji
- , Myles Dixon
- , R. Djukanovic
- , Hannah Dobson
- , S. L. Dobson
- , Annemarie B. Docherty
- , A. Donaldson
- , N. Dormand
- , Andrew Dougherty
- , Rachael Dowling
- , Stephen Drain
- , Katharine Draxlbauer
- , Katie Drury
- , Pearl Dulawan
- , A. Dunleavy
- , Sarah Dunn
- , Catherine Dupont
- , Joanne Earley
- , Nicholas Easom
- , Carlos Echevarria
- , Sarah Edwards
- , C. Edwardson
- , Claudia Efstathiou
- , Anne Elliott
- , K. Elliott
- , Yvette Ellis
- , Anne Elmer
- , Omer Elneima
- , Hosni El-Taweel
- , Teriann Evans
- , Ranuromanana Evans
- , Rachael A. Evans
- , Jonathon Evans
- , Cerys Evenden
- , Lynsey Evison
- , Laura Fabbri
- , Sara Fairbairn
- , Alexandra Fairman
- , K. Fallon
- , David Faluyi
- , Clair Favager
- , Tamanah Fayzan
- , James Featherstone
- , T. Felton
- , V. Ferreira
- , Selina Finney
- , J. Finnigan
- , L. Finnigan
- , Helen Fisher
- , S. Fletcher
- , Rachel Flockton
- , Margaret Flynn
- , David Foote
- , Amber Ford
- , D. Forton
- , Eva Fraile
- , C. Francis
- , Richard Francis
- , Susan Francis
- , Anew Frankel
- , Emily Fraser
- , N. French
- , Jonathon Fuld
- , J. Furniss
- , Lucie Garner
- , N. Gautam
- , John Geddes
- , J. George
- , P. George
- , Michael Gibbons
- , Rhyan Gill
- , Mandy Gill
- , L. Gilmour
- , F. Gleeson
- , Jodie Glossop
- , Sarah Glover
- , Nicola Goodman
- , Camelia Goodwin
- , Bibek Gooptu
- , Hussain Gordon
- , T. Gorsuch
- , M. Greatorex
- , Paul Greenhaff
- , William Greenhalf
- , Alan Greenhalgh
- , Neil J. Greening
- , John Greenwood
- , Rebecca Gregory
- , Heidi Gregory
- , D. Grieve
- , Denise Griffin
- , L. Griffiths
- , Anne-Marie Guerdette
- , Beatriz Guillen-Guio
- , Mahitha Gummadi
- , Ayushman Gupta
- , Sambasivarao Gurram
- , Elspeth Guthrie
- , Kate Hadley
- , Ahmed Haggar
- , Kera Hainey
- , Brigid Hairsine
- , Pranab Haldar
- , Lucy Hall
- , Mark Halling-Brown
- , Alyson Hancock
- , Kia Hancock
- , Neil Hanley
- , Sulaimaan Haq
- , Hayley Hardwick
- , Tim Hardy
- , Beverley Hargadon
- , Kate Harrington
- , Edward Harris
- , Victoria C. Harris
- , Ewen Harrison
- , Paul Harrison
- , Nicholas Hart
- , Alice Harvey
- , Matt Harvey
- , M. Harvie
- , L. Haslam
- , Claire Hastie
- , May Havinden-Williams
- , Jenny Hawkes
- , Nancy Hawkings
- , Jill Haworth
- , A. Hayday
- , Matthew Haynes
- , J. Hazeldine
- , Tracy Hazelton
- , Liam Heaney
- , Cheryl Heeley
- , Jonathon Heeney
- , M. Heightman
- , Simon Heller
- , Max Henderson
- , Helen Henson
- , L. Hesselden
- , Melanie Hewitt
- , Victoria Highett
- , T. Hillman
- , Ling-Pei Ho
- , Michaela Hoare
- , Amy Hoare
- , J. Hockridge
- , Philip Hogarth
- , Ailsa Holbourn
- , Sophie Holden
- , L. Holdsworth
- , D. Holgate
- , Maureen Holland
- , Leah Holloway
- , Katie Holmes
- , Megan Holmes
- , B. Holroyd-Hind
- , Anil Hormis
- , Alexander Horsley
- , Akram Hosseini
- , M. Hotopf
- , Linzy Houchen-Wolloff
- , Luke S. Howard
- , Kate Howard
- , Alice Howell
- , E. Hufton
- , Rachel Ann Hughes
- , Joan Hughes
- , Alun Hughes
- , Amy Humphries
- , Nathan Huneke
- , E. Hurditch
- , John Hurst
- , Masud Husain
- , Tracy Hussell
- , John Hutchinson
- , W. Ibrahim
- , Julie Ingham
- , L. Ingram
- , Diana Ionita
- , Karen Isaacs
- , Khalida Ismail
- , T. Jackson
- , Joseph Jacob
- , W. Y. James
- , Claire Jarman
- , Ian Jarrold
- , Hannah Jarvis
- , Roman Jastrub
- , Bhagy Jayaraman
- , Gisli Jenkins
- , P. Jezzard
- , Kasim Jiwa
- , C. Johnson
- , Simon Johnson
- , Desmond Johnston
- , Caroline Jolley
- , Ian Jones
- , Heather Jones
- , Mark Jones
- , Don Jones
- , Sherly Jose
- , Thomas Kabir
- , G. Kaltsakas
- , Vicky Kamwa
- , N. Kanellakis
- , Sabina Kaprowska
- , Zunaira Kausar
- , Natalie Keenan
- , Steven Kerr
- , Helen Kerslake
- , Angela Key
- , Fasih Khan
- , Kamlesh Khunti
- , Susan Kilroy
- , Bernie King
- , Clara King
- , Lucy Kingham
- , Jill Kirk
- , Paaig Kitterick
- , Paul Klenerman
- , Lucy Knibbs
- , Sean Knight
- , Abigail Knighton
- , Onn Min Kon
- , Samantha Kon
- , Ania Korszun
- , Ivan Koychev
- , Claire Kurasz
- , Prathiba Kurupati
- , Hanan Lamlum
- , G. Landers
- , Claudia Langenberg
- , Lara Lavelle-Langham
- , Allan Lawrie
- , Cathy Lawson
- , Claire Lawson
- , Alison Layton
- , Olivia C. Leavy
- , Ju Hee Lee
- , Elvina Lee
- , Karen Leitch
- , Rebecca Lenagh
- , Victoria Lewis
- , Joanne Lewis
- , Keir Lewis
- , N. Lewis-Burke
- , Felicity Liew
- , Tessa Light
- , Liz Lightstone
- , W. Lilaonitkul
- , S. Linford
- , Anne Lingford-Hughes
- , M. Lipman
- , Kamal Liyanage
- , Arwel Lloyd
- , Nazir I. Lone
- , Ronda Loosley
- , Janet Lord
- , Harpreet Lota
- , Wayne Lovegrove
- , Daniel Lozano-Rojas
- , Alice Lucey
- , Gardiner Lucy
- , E. Lukaschuk
- , Alison Lye
- , Ceri Lynch
- , S. MacDonald
- , G. MacGowan
- , Irene Macharia
- , J. Mackie
- , L. Macliver
- , S. Madathil
- , Gladys Madzamba
- , Nick Magee
- , Murphy Magtoto
- , N. Majeed
- , Flora Malein
- , Georgia Mallison
- , William Man
- , S. Mandal
- , K. Mangion
- , C. Manisty
- , R. Manley
- , Katherine March
- , Stefan Marciniak
- , Philip Marino
- , Myril Mariveles
- , Michael Marks
- , Elizabeth Marouzet
- , Sophie Marsh
- , M. Marshall
- , B. Marshall
- , Jane Martin
- , Adrian Martineau
- , L. M. Martinez
- , Nick Maskell
- , Darwin Matila
- , Wadzanai Matimba-Mupaya
- , Laura Matthews
- , Angeline Mbuyisa
- , Steve McAdoo
- , Hamish McAllister-Williams
- , Paul McArdle
- , Anne McArdle
- , Danny McAulay
- , Hamish J. C. McAuley
- , Gerry McCann
- , W. McCormick
- , Jacqueline McCormick
- , P. McCourt
- , Celeste McCracken
- , Lorcan McGarvey
- , Jade McGinness
- , K. McGlynn
- , Andrew McGovern
- , Heather McGuinness
- , I. B. McInnes
- , Jerome McIntosh
- , Emma McIvor
- , Katherine McIvor
- , Laura McLeavey
- , Aisling McMahon
- , Michael McMahon
- , L. McMorrow
- , Teresa Mcnally
- , M. McNarry
- , J. McNeill
- , Alison McQueen
- , H. McShane
- , Chloe Mears
- , Clare Megson
- , Sharon Megson
- , J. Meiring
- , Lucy Melling
- , Mark Mencias
- , Daniel Menzies
- , Marta Merida Morillas
- , Alice Michael
- , Benedict Michael
- , C. A. Miller
- , Lea Milligan
- , Nicholas Mills
- , Clare Mills
- , George Mills
- , L. Milner
- , Jane Mitchell
- , Abdelrahman Mohamed
- , Noura Mohamed
- , S. Mohammed
- , Philip Molyneaux
- , Will Monteiro
- , Silvia Moriera
- , Anna Morley
- , Leigh Morrison
- , Richard Morriss
- , A. Morrow
- , Paul Moss
- , Alistair Moss
- , K. Motohashi
- , N. Msimanga
- , Elizabeta Mukaetova-Ladinska
- , Unber Munawar
- , Jennifer Murira
- , Uttam Nanda
- , Heeah Nassa
- , Mariam Nasseri
- , Rashmita Nathu
- , Aoife Neal
- , Robert Needham
- , Paula Neill
- , Stefan Neubauer
- , D. E. Newby
- , Helen Newell
- , J. Newman
- , Tom Newman
- , Alex Newton-Cox
- , T. E. Nichols
- , Tim Nicholson
- , Christos Nicolaou
- , Debby Nicoll
- , Athanasios Nikolaidis
- , C. Nikolaidou
- , C. M. Nolan
- , Matthew Noonan
- , C. Norman
- , Petr Novotny
- , Kimon Ntotsis
- , Jose Nunag
- , Lorenza Nwafor
- , Uchechi Nwanguma
- , Joseph Nyaboko
- , Linda O’Brien
- , C. O’Brien
- , Natasha Odell
- , Kate O’Donnell
- , Godwin Ogbole
- , Olaoluwa Olaosebikan
- , Catherine Oliver
- , Zohra Omar
- , Peter J. M. Openshaw
- , D. P. O’Regan
- , Lorna Orriss-Dib
- , Lynn Osborne
- , Rebecca Osbourne
- , Marlies Ostermann
- , Charlotte Overton
- , Jamie Pack
- , Edmund Pacpaco
- , Stella-Maria Paddick
- , Sharon Painter
- , Erola Pairo-Castineira
- , Ashkan Pakzad
- , Sue Palmer
- , Padmasayee Papineni
- , K. Paques
- , Kerry Paradowski
- , Manish Pareek
- , Dhruv Parekh
- , H. Parfrey
- , Carmen Pariante
- , S. Parker
- , M. Parkes
- , J. Parmar
- , Sheetal Patale
- , Manish Patel
- , Suhani Patel
- , Dibya Pattenadk
- , M. Pavlides
- , Sheila Payne
- , Lorraine Pearce
- , John Pearl
- , Dan Peckham
- , Jessica Pendlebury
- , Yanchun Peng
- , Chris Pennington
- , Ida Peralta
- , Emma Perkins
- , Z. Peterkin
- , Tunde Peto
- , Nayia Petousi
- , John Petrie
- , Paul Pfeffer
- , Janet Phipps
- , S. Piechnik
- , John Pimm
- , Karen Piper Hanley
- , Riinu Pius
- , Hannah Plant
- , Tatiana Plekhanova
- , Megan Plowright
- , Krisnah Poinasamy
- , Oliver Polgar
- , Julie Porter
- , Joanna Porter
- , Sofiya Portukhay
- , Natassia Powell
- , A. Prabhu
- , James Pratt
- , Andrea Price
- , Claire Price
- , Carly Price
- , Anne Prickett
- , I. Propescu
- , J. Propescu
- , Sabrina Prosper
- , S. Pugmire
- , Sheena Quaid
- , Jackie Quigley
- , Jennifer K. Quint
- , H. Qureshi
- , I. N. Qureshi
- , K. Radhakrishnan
- , Najib Rahman
- , Markus Ralser
- , Betty Raman
- , Hazel Ramos
- , Albert Ramos
- , Jade Rangeley
- , Bojidar Rangelov
- , Liz Ratcliffe
- , Phillip Ravencroft
- , Konrad Rawlik
- , Anne Reddington
- , Heidi Redfearn
- , Dawn Redwood
- , Annabel Reed
- , Meryl Rees
- , Tabitha Rees
- , Karen Regan
- , Will Reynolds
- , Carla Ribeiro
- , A. Richards
- , Emma Richardson
- , M. Richardson
- , Pilar Rivera-Ortega
- , K. Roberts
- , Elizabeth Robertson
- , Leanne Robinson
- , Emma Robinson
- , Lisa Roche
- , C. Roddis
- , J. Rodger
- , Natalie Rogers
- , Gavin Ross
- , Alexandra Ross
- , Jennifer Rossdale
- , Anthony Rostron
- , Anna Rowe
- , J. Rowland
- , M. J. Rowland
- , A. Rowland
- , Sarah L. Rowland-Jones
- , Maura Roy
- , Igor Rudan
- , Richard Russell
- , Emily Russell
- , Gwen Saalmink
- , Ramsey Sabit
- , Beth Sage
- , T. Samakomva
- , Nilesh Samani
- , A. A. Samat
- , Claire Sampson
- , Katherine Samuel
- , Reena Samuel
- , Z. B. Sanders
- , Amy Sanderson
- , Elizabeth Sapey
- , Dinesh Saralaya
- , Jack Sargant
- , Carol Sarginson
- , Naveed Sattar
- , Kathryn Saunders
- , Peter Saunders
- , Ruth Saunders
- , Laura Saunders
- , Heather Savill
- , Avan Sayer
- , J. Schronce
- , William Schwaeble
- , Janet Scott
- , Kathryn Scott
- , Nick Selby
- , Malcolm G. Semple
- , Marco Sereno
- , Terri Ann Sewell
- , Kamini Shah
- , Ajay Shah
- , Manu Shankar-Hari
- , M. Sharma
- , Claire Sharpe
- , Michael Sharpe
- , Sharlene Shashaa
- , Alison Shaw
- , Victoria Shaw
- , Karen Shaw
- , Aziz Sheikh
- , Sarah Shelton
- , Liz Shenton
- , K. Shevket
- , Aarti Shikotra
- , Sulman Siddique
- , Salman Siddiqui
- , J. Sidebottom
- , Louise Sigfrid
- , Gemma Simons
- , Neil Simpson
- , John Simpson
- , Ananga Singapuri
- , Suver Singh
- , Claire Singh
- , Sally Singh
- , D. Sissons
- , J. Skeemer
- , Katie Slack
- , David Smith
- , Nikki Smith
- , Andrew Smith
- , Jacqui Smith
- , Laurie Smith
- , Susan Smith
- , M. Soares
- , Teresa Solano
- , Reanne Solly
- , A. R. Solstice
- , Tracy Soulsby
- , David Southern
- , D. Sowter
- , Mark Spears
- , Lisa Spencer
- , Fabio Speranza
- , Louise Stadon
- , Stefan Stanel
- , R. Steeds
- , N. Steele
- , Mike Steiner
- , David Stensel
- , G. Stephens
- , Lorraine Stephenson
- , Iain Stewart
- , R. Stimpson
- , Sue Stockdale
- , J. Stockley
- , Wendy Stoker
- , Roisin Stone
- , Will Storrar
- , Andrew Storrie
- , Kim Storton
- , E. Stringer
- , Sophia Strong-Sheldrake
- , Natalie Stroud
- , Christian Subbe
- , Catherine Sudlow
- , Zehra Suleiman
- , Charlotte Summers
- , C. Summersgill
- , Debbie Sutherland
- , D. L. Sykes
- , Nick Talbot
- , Ai Lyn Tan
- , Lawrence Tarusan
- , Vera Tavoukjian
- , Jessica Taylor
- , Abigail Taylor
- , Chris Taylor
- , John Paul Taylor
- , Amelie Te
- , Caroline Tee
- , J. Teixeira
- , Helen Tench
- , Sarah Terry
- , Susannah Thackray-Nocera
- , Favas Thaivalappil
- , David Thickett
- , David Thomas
- , S. Thomas
- , Caradog Thomas
- , Andrew Thomas
- , T. Thomas-Woods
- , A. A. Roger Thompson
- , Tamika Thompson
- , T. Thornton
- , Matthew Thorpe
- , Ryan S. Thwaites
- , Jo Tilley
- , N. Tinker
- , Gerlynn Tiongson
- , Martin Tobin
- , Johanne Tomlinson
- , Mark Toshner
- , T. Treibel
- , K. A. Tripp
- , Drupad Trivedi
- , E. M. Tunnicliffe
- , Alison Turnbull
- , Kim Turner
- , Sarah Turner
- , Victoria Turner
- , E. Turner
- , Sharon Turney
- , Lance Turtle
- , Helena Turton
- , Jacinta Ugoji
- , R. Ugwuoke
- , Rachel Upthegrove
- , Jonathon Valabhji
- , Maximina Ventura
- , Joanne Vere
- , Carinna Vickers
- , Ben Vinson
- , Ioannis Vogiatzis
- , Elaine Wade
- , Phillip Wade
- , Louise V. Wain
- , Tania Wainwright
- , Lilian Wajero
- , Sinead Walder
- , Samantha Walker
- , S. Walker
- , Tim Wallis
- , Sarah Walmsley
- , Simon Walsh
- , J. A. Walsh
- , Louise Warburton
- , T. J. C. Ward
- , Katie Warwick
- , Helen Wassall
- , Samuel Waterson
- , L. Watson
- , Ekaterina Watson
- , James Watson
- , M. Webster
- , J. Weir McCall
- , Carly Welch
- , Simon Wessely
- , Sophie West
- , Heather Weston
- , Helen Wheeler
- , Sonia White
- , Victoria Whitehead
- , J. Whitney
- , S. Whittaker
- , Beverley Whittam
- , V. Whitworth
- , Andrew Wight
- , James Wild
- , Martin Wilkins
- , Dan Wilkinson
- , Nick Williams
- , N. Williams
- , B. Williams
- , Jenny Williams
- , S. A. Williams-Howard
- , Michelle Willicombe
- , Gemma Willis
- , James Willoughby
- , Ann Wilson
- , Imogen Wilson
- , Daisy Wilson
- , Nicola Window
- , M. Witham
- , Rebecca Wolf-Roberts
- , Chloe Wood
- , F. Woodhead
- , Janet Woods
- , Dan Wootton
- , J. Wormleighton
- , J. Worsley
- , David Wraith
- , Caroline Wrey Brown
- , C. Wright
- , S. Wright
- , Louise Wright
- , Inez Wynter
- , Moucheng Xu
- , Najira Yasmin
- , S. Yasmin
- , Tom Yates
- , Kay Por Yip
- , Susan Young
- , Bob Young
- , A. J. Yousuf
- , Amira Zawia
- , Lisa Zeidan
- , Bang Zhao
- , Bang Zheng
- & O. Zongo
- , Daniel Agranoff
- , Ken Agwuh
- , Katie A. Ahmed
- , Dhiraj Ail
- , Erin L. Aldera
- , Ana Alegria
- , Beatrice Alex
- , Sam Allen
- , Petros Andrikopoulos
- , Brian Angus
- , Jane A. Armstrong
- , Abdul Ashish
- , Milton Ashworth
- , Innocent G. Asiimwe
- , Dougal Atkinson
- , Benjamin Bach
- , Siddharth Bakshi
- , Wendy S. Barclay
- , Shahedal Bari
- , Gavin Barlow
- , Samantha L. Barlow
- , Stella Barnass
- , Nicholas Barrett
- , Christopher Bassford
- , Sneha Basude
- , David Baxter
- , Michael Beadsworth
- , Jolanta Bernatoniene
- , John Berridge
- , Nicola Best
- , Debby Bogaert
- , Laura Booth
- , Pieter Bothma
- , Benjamin Brennan
- , Robin Brittain-Long
- , Katie Bullock
- , Naomi Bulteel
- , Tom Burden
- , Andrew Burtenshaw
- , Nicola Carlucci
- , Gail Carson
- , Vikki Caruth
- , Emily Cass
- , Benjamin W. A. Catterall
- , David Chadwick
- , Duncan Chambler
- , Meera Chand
- , Kanta Chechi
- , Nigel Chee
- , Jenny Child
- , Srikanth Chukkambotla
- , Richard Clark
- , Tom Clark
- , Jordan J. Clark
- , Emily A. Clarke
- , Sara Clohisey
- , Sarah Cole
- , Paul Collini
- , Marie Connor
- , Graham S. Cooke
- , Louise Cooper
- , Catherine Cosgrove
- , Audrey Coutts
- , Helen Cox
- , Jason Cupitt
- , Maria-Teresa Cutino-Moguel
- , Ana da Silva Filipe
- , Jo Dalton
- , Paul Dark
- , Christopher Davis
- , Chris Dawson
- , Thushan de Silva
- , Samir Dervisevic
- , Oslem Dincarslan
- , Alejandra Doce Carracedo
- , Cara Donegan
- , Lorna Donelly
- , Phil Donnison
- , Chloe Donohue
- , Gonçalo dos Santos Correia
- , Sam Douthwaite
- , Thomas M. Drake
- , Andrew Drummond
- , Marc-Emmanuel Dumas
- , Chris Dunn
- , Jake Dunning
- , Ingrid DuRand
- , Ahilanadan Dushianthan
- , Tristan Dyer
- , Philip Dyer
- , Angela Elliott
- , Cariad Evans
- , Anthony Evans
- , Chi Eziefula
- , Cameron J. Fairfield
- , Angie Fawkes
- , Chrisopher Fegan
- , Lorna Finch
- , Adam Finn
- , Lewis W. S. Fisher
- , Lisa Flaherty
- , Tom Fletcher
- , Terry Foster
- , Duncan Fullerton
- , Carrol Gamble
- , Isabel Garcia-Dorival
- , Atul Garg
- , Sanjeev Garg
- , Tammy Gilchrist
- , Michelle Girvan
- , Effrossyni Gkrania-Klotsas
- , Jo Godden
- , Arthur Goldsmith
- , Clive Graham
- , Tassos Grammatikopoulos
- , Christopher A. Green
- , Julian Griffin
- , Fiona Griffiths
- , Philip Gunning
- , Rishi K. Gupta
- , Katarzyna Hafezi
- , Sophie Halpin
- , Elaine Hardy
- , Ewen M. Harrison
- , Janet Harrison
- , Catherine Hartley
- , Stuart Hartshorn
- , Daniel Harvey
- , Peter Havalda
- , Daniel B. Hawcutt
- , Ross Hendry
- , Antonia Y. W. Ho
- , Maria Hobrok
- , Luke Hodgson
- , Karl Holden
- , Anthony Holmes
- , Peter W. Horby
- , Joanne Howard
- , Samreen Ijaz
- , Clare Jackson
- , Michael Jacobs
- , Susan Jain
- , Paul Jennings
- , Rebecca L. Jensen
- , Christopher B. Jones
- , Trevor R. Jones
- , Agilan Kaliappan
- , Vidya Kasipandian
- , Seán Keating
- , Stephen Kegg
- , Michael Kelsey
- , Jason Kendall
- , Caroline Kerrison
- , Ian Kerslake
- , Shadia Khandaker
- , Katharine King
- , Robyn T. Kiy
- , Stephen R. Knight
- , Susan Knight
- , Oliver Koch
- , Gouri Koduri
- , George Koshy
- , Chrysa Koukorava
- , Shondipon Laha
- , Eva Lahnsteiner
- , Steven Laird
- , Annette Lake
- , Suzannah Lant
- , Susan Larkin
- , Diane Latawiec
- , Andrew Law
- , James Lee
- , Gary Leeming
- , Daniella Lefteri
- , Tamas Leiner
- , Lauren Lett
- , Matthew Lewis
- , Sonia Liggi
- , Patrick Lillie
- , Wei Shen Lim
- , James Limb
- , Vanessa Linnett
- , Jeff Little
- , Lucia A. Livoti
- , Mark Lyttle
- , Louise MacGillivray
- , Alan Maclean
- , Michael MacMahon
- , Emily MacNaughton
- , Maria Mancini
- , Ravish Mankregod
- , Laura Marsh
- , Lynn Maslen
- , Hannah Massey
- , Huw Masson
- , Elijah Matovu
- , Nicole Maziere
- , Sarah McCafferty
- , Katherine McCullough
- , Sarah E. McDonald
- , Sarah McDonald
- , Laurence McEvoy
- , Ruth McEwen
- , John McLauchlan
- , Kenneth A. Mclean
- , Manjula Meda
- , Alexander J. Mentzer
- , Laura Merson
- , Soeren Metelmann
- , Alison M. Meynert
- , Nahida S. Miah
- , Joanna Middleton
- , Gary Mills
- , Jane Minton
- , Joyce Mitchell
- , Kavya Mohandas
- , James Moon
- , Elinoor Moore
- , Shona C. Moore
- , Patrick Morgan
- , Kirstie Morrice
- , Craig Morris
- , Katherine Mortimore
- , Samuel Moses
- , Mbiye Mpenge
- , Rohinton Mulla
- , Derek Murphy
- , Lee Murphy
- , Michael Murphy
- , Ellen G. Murphy
- , Thapas Nagarajan
- , Megan Nagel
- , Mark Nelson
- , Lisa Norman
- , Lillian Norris
- , Lucy Norris
- , Mahdad Noursadeghi
- , Michael Olanipekun
- , Wilna Oosthuyzen
- , Anthonia Osagie
- , Matthew K. O’Shea
- , Igor Otahal
- , Mark Pais
- , Massimo Palmarini
- , Carlo Palmieri
- , Selva Panchatsharam
- , Danai Papakonstantinou
- , Hassan Paraiso
- , Brij Patel
- , Natalie Pattison
- , William A. Paxton
- , Rebekah Penrice-Randal
- , Justin Pepperell
- , Mark Peters
- , Mandeep Phull
- , Jack Pilgrim
- , Stefania Pintus
- , Tim Planche
- , Daniel Plotkin
- , Georgios Pollakis
- , Frank Post
- , Nicholas Price
- , David Price
- , Tessa Prince
- , Rachel Prout
- , Nikolas Rae
- , Andrew Rambaut
- , Henrik Reschreiter
- , Tim Reynolds
- , Neil Richardson
- , P. Matthew Ridley
- , Mark Roberts
- , Stephanie Roberts
- , Devender Roberts
- , David L. Robertson
- , Alistair Rose
- , Guy Rousseau
- , Bobby Ruge
- , Clark D. Russell
- , Brendan Ryan
- , Debby Sales
- , Taranprit Saluja
- , Vanessa Sancho-Shimizu
- , Caroline Sands
- , Egle Saviciute
- , Matthias Schmid
- , Janet T. Scott
- , James Scott-Brown
- , Aarti Shah
- , Prad Shanmuga
- , Anil Sharma
- , Catherine A. Shaw
- , Victoria E. Shaw
- , Anna Shawcross
- , Rebecca K. Shears
- , Jagtur Singh Pooni
- , Jeremy Sizer
- , Benjamin Small
- , Richard Smith
- , Catherine Snelson
- , Tom Solomon
- , Rebecca G. Spencer
- , Nick Spittle
- , Shiranee Sriskandan
- , Nikki Staines
- , Tom Stambach
- , Richard Stewart
- , David Stuart
- , Krishanthi S. Subramaniam
- , Pradeep Subudhi
- , Olivia V. Swann
- , Tamas Szakmany
- , Agnieska Szemiel
- , Aislynn Taggart
- , Sarah Tait
- , Zoltan Takats
- , Panteleimon Takis
- , Jolanta Tanianis-Hughes
- , Kate Tatham
- , Richard S. Tedder
- , Jo Thomas
- , Jordan Thomas
- , Robert Thompson
- , Chris Thompson
- , Emma C. Thomson
- , Ascanio Tridente
- , Erwan Trochu
- , Darell Tupper-Carey
- , Lance C. W. Turtle
- , Mary Twagira
- , Nick Vallotton
- , Libby van Tonder
- , Rama Vancheeswaran
- , Rachel Vincent
- , Lisa Vincent-Smith
- , Shico Visuvanathan
- , Alan Vuylsteke
- , Sam Waddy
- , Rachel Wake
- , Andrew Walden
- , Ingeborg Welters
- , Murray Wham
- , Tony Whitehouse
- , Paul Whittaker
- , Ashley Whittington
- , Meme Wijesinghe
- , Eve Wilcock
- , Martin Williams
- , Lawrence Wilson
- , Stephen Winchester
- , Martin Wiselka
- , Adam Wolverson
- , Daniel G. Wootton
- , Andrew Workman
- , Nicola Wrobel
- , Bryan Yates
- , Peter Young
- , Maria Zambon
- & J. Eunice Zhang
Contributions
F.L. recruited participants, acquired clinical samples, analyzed and interpreted data and cowrote the manuscript, including all drafting and revisions. C.E. analyzed and interpreted data and cowrote this manuscript, including all drafting and revisions. S.F. and M.R. supported the analysis and interpretation of data as well as drafting and revisions. D.S., J.K.S., S.C.M., S.A., N.M., J.N., C.K., O.C.L., O.E., H.J.C.M., A. Shikotra, A. Singapuri, M.S., V.C.H., M.T., N.J.G., N.I.L. and C.C. contributed to acquisition of data underlying this study. L.H.-W., A.A.R.T., S.L.R.-J., L.S.H., O.M.K., D.G.W., T.I.d.S. and A. Ho made substantial contributions to conception/design and implementation of this work and/or acquisition of clinical samples for this work. They have supported drafting and revisions of the manuscript. E.M.H., J.K.Q. and A.B.D. made substantial contributions to the study design as well as data access, linkage and analysis. They have supported drafting and revisions of this work. J.D.C., L.-P.H., A. Horsley, B.R., K.P., M.M. and W.G. made substantial contributions to the conception and design of this work and have supported drafting and revisions of this work. J.K.B. obtained funding for ISARIC4C, is ISARIC4C consortium co-lead, has made substantial contributions to conception and design of this work and has supported drafting and revisions of this work. M.G.S. obtained funding for ISARIC4C, is ISARIC4C consortium co-lead, sponsor/protocol chief investigator, has made substantial contributions to conception and design of this work and has supported drafting and revisions of this work. R.A.E. and L.V.W. are co-leads of PHOSP-COVID, made substantial contributions to conception and design of this work, the acquisition and analysis of data, and have supported drafting and revisions of this work. C.B. is the chief investigator of PHOSP-COVID and has made substantial contributions to conception and design of this work. R.S.T. and L.T. made substantial contributions to the acquisition, analysis and interpretation of the data underlying this study and have contributed to drafting and revisions of this work. P.J.M.O. obtained funding for ISARIC4C, is ISARIC4C consortium co-lead, sponsor/protocol chief investigator and has made substantial contributions to conception and design of this work. R.S.T. and P.J.M.O. have also made key contributions to interpretation of data and have co-written this manuscript. All authors have read and approve the final version to be published. All authors agree to accountability for all aspects of this work. All investigators within ISARIC4C and the PHOSP-COVID consortia have made substantial contributions to the conception or design of this study and/or acquisition of data for this study. The full list of authors within these groups is available in Supplementary Information .
Corresponding authors
Correspondence to Ryan S. Thwaites or Peter J. M. Openshaw .
Ethics declarations
Competing interests.
F.L., C.E., D.S., J.K.S., S.C.M., C.D., C.K., N.M., L.N., E.M.H., A.B.D., J.K.Q., L.-P.H., K.P., L.S.H., O.M.K., S.F., T.I.d.S., D.G.W., R.S.T. and J.K.B. have no conflicts of interest. A.A.R.T. receives speaker fees and support to attend meetings from Janssen Pharmaceuticals. S.L.R.-J. is on the data safety monitoring board for Bexero trial in HIV+ adults in Kenya. J.D.C. is the deputy chief editor of the European Respiratory Journal and receives consulting fees from AstraZeneca, Boehringer Ingelheim, Chiesi, GSK, Insmed, Janssen, Novartis, Pfizer and Zambon. A. Horsley is deputy chair of NIHR Translational Research Collaboration (unpaid role). B.R. receives honoraria from Axcella therapeutics. R.A.E. is co-lead of PHOSP-COVID and receives fees from AstraZenaca/Evidera for consultancy on LC and from AstraZenaca for consultancy on digital health. R.A.E. has received speaker fees from Boehringer in June 2021 and has held a role as European Respiratory Society Assembly 01.02 Pulmonary Rehabilitation secretary. R.A.E. is on the American Thoracic Society Pulmonary Rehabilitation Assembly program committee. L.V.W. also receives funding from Orion pharma and GSK and holds contracts with Genentech and AstraZenaca. L.V.W. has received consulting fees from Galapagos and Boehringer, is on the data advisory board for Galapagos and is Associate Editor for the European Respiratory Journal . A. Ho is a member of NIHR Urgent Public Health Group (June 2020–March 2021). M.M. is an applicant on the PHOSP study funded by NIHR/DHSC. M.G.S. acts as an independent external and nonremunerated member of Pfizer’s External Data Monitoring Committee for their mRNA vaccine program(s), is Chair of Infectious Disease Scientific Advisory Board of Integrum Scientific LLC, and is director of MedEx Solutions Ltd. and majority owner of MedEx Solutions Ltd. and minority owner of Integrum Scientific LLC. M.G.S.’s institution has been in receipt of gifts from Chiesi Farmaceutici S.p.A. of Clinical Trial Investigational Medicinal Product without encumbrance and distribution of same to trial sites. M.G.S. is a nonrenumerated member of HMG UK New Emerging Respiratory Virus Threats Advisory Group and has previously been a nonrenumerated member of the Scientific Advisory Group for Emergencies (SAGE). C.B. has received consulting fees and/or grants from GSK, AstraZeneca, Genentech, Roche, Novartis, Sanofi, Regeneron, Chiesi, Mologic and 4DPharma. L.T. has received consulting fees from MHRA, AstraZeneca and Synairgen and speakers’ fees from Eisai Ltd., and support for conference attendance from AstraZeneca. L.T. has a patent pending with ZikaVac. P.J.M.O. reports grants from the EU Innovative Medicines Initiative 2 Joint Undertaking during the submitted work; grants from UK Medical Research Council, GSK, Wellcome Trust, EU Innovative Medicines Initiative, UK National Institute for Health Research and UK Research and Innovation–Department for Business, Energy and Industrial Strategy; and personal fees from Pfizer, Janssen and Seqirus, outside the submitted work.
Peer review
Peer review information.
Nature Immunology thanks Ziyad Al-Aly and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Ioana Staicu was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended data fig. 1 penalized logistic regression performance..
Graphs show classification error and Area under curve (AUC) from the 50 repeats tenfold nested cross-validation used to optimise and assess the performance of PLR testing associations with each LC outcome relative to Recovered (n = 233): Cardio_Resp (n = 398), Fatigue (n = 384), Anxiety/Depression (n = 202), GI (n = 132), ( e ) Cognitive (n = 6). The distributions of classification error and area under curve (AUC) from the nested cross-validation are shown. Box plot centre line represents the Median and boundaries of the box represent interquartile range (IQR), the whisker length represent 1.5xIQR.
Extended Data Fig. 2 Associations with long COVID symptoms in full study cohort.
( a ) Fibrinogen levels at 6 months were compared between pooled LC cases (n = 295) and Recovered (n = 233) and between the Cognitive group (n = 41) and Recovered (n = 233). Box plot centre line represent the Median and boundaries of the box represent interquartile range (IQR), the whisker length represents 1.5xIQR, any outliers beyond the whisker range are shown as individual dots. Median differences were compared using two-sided Wilcoxon signed-rank test *= p < 0·05, **= p < 0·01, ***= p < 0·001, ****= p < 0·0001. Unadjusted p-values are reported. b ) Distribution of time from COVID-19 hospitalisation at sample collection applying CDC and NICE definitions of LC (n = 719) ( c ) Upset plot of symptom groups. Horizontal coloured bars represent the number of patients in each symptom group: Cardiorespiratory (Cardio_Resp), Fatigue, Cognitive, Gastrointestinal (GI) and Anxiety/Depression (Anx_Dep). Vertical black bars represent the number of patients in each symptom combination group. To prevent patient identification, where less than 5 patients belong to a combination group, this has been represented as ‘<5’. The Recovered group (n = 250) were used as controls. Forest plots show Olink protein concentrations (NPX) associated with ( d ) Cardio_Resp (n = 398), ( e ) Fatigue (n = 342), ( f ) Anx_Dep (n = 219), ( g ) GI (n = 134), and ( h ) Cognitive (n = 65). Error bars represent the median accuracy of the model.
Extended Data Fig. 3 Validation of olink measurements using conventional assays in plasma.
Olink measured protein (NPX) were compared to chemiluminescence assays (ECL or ELISA, log2[pg/mL]) to validate our findings, where contemporaneously collected plasma samples were available (n = 58). Results from key mediators associated with LC groups were validated: CSF3, IL1R2, IL2, IL3RA, TNFa, TFF2. R = spearman rank correlation coefficient and shaded areas indicated the 95% confidence interval. Samples that fell below the lower limit of detection for a given assay were excluded and the ‘n’ value on each panel indicates the number of samples above this limit.
Extended Data Fig. 4 Univariate analysis of proteins associated with each symptom.
Olink measured plasma protein levels (NPX) compared between LC groups (Cardio_Resp, n = 398, Fatigue n = 384, Anxiety/Depression, n = 202, GI, n = 132 and Cognitive, n = 60) and Recovered (n = 233). Proteins identified by PLR were compared between groups. Median differences were compared using two-sided Wilcoxon signed-rank test. * = p < 0·05, ** = p < 0·01, *** = p < 0·001, ****= p < 0·0001 after FDR adjustment. Box plot centre line represent the Median and boundaries of the box represent interquartile range (IQR), the whisker length represents 1.5xIQR, any outliers beyond the whisker range are shown as individual dots.
Extended Data Fig. 5 Unadjusted Penalised Logistic Regression.
Olink measured proteins (NPX) and their association with Cardio_Resp (n = 398), Fatigue (n = 342), Anx_Dep (n = 219), GI (n = 134), and Cognitive (n = 65). Forest plots show odds of each LC outcome vs Recovered (n = 233), using PLR without adjusting for clinical co-variates. Error bars represent the median accuracy of the model.
Extended Data Fig. 6 Partial Least Squares analysis.
Olink measured proteins (NPX) and their association with Cardio_Resp (n = 398), Fatigue (n = 342), Anx_Dep (n = 219), GI (n = 134), and Cognitive (n = 65) groups. Forest plots show odds of LC outcome vs Recovered (n = 233), using PLS analysis. Error bars represent the standard error of the coefficient estimate.
Extended Data Fig. 7 Network analysis centrality.
Each graph shows the centrality score for each Olink measured protein (NPX) found to have significant associations with other proteins that were elevated in the Cardio_Resp (n = 398), Fatigue (n = 342), Anx_Dep (n = 219), GI (n = 134), and Cognitive (n = 65) groups relative to Recovered (n = 233).
Extended Data Fig. 8 Inflammation in men and women with long COVID.
Olink measured plasma protein levels (NPX) between men and women with symptoms, divided by age (<50 or >=50years): (a) shows IL1R2 and MATN2 in the Anxiety/Depression group (<50 n = 55, >=50 n = 133), (b) shows CTSO and NFASC in the Cognitive group (<50 n = 11, >=50 n = 50). Median values were compared between men and women using two-sided Wilcoxon signed-rank test. Box plot centre line represent the Median and boundaries represent interquartile range (IQR), the whisker length represents 1.5xIQR.
Extended Data Fig. 9 Inflammation in the upper respiratory tract.
Nasal cytokines measured by immunoassay in the CardioResp Group (n = 29) and Recovered (n = 31): ( a ) shows IL1a, IL1b, IL-6, APO-2, TGFa, TFF2. Median differences were compared using two-sided Wilcoxon signed-rank test. Box plot centre line represents the Median and boundaries of the box represent interquartile range (IQR), the whisker length represent 1.5xIQR. ( b ) Shows cytokines measured by immunoassay in paired plasma and nasal (n = 70). Correlations between IL1a, IL1b, IL-6, APO-2, TGFa and TFF2 in nasal and plasma samples were compared using Spearman’s rank correlation coefficient ( R ). Shaded areas indicated the 95% confidence interval of R.
Extended Data Fig. 10 Graphical abstract.
Summary of interpretation of key findings from Olink measured proteins and their association with CardioResp (n = 398), Fatigue (n = 342), Anx/Dep (n = 219), GI (n = 134), and Cognitive (n = 65) groups relative to Recovered (n = 233).
Supplementary information
Supplementary information.
Supplementary Methods, Statistics and reproducibility statement, Supplementary Results, Supplementary Tables 1–7, Extended data figure legends, Appendix 1 (Supplementary Table 8), Appendix 2 (PHOSP-COVID author list) and Appendix 3 (ISARIC4C author list).
Reporting Summary
Rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Liew, F., Efstathiou, C., Fontanella, S. et al. Large-scale phenotyping of patients with long COVID post-hospitalization reveals mechanistic subtypes of disease. Nat Immunol 25 , 607–621 (2024). https://doi.org/10.1038/s41590-024-01778-0
Download citation
Received : 11 August 2023
Accepted : 06 February 2024
Published : 08 April 2024
Issue Date : April 2024
DOI : https://doi.org/10.1038/s41590-024-01778-0
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
This article is cited by
Immune dysregulation in long covid.
- Laura Ceglarek
- Onur Boyman
Nature Immunology (2024)
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.

IMAGES
VIDEO
COMMENTS
Immunology is a broad branch of biomedical science that covers the study of all aspects of the immune system in all organisms. ... Explore the latest full-text research PDFs, articles, conference ...
Pillars of Immunology. The Journal of Immunology publishes commentaries highlighting scientific reports that were published more than 15 years ago and that have made a major impact on the course of immunological research today. Read published commentaries here . Don't miss the exciting Brief Review Collection, coming in the January 15, 2024 ...
Imbalanced immune responses are involved in developing preeclampsia (PE). We wish to explore the expression and potential changes of immune checkpoint molecules TIGIT, CD226 and CD155 in PE patients. Cui Li, Haiyan Liu and Zhongliang Duan. BMC Immunology 2024 25 :12. Research Published on: 7 February 2024.
Nature Immunology is a multidisciplinary journal that publishes papers of the highest quality and significance in all areas of immunology. Priority is given to work that provides fundamental ...
Abstract. Beyond structural and chemical barriers to pathogens, the immune system has two fundamental lines of defense: innate immunity and adaptive immunity. Innate immunity is the first immunological mechanism for fighting against an intruding pathogen. It is a rapid immune response, initiated within minutes or hours after aggression, that ...
IL-23 promotes tumor growth in preclinical cancer models and correlates with adverse clinical outcomes. Here, Becher and colleagues find that IL-23 produced by tumor-associated macrophages ...
Current Research in Immunology (CRIMMU) is an international peer-reviewed open access journal publishing original papers, short communications, and reviews that cover all aspects of molecular and cellular immunology. It is a companion to the highly regarded review journal Current Opinion in Immunology and is part of the Current Opinion and Research (CO+RE) suite of journals.
The mammalian immune system is a remarkable sensory system for the detection and neutralization of pathogens. History is replete with the devastating effects of plagues, and the coronavirus disease 2019 (COVID-19) pandemic is a defining global health crisis of our time. Although the development of effective vaccines has saved many lives, the ...
Several research groups and companies developed chimeric and humanized mAbs (Table 2), and these mAbs included additional modifications, such as changes in the carbohydrates (glycosylation) and/or ...
Harmonization of ANA testing challenge: quantification strategy to accurately predict end-point titers avoiding serial dilution. Immunologic Research is a transformative journal that covers the entire spectrum of immunology, from molecular and cellular levels to diseases and clinical ...
01 Apr 2024. 27 Mar 2024. 25 Mar 2024. 21 Mar 2024. 14 Mar 2024. Journal of Immunology Research provides a platform for scientists and clinicians working in different and diverse areas of immunology and therapy.
Cancer immunology is the study of interactions between the immune system and cancer cells, which is a rapid growing field of research that aims to identify biomarkers in cancer immunodiagnosis and to develop innovative cancer immunotherapeutic strategies. The immune response, including the recognition of cancer-specific antigens, is of ...
First release papers; Archive; ... Science Immunology; Research Article or Resource; clear all. Article Type. Research And Reviews 743; Peer Reviewed. Yes 743; Publication Date. 2016. 2024. ... NSF tests ways to improve research security without disrupting peer review. 5 Apr 2024 By . Elizabeth Finkel;
Welcome to the Special Collections page for Science Immunology. The Science Immunology editors have selected 7 papers published in Science Immunology that summarize and synthesize research on topics of broad interest within the field of immunology. Articles are freely available online to read until the end of June 2024.
Researchers employ advanced technologies to uncover promising biomarkers with implications for treatment response and prognosis in cancer patients. Systematic reviews and original papers shed light on the multifaceted landscape of tumor immunology, exploring biomarkers ranging from traditional immunological markers to emerging candidate biomarkers.
Immunology is the branch of biomedical sciences concerned with all aspects of the immune system in all multicellular organisms. Profiling of plasma proteins in individuals with COVID-19 shows that ...
In addition, Scopus and Web of Science search engines are used for reliable results. For searching relevant articles, keywords used are healthcare AI, immunology, deep learning, machine learning, etc. The rest of the paper is organized as follows. Section 4 provides a discussion on what are health diseases and their subcategorizations.
From innovations in basic immunology and structural biology to treatments for immune-mediated diseases and cancer, the conceptual and technological advances resulting from HIV/AIDS research have had an enormous impact on the research and public and global health communities over and above the field of HIV/AIDS.
Mixture-of-Depths:Dynamicallyallocatingcomputeintransformer-basedlanguagemodels We leverage an approach akin to Mixture of Experts (MoE) transformers, in which dynamic
In major dysmood disorder, physiosomatic, chronic fatigue and fibromyalgia symptoms are driven by immune activation and increased immune-associated neurotoxicity. Michael Maes. , Abbas F. Almulla ...
National Bureau of Economic Research
A lot of research 10 - 12 has been geared toward the understanding of this challenge, but the complexity of it requires a different dimension of approach. 13 An approach that must accommodate many factors affecting vaccine development like pathogen antigenic variability, the emergence of infectious disease, human genetic variation is the goal ...
Eran Ophir, Ph.D., Chief Scientific Officer at Compugen added, "Our paper published online yesterday in Cancer Immunology Research describes how COM503, a potential first-in-class high affinity ...
This Review, aimed at a broad scientific audience, provides an introductory guide to the history, development and immunological basis of vaccines, immunization and related issues to provide ...
View PDF HTML (experimental) Abstract: Scientific publishing lays the foundation of science by disseminating research findings, fostering collaboration, encouraging reproducibility, and ensuring that scientific knowledge is accessible, verifiable, and built upon over time. Recently, there has been immense speculation about how many people are using large language models (LLMs) like ChatGPT in ...
View a PDF of the paper titled ReALM: Reference Resolution As Language Modeling, by Joel Ruben Antony Moniz and 7 other authors. View PDF HTML (experimental) Abstract: Reference resolution is an important problem, one that is essential to understand and successfully handle context of different kinds. This context includes both previous turns ...
Other grants that have supported this work include the UK Coronavirus Immunology Consortium (funder reference 1257927), the Imperial Biomedical Research Centre (NIHR Imperial BRC, grant IS-BRC ...